Impact Factor ISSN: 1837-9664
J Cancer 2022; 13(10):2988-2999. doi:10.7150/jca.72789 This issue Cite
Research Paper
Circ_0039908/miR-let-7c/RRM2 axis was identified played an important role in lung adenocarcinoma by integrated analysis
1. Department of Pulmonary and Critical Care Medicine, the First Affiliated Hospital of Soochow University, Suzhou, 215006, China.
2. Suzhou Key Laboratory for Respiratory Diseases, Suzhou, 215006, China.
3. Institute of Respiratory Diseases, Soochow University, Suzhou, 215006, China.
4. Department of Thoracic Surgery, The First Affiliated Hospital of Soochow University, Medical College of Soochow University, Suzhou, China.
# These authors contributed equally: Weijie Zhang and Jun Chen
Abstract
Background: Circular RNAs (circRNAs) are shown to play a significant role in cancer initiation and progression by interacting on microRNAs (miRNAs) which act as one kind of competing endogenous RNAs (ceRNAs) for the regulation effect on target gene expressions. This study was performed to explore the prognosis-related circRNAs in lung adenocarcinoma (LUAD) patients by integrated analysis and find the mechanism it worked.
Methods: The miRNAs and mRNAs, accompanied with circRNAs expressions were obtained through The Cancer Genome Atlas (TCGA) and the Gene Expression Omnibus (GEO) database, The cytoHubba app of Cytoscape was used to identify hubgenes. Quantitative real-time PCR (q-RT PCR) was performed to identify the expression of circRNA, miRNA and mRNA, Cell Counting Kit-8 (CCK-8) and clone formation assays were used to evaluate the proliferation ability of different kinds of cells in vitro. Transwell assays were utilized to assess the motility of tumor cells.
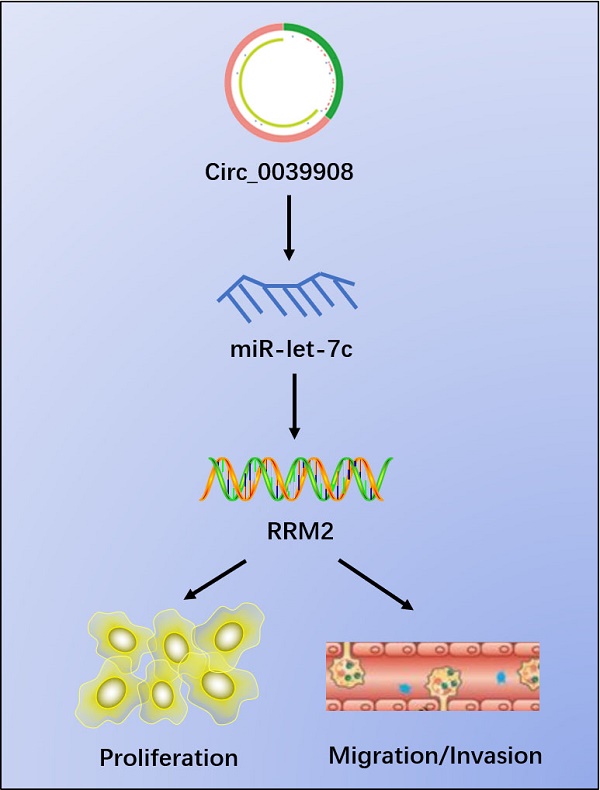
Results: Finally, circRNA_0039908/let7c-5p/RRM2 axis was identified in our research, it can play an important role in the LUAD pathogenesis progression and we found that the proliferation, invasion and migration abilities of LUAD cells can be suppressed after knockdown of circRNA_0039908. This work indicates that circRNA_0039908/let7c-5p/RRM2 axis may be a promising target in the prognosis and treatment of LUAD patients.
Conclusions: Circ_0039908/miR-let-7c/RRM2 axis can promote the ability of proliferation, migration and invasion of LUAD cells.
Keywords: Lung adenocarcinoma, circRNA, competitive endogenous RNA, bioinformatics analysis

 Global reach, higher impact
Global reach, higher impact