Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(7):2151-2164. doi:10.7150/jca.48045 This issue Cite
Research Paper
Comprehensive investigation of the clinical significance of long non-coding RNA HOXA-AS2 in acute myeloid leukemia using genome-wide RNA sequencing dataset
1. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning, 530021, Guangxi Zhuang Autonomous Region, People's Republic of China.
2. Department of Hepatobiliary Surgery, The First Affiliated Hospital of Guangxi Medical University, Nanning, 530021, Guangxi Zhuang Autonomous Region, People's Republic of China.
Abstract
Objective: The present study aimed to determine the prognostic value of HOXA cluster antisense RNA2 (HOXA-AS2) in acute myeloid leukemia (AML), and to explore its potential molecular mechanisms. We also screening of potential drugs targeting HOXA-AS2 in AML.
Methods: The level 3 raw genome-wide RNA sequencing dataset of AML was download from The Cancer Genome Atlas (TCGA) Data Portal, and the potential molecular mechanisms and drugs prediction of HOXA-AS2 in AML were explored using multiple bioinformatics analysis approaches.
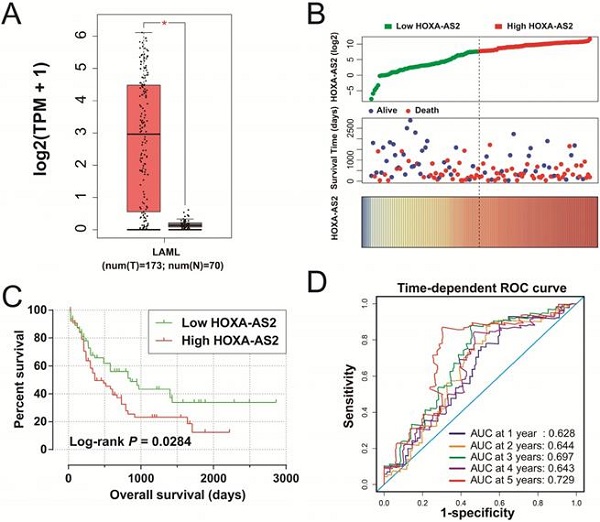
Results: TCGA AML cohort dataset indicated that HOXA-AS2 was significantly up-regulated in AML bone marrow tissues, and high HOXA-AS2 expression was related to poor overall survival (log-rank P=0.0284, hazard ratio 1.640, 95% confidence interval 1.046-2.573). Functional enrichment of differentially expressed genes (DEGs) suggested that the difference in prognosis between AML patients with high- and low-HOXA-AS2 expression may be due to differences in biological processes and pathways, including cell adhesion, angiogenesis, mitogen-activated protein kinase, cell differentiation, and other biological processes, and phosphatidylinositol 3 kinase-protein kinase B and Wnt signaling pathways. We also screened out three potential HOXA-AS2-targeted therapeutic drugs for AML, megestrol, carmustine, and cefoxitin, based on these DEGs. Functional enrichment analysis of HOXA-AS2-co-expressed genes revealed that HOXA-AS2 may act a part in AML by regulating nuclear factor-κB transcription factor activity, DNA methylation, angiogenesis, apoptosis, cell migration, Toll-like receptor 4, and Wnt signaling pathways.
Conclusion: Our findings suggest that HOXA-AS2 is up-regulated in the bone marrow in patients with AML, and may serve as a novel prognostic biomarker for AML.
Keywords: HOXA-AS2, acute myeloid leukemia, The Cancer Genome Atlas, molecular mechanism, drug prediction

 Global reach, higher impact
Global reach, higher impact