Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(19):5929-5937. doi:10.7150/jca.57951 This issue Cite
Review
Research progress, challenges and perspectives on PNPLA3 and its variants in Liver Diseases
Institute of Interdisciplinary Integrative Medicine Research, Shanghai University of Traditional Chinese Medicine, Shanghai 201203, China.
Received 2021-1-7; Accepted 2021-7-26; Published 2021-8-13
Abstract
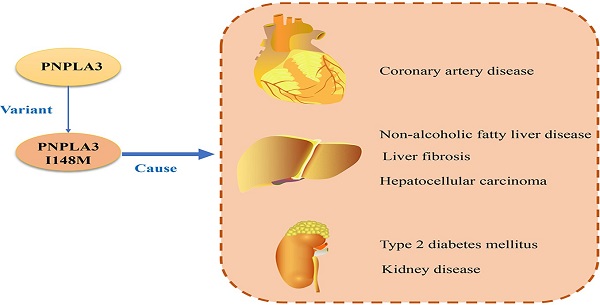
The human patatin-like phospholipase domain-containing 3 gene (PNPLA3) is highly expressed in liver and adipose tissue and encodes a transmembrane polypeptide chain containing 481 amino acids. The I148M variant of PNPLA3 is a single nucleotide polymorphism, which is related to a variety of liver and cardiovascular diseases and their complications (such as non-alcoholic fatty liver disease, liver fibrosis, coronary artery disease). This review mainly describes the pathophysiological effects of PNPLA3 and its variants, and their roles in the progression of liver disease and its complications.
Keywords: PNPLA3, NAFLD, type 2 diabetes, liver fibrosis, hepatocellular carcinoma
Introduction
Liver diseases have a significant impact on people's quality of life. Deaths caused by cirrhosis and liver cancer account for 3.5% of the global mortality rate [1]. The occurrence of liver disease has a complicated relationship with environmental and genetic factors. Patatin-like phospholipase domain-containing 3 gene (PNPLA3) and its variants (PNPLA3 I148M) have been confirmed to be closely related to the occurrence of liver disease. The human PNPLA3 is a protein that contains a patatin-like phospholipase domain and has nine types [2]. Patatin, as the main protein of potato tuber, has nonspecific lipoacyl hydrolase activity [3, 4]. In 2001, Baulande et al. [5] identified genes that participated in the reprogramming of gene expression during the adipose differentiation of the murine 3T3 preadipocyte cell line, and it was then that PNPLA3 was first identified and cloned.
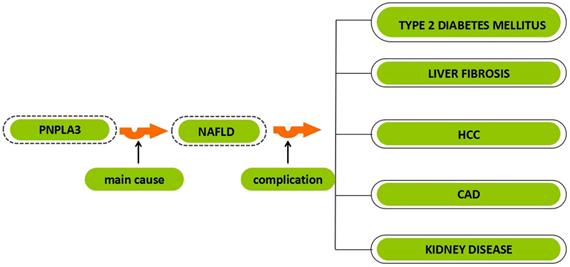
This article reviews the discovery and role shaping research progress and challenges about PNPLA3, and mainly elaborates the role of a non-synonymous variant of PNPLA3 (rs738409, I148M) in liver disease and related diseases. We also summarized the discovery and history of knowledge about PNPLA3 (Figure 1, Ttable 1).
The relationship between PNPLA3 and different diseases. PNPLA3, patatin-like phospholipase domain-containing 3 gene; NAFLD, non-alcoholic fatty liver disease; HCC, hepatocellular carcinoma; CAD, coronary artery disease.
Structure of PNPLA3
The PNPLA3 gene encodes a protein of 481 amino acids [6], and it is highly expressed in liver and adipose tissues. In humans, PNPLA3 gene is located on the long arm (q) and (22q. 13.31) of chromosome 22. Its molecular weight is predicted to be 52.8 kDa [5]. It has 9 exons and a transcription length of 2805 base pairs [7]. The N-terminal of PNPLA3 contains a patatin like domain with the common sequence of ser-Asp catalytic dimer (Gly-X-Ser-X-Gly and Asp-X-Gly/Ala) [8]. The non-synonymous variation of PNPLA3 (rs738409, I148M) is caused by the substitution of isoleucine (I) with methionine (M) in the amino acid coding sequence 148 (I148M). PNPLA3 I148M has received widespread attention due to its high correlation with the risk of non-alcoholic fatty liver disease (NAFLD) [9, 10]. The wild-type PNPLA3 is mainly distributed between membrane and lipid droplets, and closely bound with membrane. The allocation and positioning of PNPLA3 I148M have not changed significantly [11, 12].
Physiological function of PNPLA3
The PNPLA3 protein is also called adiponutrin. In the liver, it is mainly found in hepatocytes [13], stellate cells [14, 15] and sinusoidal cells [16]. PNPLA3 levels in stellate cells were lower than stellate hepatocytes [17]. Through biochemical grading studies, it was found that PNPLA3 is mostly present in lipid droplets in liver cells [11], and PNPLA3 I148M has relation to droplets of larger size with reduced triglyceride (TG) hydrolysis, as shown by recent studies [18, 19]. Thus, PNPLA3 protein is mainly connected with lipid droplets of hepatocytes.
