Impact Factor ISSN: 1837-9664
J Cancer 2024; 15(1):275-292. doi:10.7150/jca.89338 This issue Cite
Research Paper
Interplay of Sphingolipid Metabolism in Predicting Prognosis of GBM Patients: Towards Precision Immunotherapy
1. Department of Gastroenterology, Affiliated Hospital of Jiangsu University, Jiangsu University, Zhenjiang, China.
2. Department of Neurosurgery, the Affiliated Hospital of Youjiang Medical University for Nationalities, Guangxi, China.
3. Department of Cell Biology, School of Medicine, Jiangsu University, Zhenjiang, China.
4. School of Clinical Medicine, Guizhou Medical University, Guiyang, China.
5. Faculty of Dentistry, University of Debrecen, Debrecen, Hungary.
6. Department of Oral and Maxillofacial-Head and Neck Oncology, Shanghai Ninth People's Hospital, Shanghai Jiao Tong University School of Medicine, College of Stomatology, Shanghai Jiao Tong University, Shanghai, China.
7. National Center for Stomatology & National Clinical Research Center for Oral Diseases, Shanghai, China.
8. Shanghai Key Laboratory of Stomatology, Shanghai, China.
9. Department of Clinical Veterinary Medicine, Huazhong Agricultural University, Wuhan, China.
10. Faculty of Medicine, University of Debrecen, Debrecen, Hungary.
11. Xinxiang Medical University, Xinxiang, China.
12. Department of Endocrinology, Affiliated Hospital of Jiangsu University, Jiangsu University, Zhenjiang, China.
#Equal Contribution: Qi Wang, Chuanhua Zheng, Hanjin Hou, Xin Bao
Abstract
Background: In spite of numerous existing bio-surveillance systems for predicting glioma (GBM) prognosis, enhancing the efficacy of immunotherapy remains an ongoing conundrum. The continual scrutiny of the dynamic interplay between the sphingolipid metabolic pathway and tumor immunophenotypes has unveiled potential implications. However, the intricate orchestration of functional and regulatory mechanisms by long non-coding RNAs (lncRNAs) in GBM, particularly in the context of sphingolipid metabolism, remains cryptic.
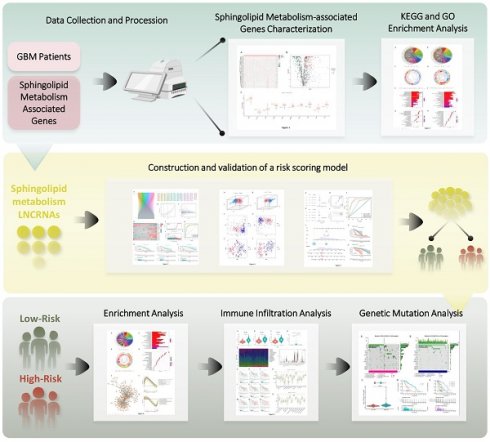
Methods: We harnessed established R packages to intersect gene expression profiles of GBM patients within the The Cancer Genome Atlas (TCGA) database with the compilation of sphingolipid metabolism genes from GeneCards. This enabled us to discern markedly distinct lncRNAs, which were subsequently deployed to construct a robust prognostic model utilizing Lasso-Cox regression analysis. We then scrutinized the immune microenvironment across various risk strata using the ssGSEA and CIBERSORT algorithms. To evaluate mutation patterns and drug resistance profiles within patient subgroups, we devised the "Prophytic" and "Maftools" packages, respectively.
Results: Our investigation scrutinized lncRNAs linked to sphingolipid metabolism, utilizing glioma specimens from TCGA. We meticulously curated 1224 sphingolipid-associated genes gleaned from GeneCards and pinpointed 272 differentially expressed mRNAs via transcriptomic analysis. Enrichment analyses underscored their significance in sphingolipid processes. A prognostic model founded on 17 meticulously selected lncRNAs was systematically constructed and validated. This model adeptly stratified GBM patients into high- and low-risk categories, yielding highly precise prognostic insights. We also discerned correlations between immune cell infiltration and genetic mutation discrepancies, along with distinct therapeutic responses through drug sensitivity analysis. Notably, computational findings were corroborated through experimental validation by RT-PCR.
Conclusion: In summation, our exhaustive inquiry underscores the multifaceted utility of the sphingolipid metabolic pathway as an autonomous diagnostic and prognostic indicator for glioma patients. Furthermore, we amalgamate a profusion of substantiated evidence concerning immune infiltration and gene mutations, thereby reinforcing the proposition that sphingolipid metabolism may function as a pivotal determinant in the panorama of immunotherapeutic interventions.
Keywords: Sphingolipid, lncRNAs, GBM, Precision immunotherapy, Biomarkers

 Global reach, higher impact
Global reach, higher impact