In terms of the role of PNPLA3, it was thought to help regulated the development of adipocytes and the production and breakdown of fats in hepatocytes and adipocytes, through the processes known as lipogenesis and lipolysis. PNPLA3 protein has lipase activity and can hydrolyze triglycerides. This lipase activity is mainly for monounsaturated fatty acids and polyunsaturated fatty acids [12, 20]. Mitsche et al. found that the transfer of vLCPUFAs from TG to phospholipids (PLs) in the liver requires the participation in PNPLA3. This effect was confirmed in PNPLA3 inactivated mice and PNPLA3-/- mice [21]. On the other hand, PNPLA3 also participates in the remodeling of TG and PL in the form of vLCPUFA-specific TG hydrolase [21]. Chen et al. [22] indicated that PNPLA3 mRNA levels decreased after fasting and that PNPLA3 expression increased with refeeding of mice by both insulin and glucose [23]. It was also shown that the PNPLA3 was used to help process and store fats in the diet. In the liver, levels of PNPLA3 were positively correlated with body mass index (BMI) [24]. However, some studies have found that after knocking out PNPLA3, the TG content and TG hydrolysis in fat and liver tissues did not change significantly [25, 26]. At the same time, PNPLA3 also had lysophosphatidic acid acyltransferase activity (LPAAT) [12]. In some studies, it was mentioned that the change of LPAAT activity of PNPLA3 was related to the accumulation of TG. They proposed that the change of TG in lipid droplets was not related to its lipase activity, but was mainly determined by its LPAAT activity [27]. BasuRay et al. [28] denied that the accumulation of TG was related to the activity of LPAAT. They proposed that PNPLA3 regulated the accumulation of TG in lipid droplets (LDs) through ubiquitination. From the above studies, it was found that the mechanism of PNPLA3 regulating TG was complicated. In other studies, it was also found that PNPLA3 also had retinyl esterase activity and participates in the metabolism of vitamin A in the body [29, 30]. In addition, Huang et al. [17] revealed that PNPLA3 achieves nutritional control through a feed-forward loop, which started from transcriptional upregulation and expands through coordinated inhibition of protein degradation.
Study on the related mechanism of PNPLA3 and its variants in promoting NAFLD
| Key Finding | Year | Reference |
|---|---|---|
| Found a non-synonymous variant of PNPLA3, PNPLA3 (rs738409, I148M). | 2008 | [35] |
| PNPLA3 deletion is not associated with fatty liver, lipid homeostasis and insulin resistance in mice. | 2010, 2011 | [25,26] |
| PNPLA3 I148M significantly reduces the activity of TAGs and DAG hydrolase and changes the liver TAG-fatty acid profile. | 2012 | [44] |
| The PNPLA3-I148M variant loses its triglyceride hydrolysis activity. | 2010, 2015, 2016 | [11,45,46] |
| The I148M variant showed increased LPAAT activity leading to an increase in cellular lipid accumulation. | 2012 | [27] |
| PNPLA3- I148M can avoid ubiquitination and proteasome degradation, resulting in a decrease in TG mobilization in LDs. | 2017 | [28] |
| PNPLA3 promotes the transfer of vLCPUFAs from TG to PLs in liver lipid droplets, and PNPLA3-I148M enhances this effect. | 2018 | [21] |
PNPLA3, patatin-like phospholipase domain-containing protein 3; TAG, triacylglyceride; DAG, diacylglycerol; LPAAT, lysophophosphatidic acid acyltransferase; TG, triacylglycerol; LDs, lipid droplets; vLCPUFAs, very long-chain polyunsaturated fatty acids; PLs, phospholipids.
PNPLA3 I148M and the occurrence of diseases. The orange arrow represents the role of PNPLA3, and the green arrow represents the role of PNPLA3 I148M. PNPLA3, patatin-like phospholipase domain-containing 3 gene; NAFLD, non-alcoholic fatty liver disease; HCC, hepatocellular carcinoma;T2DM, type 2 diabetes mellitus; CAD, coronary artery disease; CKD, chronic kidney disease.
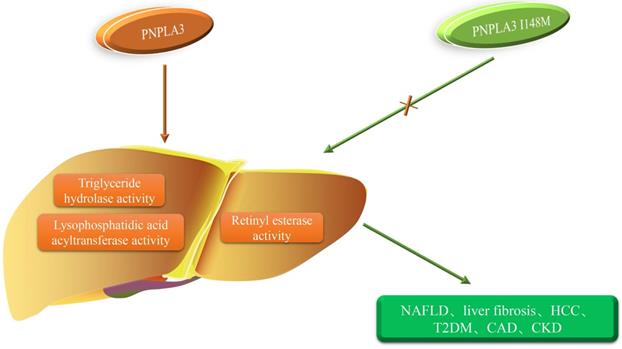
PNPLA3 and liver disease
PNPLA3 and NAFLD
The occurrence of NAFLD is related to many factors, mainly manifested as the accumulation of TG in liver cells and symptoms related to metabolic syndrome [31]. NAFLD initially manifests as liver steatosis, which may develop into non-alcoholic steatohepatitis (NASH). It has the following characteristics: hepatocyte damage, hepatocyte swelling ('ballooning'), inflammatory infiltrate and the early development of perisinusoidal ('chicken wire') fibrosis. If the damage to the liver continues to increase, there is a high probability of causing liver cirrhosis, liver decompensation, liver cancer, etc. [32].
A study using PNPLA3-KO mice found that when endoplasmic reticulum stresses occured, PNPLA3 participated in liver fatty acid metabolism and triglyceride accumulation [33]. Another study found that when mice were given a high-fat diet, the level of PNPLA3 in the liver increased significantly, suggesting that PNPLA3 may be involved in lipid metabolism under conditions of excess lipids [34]. But most studies have shown that the occurrence of NAFLD is related to rs738409. Romeo et al. [35] found a non-synonymous variant of PNPLA3 (rs738409, I148M) in a 2008 study, which was a major risk factor for the development of NAFLD. They suggested that the incidence of PNPLA3 I148M in the Hispanic population was higher than that of European Americans and African Americans [35]. Hotta et al. [36] genotyped 253 NAFLD patients and 578 control subjects using TaqMan analysis. They found that individuals with a G-allele of rs738409 were more likely to develop NAFLD in the Japanese population. Moreover, rs738409 individuals also had higher plasma alanine aminotransferase (ALT), aspartate aminotransferase (AST), ferritin, and histological fibrosis stages. Jain et al. [37] compared 218 obese Asian Indian adolescents with 86 healthy lean people without fatty liver and found that rs738409 C>G of PNPLA3 may be closely related to obesity and NAFLD in Asian Indian adolescents. Similar experiments were conducted in America [38], Italy [39], and Malaysia [40], and their results were similar.
Sookoian et al. [41] started to study whether PNPLA3 I148M was related to the severity of NAFLD in 2011. Through histological evaluation, they found that PNPLA3 I148M and NAFLD severity had a strong positive correlation that was independent of BMI, sex, age and insulin sensitivity. Recently, Arida et al. [42] performed NAFLD liver fat score and fibrosis score on 13,298 adults without viral hepatitis who had fasted for at least 4 hours. The results showed that patients with PNPLA3 I148M genotype were more likely to have liver fat and fibrosis, and have a higher mortality rate from liver disease. Studies have also shown that NAFLD patients with the PNPLA3 rs738409 G allele carrying the PNPLA3 148M variant have a subtle response to treatment involving dipeptidyl peptidase-4 inhibitors, lifestyle changes and bariatric surgery [43]. This suggests that the genetic variant PNPLA3 I148M may be used as one of the markers for monitoring NAFLD.
The key mechanism of PNPLA3-148M causing steatosis is very complicated. Generally speaking, it is related to the relative or absolute increase of TG accumulation in the liver. The first mechanism is related to the lipase activity of PNPLA3. Studies have shown that the PNPLA3-148M variant has reduced TG hydrolase activity and increased TG accumulation in the liver. Huang et al. [20] found that the mutation of PNPLA3 at position 148 (PNPLA3 I148M) would greatly reduce the activity of triacylglycerols (TAGs) and diacyl-glycerol (DAG) hydrolases. Other studies suggested that the accumulation of TAG in the liver of PNPLA3 I148M transgenic mice was not only an increase in TAG synthesis [44]. The I148M variant can promote liver steatosis by changing the liver TAG-fatty acid profile and increasing fatty acid formation [44]. He et al. [11] found through in vitro experiments that PNPLA3-I148M lost the activity of hydrolyzing triglycerides, causing TGs to accumulate in the liver and gradually form fatty degeneration. In several other studies, it was also found that overexpression of PNPLA3-148M in mice and humans could promote the occurrence of steatosis [45, 46]. The second mechanism is that PNPLA3-148M will increase the activity of LPAAT to promote the synthesis of TG. Kumari et al. [27] found that the I148M variant had higher LPAAT activity than the wild type. It is worth noting that the LPAAT activity of this variant was much higher than the TG hydrolase activity [27]. The third mechanism is related to the reduced ubiquitination of I148M. BasuRay et al. [28] found that the main catabolic pathway of PNPLA3 was related to ubiquitination and proteasome degradation. After I148M transgenic mice were treated with the proteasome inhibitor bortezomib, the ubiquitination ratio of PNPLA3 was much lower than that of WT mice [28]. They proposed that PNPLA3-148M can avoid ubiquitination and proteasome degradation, resulting in the accumulation of PNPLA3 in the liver and the reduction of TG mobilization in LDs [28, 47]. In short, PNPLA3 I148M, as a high-risk factor for NAFLD, will be a research focus on the treatment of NAFLD in the future.
PNPLA3 and liver fibrosis
Fibrosis is the end result of many inflammatory and tissue repair reactions, as well as most chronic inflammatory diseases [48], manifested by extracellular Matrix [49]. The causes of liver fibrosis include alcoholic liver injury (ALD), NAFLD and hepatitis.
ALD is very common in liver disease. Alcoholic fatty liver (AFL) can develop into alcoholic steatohepatitis (ASH), which is a disease with inflammatory changes in the liver. The long-term existence of ASH will eventually lead to fibrosis and cirrhosis, and even develop into hepatocellular carcinoma (HCC) [50]. Kolla et al. [51] conducted a case-control study on patients of European ancestry with a history of ALD, and genotyping single pairs of nucleotide polymorphisms (SNPs), and determined that PNPLA3 SNP rs738409 has a strong correlation between the occurrences of ALD. Trépo et al. [52] selected 328 healthy people and 330 ALD patients in the rs738409 polymorphism genotyping study among European whites, and found that the frequency of rs738409 G alleles in the control group was significantly lower than that of ALD, and rs738409 G alleles genes were also significantly connected with steatosis, fibrosis, and a higher risk of liver cirrhosis. Chamorro et al. [53] conducted a meta-analysis and also confirmed the correlation between the rs738409 G allele and ALD and alcoholic cirrhosis (ALC). Some other studies have also confirmed that PNPLA3 rs738409 was closely related to the occurrence of ALD in whites [54] and Chinese Han males [55].
Hepatic stellate cells (HSCs) are the main participants in liver fibrosis. PNPLA3 is highly expressed in HSCs and participates in HSCs-related retinol metabolism [13, 15]. When cells are stimulated by insulin, the expression of PNPLA3 in HSCs increased to promote the release of extracellular retinol [15]. PNPLA3 can promote the remodeling of extracellular matrix through this process, thereby preventing the occurrence of fibrosis [15]. The PNPLA3 I148M variant's ability to participate in the metabolism of retinol was significantly reduced, which increased the risk of fibrosis [13, 15]. In other cell experiments, it was also confirmed that PNPLA3 was essential for activating HSCs. The proliferation and migration of HSCs will be enhanced by the I148M variant, which promotes the development of fibrosis [14]. Manchiero et al. [56] tested 290 patients in the clinic hospital of the University of São Paulo and found that the PNPLA3 rs738409 GG genotype plays a major role in the development of hepatitis C (HCV) steatosis and advanced fibrosis. Yasuiet al. [57] also confirmed that this genotype was an independent risk factor for the development of steatosis, severe necrotic inflammation, and advanced liver fibrosis from 276 chronic HCV patients in Japan. In the ordinal logistic regression analysis, the more advanced stage of fibrosis was related to the PNPLA3 rs738409 genotype [58]. It is worth noting that in their study, there was no obvious relationship between the PNPLA3 rs738409 genotype and histological necrotizing inflammatory activity [58]. Ampuero et al. [59] also proposed that PNPLA3 rs738409 had a greater contribution to steatosis in patients with HCV-1 and IL28B-CT/TT genotypes. Mazo et al. [60] conducted a multi-center cross-sectional study [58]. Among them, research on NAFLD patients showed that a CG or GG at rs738409 PNPLA3 makes Brazilians more susceptible to NAFLD. In addition, NASH patients with PNPLA3 GG had more significant increased in liver enzymes and fibrosis. Kupcinskas et al. [61] in the study of Eastern Europeans also proposed that PNPLA3 rs738409 and RNF7 rs16851720 were significantly related to the occurrence of liver fibrosis and cirrhosis. In conclusion, the PNPLA3 rs738409 genotype is significantly related to the progression of ALD, NAFLD and HCV to liver fibrosis, and may be related to the severity of liver fibrosis.
PNPLA3 and hepatocellular carcinoma
HCC is one of the main causes of death from cancer worldwide [62]. HCC caused by viral hepatitis B (HBV) and HCV is the cause of death in developing countries, while HCC caused by large amounts of alcohol intake is the leading cause of death in Europeans and Americans [63-65]. A final consequence of chronic hepatitis C (CHC) or chronic hepatitis B (CHB) is sometimes development of HCC, which especially occurs to developing countries (over 80% of HCCs occurs to such regions) [65]. In 2011, Valenti et al. [58] proved that the PNPLA3 I148M gene mutation promoted the occurrence of HCC, and they also revealed that this gene mutation can promoted the development of HCV-related cirrhosis into HCC. In 2014, Liu et al. [66] showed that carriers of the PNPLA3 rs738409 C>G polymorphism had a higher risk of HCC. In the meta-analysis, the PNPLA3 rs738409 C>G polymorphism was found to be significantly associated with an increased risk of HCC, and the PNPLA3 rs738409 polymorphism may be a potential biomarker for HCC in whites [67, 68].
Valenti et al. [69] concluded the PNPLA3 148M allele was associated with a shorter history of liver cirrhosis, advanced liver cirrhosis, and lower differentiation of liver cancer in patients with ALD and NAFLD. And the homozygosity of PNPLA3 148M significantly reduced the survival rate of ALD and NAFLD patients. Nischalke et al. [70] also confirmed that the PNPLA3 148M variant was an important risk factor of the development of alcoholic cirrhosis into HCC. Further, the G/G genotype of PNPLA3 rs738409 SNP was correlated with the proceeding of HCC in patients with non-hepatitis B virus (NBNC), and showed a poor prognosis [71]. The same conclusion were obtained in the prospective study of Guyot et al. [72], and older age, higher BMI and men were more likely to develop HCC. Takeuchi et al. [71] also mentioned the correlation between BMI and HCC. Burza et al. [73] also pointed out that the PNPLA3 I148M gene mutation was a risk factor for severely obese people suffering from HCC. This indicates that the PNPLA3 148M gene mutation may not be directly related to the occurrence of tumors. From previous studies, it can be found that the mutation of PNPLA3 148M gene may not be directly related to the occurrence of tumors. PNPLA3 148M gene mutation may increase the probability of HCC by increasing steatosis, promoting fibrosis, etc. [11, 14, 20, 71, 73].
PNPLA3 and other diseases
PNPLA3 and type 2 diabetes mellitus
Diabetes is one of the common chronic diseases, which is related to decreased insulin secretion and weakened or disappeared insulin action [74]. Type 2 diabetes (T2DM) patients account for 90% to 95% of all diabetes patients, and the number is still increasing, which may be related to people's lifestyle changes. The occurrence of T2DM is not only related to environmental factors, but also related to genetic factors. The mutation of PNPLA3 as a genetic variation also has an important impact on the occurrence and development of T2DM [75, 76].
Bellan et al. suggested that T2DM patients with rs738409 mutant may be more likely to develop NAFLD [77]. Xia et al. [78] found that among participants with poor metabolism, the PNPLA3 genotype played a subtle role in weight changes and exacerbated steatosis. It is worth noting that this genotype was found in this study to reduce the risk of T2DM. Petit et al. [79] also showed that the increase in liver fat contented in T2DM patients was related to PNPLA3 rs738409, and the liver steatosis of minor G allele carriers may be more obvious. Machado et al. [80] showed that the PNPLA3 rs738409c>G polymorphism seemed to be related to the development of complications of T2DM, and the GG genotype was more closely related to the occurrence of liver fibrosis, and both CG and GG genotypes were associated with cardiovascular complications related. It is worth noting that patients with G alleles have better blood glucose control [80]. It is also mentioned in other studies that PNPLA3 SNP rs738409 has no obvious correlation with changes in ALT, AST, ALP and other indicators [81]. These results suggest that the T2DM-related complications caused by this genetic mutation may not have much to do with metabolism in the body.
In some studies, the relationship between PNPLA3 SNP rs738409 and insulin resistance was also discussed. It was found that although this gene is not only not related to insulin sensitivity, it is also not related to the levels of liver enzymes, high-sensitivity c-reactive protein and fetuin-A, suggested that PNPLA3 SNP rs738409 mutation did not cause insulin resistance [35, 82, 83]. Previous experiments have shown that PNPLA3 I148M does not seem to cause high liver fat content and insulin resistance [35, 82], and recent research showed a deeper relationship. Barata et al. [84] found through meta-analysis that reducing insulin resistance and increasing insulin levels in non-diabetic patients carrying this gene can improve liver steatosis.
From the above results, PNPLA3 I148M was found to be not directly related to the occurrence of diabetes, and the development of diabetes-related complications will be promoted by it. This association is still related to the aforementioned PNPLA3 I148M variant promoting lipid accumulation.
PNPLA3 and coronary artery disease
Cardiovascular atherosclerosis (CAD) causes a large proportion of deaths in both developing and developed countries [85]. Many factors are related to the increase in the incidence of adverse events. A large number of studies have found that in addition to showing liver pathological manifestations, NAFLD also increases the risk of cardiovascular diseases including CAD [86, 87].
According to previous research, PNPLA3 was a factor that was significantly related to susceptibility to NAFLD, and CAD was the main contributor to the death of NAFLD patients. Posadas-Sánchez et al. [88] evaluated whether the PNPLA3 gene can be used as an early diagnostic marker for early coronary artery disease (pCAD) and cardiovascular risk factors. They used the exonuclease TaqMan assay to test the genotype polymorphisms of 2572 subjects. The results showed that PNPLA3 (rs738409) polymorphism could be used as a marker to help diagnose pCAD. Machado et al. [80] performed an analysis on 303 patients with diabetes mellitus and NAFLD (118 males, mean age 59±9.5 years). For patients with diabetes and NAFLD, on the one hand, the PNPLA3 gene rs738409 C>G polymorphism increased the risk of cardiovascular disease, on the other hand, it can play a beneficial role in blood glucose control. Petta et al. [89] also found that among NAFLD patients, patients with PNPLA3 rs738409 GG genotype were more likely to develop carotid atherosclerosis. In summary, NAFLD patients with PNPLA3 rs738409 variants are at greater risk of developing CAD. Brouwers et al. [90] concluded that the NAFLD susceptibility gene (PNPLA3 I148M) itself did not cause CAD, and the relationship between PNPLA3 and CAD was mainly through plasma lipids. In short, the PNPLA3 I148M variant can promote the development of NAFLD, thereby increasing the risk of CAD in patients.
PNPLA3 and kidney disease
Chronic kidney disease (CKD), as usual chronic disease, affects approximately 8-16% of adults worldwide, and the prevalence increases sharply with age [91]. At present, with the accumulation of research on PNPLA3, NAFLD is more closely related to kidney function, and the growth of chronic kidney disease (CKD) may also be related to PNPLA3 [92, 93].
Sun et al. [94] attempted to explore which biomarkers related to renal tubular injury (RTI) and glomerular function changes were affected by the PNPLA3 genotype. Their patients were those who had NAFLD with either normal alanine aminotransferase levels (nALT) or abnormal alanine aminotransferase levels (abnALT). They studied 217 patients with histologically confirmed NAFLD, 75 of whom had sustained nALT levels (three months below the normal upper limit). The results showed that the nALT patient groups had higher albuminuria than the abnALT group and had a higher prevalence of CKD. They concluded that adult patients with the PNPLA3 rs738409G allele, NAFLD, and persistent nALT are more likely to develop early glomerular and tubular damage. The molecular mechanisms remain unknown. Mantovani et al. [95] studied 101 postmenopausal white women with T2DM and found that the rs738409 G/G genotype of the PNPLA3 gene was connected with a lower level of e-GFRCKD-EPI and a higher risk of CKD. There was no obvious correlation with NAFLD and other risk factors, which has also been confirmed in other studies [96]. In addition, studies have shown that this genotype was related to the decline in renal function of the elderly in Japan [97].
In addition, Targher et al. [98] performed a similar study on Caucasian children with biopsy-proven NAFLD, and showed for the first time that decreased e-GFR and increased 24-hour urinary protein excretion were strongly associated with the G allele of rs738409. Costanzo et al. [99] also obtained the same result through a study of 230 obese children and adolescents, that is, the PNPLA3 rs738409 variant increases the risk of impaired renal function in children with NAFLD. Regarding the cause of renal damage, research speculated that it may be related to retinol metabolism, lipid metabolism and podocyte damage [96]. NAFLD is considered to be an independent risk factor for CKD. Based on the above research, it is speculated that the PNPLA3 I148M variant promotes the development of CKD is still related to the susceptibility of NAFLD.
Further challenges
The fact that PNPLA3 is a factor that is significantly related to the susceptibility of NAFLD is now well established [100-102]. Thus, some researchers think they can study precision medicine in NAFLD with PNPLA3 as a therapeutic target [103]. Although the PNPLA3 I148M in the pathology of fatty liver disease is well understood, further research is clearly needed to further expand the molecular mechanisms associated with these new findings and to further validate these results. PNPLA3 I148M is closely related to liver disease and its complications, and according to the current research, it is mainly related to the accumulation of lipids in the liver. But its specific mechanism also needs further research to clarify.
The recent research in murine models shows that antisense oligonucleotide therapy for PNPLA3 can delay the progress of NAFLD and improve liver fibrosis. Importantly, it can also inhibit the expression of PNPLA3 148M [104]. Although this experiment was conducted in mice, in future, it may also open to a precision medicine approach from NASH.
Studies of inhibitors or agonists of PNPLA3 are relatively few, and even now, people understand that there is a strong association between PNPLA3 and those diseases, thus research on inhibitors or agonists of PNPLA3 and its variant could be further investigated in the next step.
Abbreviations
PNPLA3: patatin-like phospholipase domain-containing 3 gene; TG: triglyceride; PL: phospholipid; BMI: body mass index; LPAAT: lysophosphatidic acid acyltransferase activity; NAFLD: non-alcoholic fatty liver disease; NASH: non-alcoholic steatohepatitis; ALT: alanine transaminase; AST: aspartate transaminase; CGI-58: comparative gene identification-58; ATGL: adipose triglyceride lipase; ALD: alcoholic liver injury; CHC: chronic hepatitis C; CHB: chronic hepatitis B; HCC: hepatocellular carcinoma; SNPs: single pairs of nucleotide polymorphisms; HCV: hepatitis C virus; CBM: Chinese Biomedical Database; T2DM: type 2 diabetes mellitus; CAD: coronary artery disease; CKD: chronic kidney disease; RTI: renal tubular injury; nALT: normal alanine aminotransaminase levels; abnALT: abnormal alanine aminotransaminase levels; e-GFR: estimated Glomerular Filtration Rate.
Acknowledgements
Authors' contributions
T.W. contributed to design the research and review of the final version of the manuscript. H.J.X. contributed to conduct/data collection, analysis of the research data and wrote the manuscript. Z.C.W and J.M.W. contributed to data collection. All authors contributed to and approved the final manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (81873076), Shanghai Talents development fund Project (2017090) and Hundred Talents Program of Shanghai University of Traditional Chinese Medicine.
Availability of data and materials
All data presented herein is available from the referenced sources.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Asrani SK, Devarbhavi H, Eaton J, Kamath PS. Burden of liver diseases in the world. Journal of hepatology. 2019;70:151-71
2. Kienesberger PC, Oberer M, Lass A, Zechner R. Mammalian patatin domain containing proteins: a family with diverse lipolytic activities involved in multiple biological functions. Journal of lipid research. 2009;50(Suppl):S63-S8
3. Strickland JA, Orr GL, Walsh TA. Inhibition of Diabrotica Larval Growth by Patatin, the Lipid Acyl Hydrolase from Potato Tubers. Plant Physiol. 1995;109:667-74
4. Rydel TJ, Williams JM, Krieger E, Moshiri F, Stallings WC, Brown SM. et al. The crystal structure, mutagenesis, and activity studies reveal that patatin is a lipid acyl hydrolase with a Ser-Asp catalytic dyad. Biochemistry. 2003;42:6696-708
5. Baulande S, Lasnier F, Lucas M, Pairault J. Adiponutrin, a transmembrane protein corresponding to a novel dietary- and obesity-linked mRNA specifically expressed in the adipose lineage. The Journal of biological chemistry. 2001;276:33336-44
6. Tortora R, Rispo A, Alisi A, Imperatore N, Crudele A, Ferretti F. et al. PNPLA3 rs738409 Polymorphism Predicts Development and Severity of Hepatic Steatosis but Not Metabolic Syndrome in Celiac Disease. Nutrients. 2018 10
7. Sookoian S, Pirola CJ. PNPLA3, the triacylglycerol synthesis/hydrolysis/storage dilemma, and nonalcoholic fatty liver disease. World J Gastroenterol. 2012;18:6018-26
8. Wilson PA, Gardner SD, Lambie NM, Commans SA, Crowther DJ. Characterization of the human patatin-like phospholipase family. J Lipid Res. 2006;47:1940-9
9. Basu Ray S. PNPLA3-I148M: a problem of plenty in non-alcoholic fatty liver disease. Adipocyte. 2019;8:201-8
10. Sahlman P, Nissinen M, Puukka P, Jula A, Salomaa V, Männistö S. et al. Genetic and lifestyle risk factors for advanced liver disease among men and women. J Gastroenterol Hepatol. 2020;35:291-8
11. He S, McPhaul C, Li JZ, Garuti R, Kinch L, Grishin NV. et al. A sequence variation (I148M) in PNPLA3 associated with nonalcoholic fatty liver disease disrupts triglyceride hydrolysis. J Biol Chem. 2010;285:6706-15
12. Pingitore P, Pirazzi C, Mancina RM, Motta BM, Indiveri C, Pujia A. et al. Recombinant PNPLA3 protein shows triglyceride hydrolase activity and its I148M mutation results in loss of function. Biochim Biophys Acta. 2014;1841:574-80
13. Pingitore P, Dongiovanni P, Motta BM, Meroni M, Lepore SM, Mancina RM. et al. PNPLA3 overexpression results in reduction of proteins predisposing to fibrosis. Hum Mol Genet. 2016;25:5212-22
14. Bruschi FV, Claudel T, Tardelli M, Caligiuri A, Stulnig TM, Marra F. et al. The PNPLA3 I148M variant modulates the fibrogenic phenotype of human hepatic stellate cells. Hepatology. 2017;65:1875-90
15. Pirazzi C, Valenti L, Motta BM, Pingitore P, Hedfalk K, Mancina RM. et al. PNPLA3 has retinyl-palmitate lipase activity in human hepatic stellate cells. Hum Mol Genet. 2014;23:4077-85
16. Gunn PJ, Green CJ, Pramfalk C, Hodson L. In vitro cellular models of human hepatic fatty acid metabolism: differences between Huh7 and HepG2 cell lines in human and fetal bovine culturing serum. Physiol Rep. 2017 5
17. Huang Y, He S, Li JZ, Seo Y-K, Osborne TF, Cohen JC. et al. A feed-forward loop amplifies nutritional regulation of PNPLA3. Proc Natl Acad Sci USA. 2010;107:7892-7
18. Chamoun Z, Vacca F, Parton RG, Gruenberg J. PNPLA3/adiponutrin functions in lipid droplet formation. Biol Cell. 2013;105:219-33
19. Ruhanen H, Perttilä J, Hölttä-Vuori M, Zhou Y, Yki-Järvinen H, Ikonen E. et al. PNPLA3 mediates hepatocyte triacylglycerol remodeling. Journal of lipid research. 2014;55:739-46
20. Huang Y, Cohen JC, Hobbs HH. Expression and characterization of a PNPLA3 protein isoform (I148M) associated with nonalcoholic fatty liver disease. J Biol Chem. 2011;286:37085-93
21. Mitsche MA, Hobbs HH, Cohen JC. Patatin-like phospholipase domain-containing protein 3 promotes transfer of essential fatty acids from triglycerides to phospholipids in hepatic lipid droplets. J Biol Chem. 2018;293:6958-68
22. Chen L-Z, Xin Y-N, Geng N, Jiang M, Zhang D-D, Xuan S-Y. PNPLA3 I148M variant in nonalcoholic fatty liver disease: demographic and ethnic characteristics and the role of the variant in nonalcoholic fatty liver fibrosis. World journal of gastroenterology. 2015;21:794-802
23. Moldes M, Beauregard G, Faraj M, Peretti N, Ducluzeau P-H, Laville M. et al. Adiponutrin gene is regulated by insulin and glucose in human adipose tissue. Eur J Endocrinol. 2006;155:461-8
24. Kotronen A, Johansson LE, Johansson LM, Roos C, Westerbacka J, Hamsten A. et al. A common variant in PNPLA3, which encodes adiponutrin, is associated with liver fat content in humans. Diabetologia. 2009;52:1056-60
25. Basantani MK, Sitnick MT, Cai L, Brenner DS, Gardner NP, Li JZ. et al. Pnpla3/Adiponutrin deficiency in mice does not contribute to fatty liver disease or metabolic syndrome. Journal of lipid research. 2011;52:318-29
26. Chen W, Chang B, Li L, Chan L. Patatin-like phospholipase domain-containing 3/adiponutrin deficiency in mice is not associated with fatty liver disease. Hepatology. 2010;52:1134-42
27. Kumari M, Schoiswohl G, Chitraju C, Paar M, Cornaciu I, Rangrez AY. et al. Adiponutrin functions as a nutritionally regulated lysophosphatidic acid acyltransferase. Cell Metab. 2012;15:691-702
28. BasuRay S, Smagris E, Cohen JC, Hobbs HH. The PNPLA3 variant associated with fatty liver disease (I148M) accumulates on lipid droplets by evading ubiquitylation. Hepatology. 2017;66:1111-24
29. Saeed A, Dullaart RPF, Schreuder T, Blokzijl H, Faber KN. Disturbed Vitamin A Metabolism in Non-Alcoholic Fatty Liver Disease (NAFLD). Nutrients. 2017 10
30. Shajari S, Saeed A, Smith-Cortinez NF, Heegsma J, Sydor S, Faber KN. Hormone-sensitive lipase is a retinyl ester hydrolase in human and rat quiescent hepatic stellate cells. Biochim Biophys Acta Mol Cell Biol Lipids. 2019;1864:1258-67
31. Hamaguchi M, Kojima T, Takeda N, Nakagawa T, Taniguchi H, Fujii K. et al. The metabolic syndrome as a predictor of nonalcoholic fatty liver disease. Ann Intern Med. 2005;143:722-8
32. Cohen JC, Horton JD, Hobbs HH. Human fatty liver disease: old questions and new insights. Science. 2011;332:1519-23
33. Ochi T, Munekage K, Ono M, Higuchi T, Tsuda M, Hayashi Y. et al. Patatin-like phospholipase domain-containing protein 3 is involved in hepatic fatty acid and triglyceride metabolism through X-box binding protein 1 and modulation of endoplasmic reticulum stress in mice. Hepatol Res. 2016;46:584-92
34. Hoekstra M, Li Z, Kruijt JK, Van Eck M, Van Berkel TJC, Kuiper J. The expression level of non-alcoholic fatty liver disease-related gene PNPLA3 in hepatocytes is highly influenced by hepatic lipid status. Journal of hepatology. 2010;52:244-51
35. Romeo S, Kozlitina J, Xing C, Pertsemlidis A, Cox D, Pennacchio LA. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nature genetics. 2008;40:1461-5
36. Hotta K, Yoneda M, Hyogo H, Ochi H, Mizusawa S, Ueno T. et al. Association of the rs738409 polymorphism in PNPLA3 with liver damage and the development of nonalcoholic fatty liver disease. BMC Med Genet. 2010;11:172
37. Jain V, Kumar A, Ahmad N, Jana M, Kalaivani M, Kumar B. et al. Genetic polymorphisms associated with obesity and non-alcoholic fatty liver disease in Asian Indian adolescents. J Pediatr Endocrinol Metab. 2019;32:749-58
38. Rotman Y, Koh C, Zmuda JM, Kleiner DE, Liang TJ. The association of genetic variability in patatin-like phospholipase domain-containing protein 3 (PNPLA3) with histological severity of nonalcoholic fatty liver disease. Hepatology. 2010;52:894-903
39. Valenti L, Alisi A, Galmozzi E, Bartuli A, Del Menico B, Alterio A. et al. I148M patatin-like phospholipase domain-containing 3 gene variant and severity of pediatric nonalcoholic fatty liver disease. Hepatology. 2010;52:1274-80
40. Zain SM, Mohamed R, Mahadeva S, Cheah PL, Rampal S, Basu RC. et al. A multi-ethnic study of a PNPLA3 gene variant and its association with disease severity in non-alcoholic fatty liver disease. Hum Genet. 2012;131:1145-52
41. Sookoian S, Pirola CJ. Meta-analysis of the influence of I148M variant of patatin-like phospholipase domain containing 3 gene (PNPLA3) on the susceptibility and histological severity of nonalcoholic fatty liver disease. Hepatology. 2011;53:1883-94
42. Unalp-Arida A, Ruhl CE. Patatin-Like Phospholipase Domain-Containing Protein 3 I148M and Liver Fat and Fibrosis Scores Predict Liver Disease Mortality in the U.S. Population. Hepatology. 2020;71:820-34
43. Wang J-Z, Cao H-X, Chen J-N, Pan Q. PNPLA3 rs738409 underlies treatment response in nonalcoholic fatty liver disease. World J Clin Cases. 2018;6:167-75
44. Li JZ, Huang Y, Karaman R, Ivanova PT, Brown HA, Roddy T. et al. Chronic overexpression of PNPLA3I148M in mouse liver causes hepatic steatosis. J Clin Invest. 2012;122:4130-44
45. Liu W, Anstee QM, Wang X, Gawrieh S, Gamazon ER, Athinarayanan S. et al. Transcriptional regulation of PNPLA3 and its impact on susceptibility to nonalcoholic fatty liver Disease (NAFLD) in humans. Aging (Albany NY). 2016;9:26-40
46. Smagris E, BasuRay S, Li J, Huang Y, Lai KM, Gromada J. et al. Pnpla3I148M knockin mice accumulate PNPLA3 on lipid droplets and develop hepatic steatosis. Hepatology. 2015;61:108-18
47. BasuRay S, Wang Y, Smagris E, Cohen JC, Hobbs HH. Accumulation of PNPLA3 on lipid droplets is the basis of associated hepatic steatosis. Proc Natl Acad Sci USA. 2019;116:9521-6
48. Shimamura T, Fujisawa T, Husain SR, Kioi M, Nakajima A, Puri RK. Novel role of IL-13 in fibrosis induced by nonalcoholic steatohepatitis and its amelioration by IL-13R-directed cytotoxin in a rat model. J Immunol. 2008;181:4656-65
49. Wynn TA, Ramalingam TR. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat Med. 2012;18:1028-40
50. Seitz HK, Bataller R, Cortez-Pinto H, Gao B, Gual A, Lackner C. et al. Alcoholic liver disease. Nat Rev Dis Primers. 2018;4:16
51. Kolla BP, Schneekloth TD, Biernacka J, Shah V, Lazaridis KN, Geske J. et al. PNPLA3 Association with Alcoholic Liver Disease in a Cohort of Heavy Drinkers. Alcohol Alcohol. 2018;53:357-60
52. Trepo E, Gustot T, Degre D, Lemmers A, Verset L, Demetter P. et al. Common polymorphism in the PNPLA3/adiponutrin gene confers higher risk of cirrhosis and liver damage in alcoholic liver disease. J Hepatol. 2011;55:906-12
53. Chamorro AJ, Torres JL, Miron-Canelo JA, Gonzalez-Sarmiento R, Laso FJ, Marcos M. Systematic review with meta-analysis: the I148M variant of patatin-like phospholipase domain-containing 3 gene (PNPLA3) is significantly associated with alcoholic liver cirrhosis. Aliment Pharmacol Ther. 2014;40:571-81
54. Stickel F, Buch S, Lau K, Meyer zu Schwabedissen H, Berg T, Ridinger M. et al. Genetic variation in the PNPLA3 gene is associated with alcoholic liver injury in caucasians. Hepatology. 2011;53:86-95
55. Zhang Y, Guo T, Yang F, Mao Y, Li L, Liu C. et al. Single-nucleotide rs738409 polymorphisms in the PNPLA3 gene are strongly associated with alcoholic liver disease in Han Chinese males. Hepatol Int. 2018;12:429-37
56. Manchiero C, Nunes A, Magri MC, Dantas BP, Mazza CC, Barone AA. et al. The rs738409 polymorphism of the PNPLA3 gene is associated with hepatic steatosis and fibrosis in Brazilian patients with chronic hepatitis C. BMC Infect Dis. 2017;17:780
57. Yasui K, Kawaguchi T, Shima T, Mitsuyoshi H, Seki K, Sendo R. et al. Effect of PNPLA3 rs738409 variant (I148 M) on hepatic steatosis, necroinflammation, and fibrosis in Japanese patients with chronic hepatitis C. J Gastroenterol. 2015;50:887-93
58. Valenti L, Rumi M, Galmozzi E, Aghemo A, Del Menico B, De Nicola S. et al. Patatin-like phospholipase domain-containing 3 I148M polymorphism, steatosis, and liver damage in chronic hepatitis C. Hepatology. 2011;53:791-9
59. Ampuero J, Campo JAD, Rojas L, García-Lozano JR, Solá R, Andrade R. et al. PNPLA3 rs738409 causes steatosis according to viral & IL28B genotypes in hepatitis C. Annals of Hepatology. 2014;13:356-63
60. Mazo DF, Malta FM, Stefano JT, Salles APM, Gomes-Gouvea MS, Nastri ACS. et al. Validation of PNPLA3 polymorphisms as risk factor for NAFLD and liver fibrosis in an admixed population. Annals of hepatology. 2019;18:466-71
61. Kupcinskas J, Valantiene I, Varkalaite G, Steponaitiene R, Skieceviciene J, Sumskiene J. et al. PNPLA3 and RNF7 Gene Variants are Associated with the Risk of Developing Liver Fibrosis and Cirrhosis in an Eastern European Population. J Gastrointestin Liver Dis. 2017;26:37-43
62. Schutte K, Bornschein J, Malfertheiner P. Hepatocellular carcinoma-epidemiological trends and risk factors. Dig Dis. 2009;27:80-92
63. Altekruse SF, McGlynn KA, Reichman ME. Hepatocellular carcinoma incidence, mortality, and survival trends in the United States from 1975 to 2005. J Clin Oncol. 2009;27:1485-91
64. Espey DK, Wu XC, Swan J, Wiggins C, Jim MA, Ward E. et al. Annual report to the nation on the status of cancer, 1975-2004, featuring cancer in American Indians and Alaska Natives. Cancer. 2007;110:2119-52
65. Yang JD, Roberts LR. Hepatocellular carcinoma: A global view. Nat Rev Gastroenterol Hepatol. 2010;7:448-58
66. Liu YL, Patman GL, Leathart JBS, Piguet AC, Burt AD, Dufour JF. et al. Carriage of the PNPLA3 rs738409 C >G polymorphism confers an increased risk of non-alcoholic fatty liver disease associated hepatocellular carcinoma. Journal of hepatology. 2014;61:75-81
67. Li J-F, Zheng E-Q, Xie M. Association between rs738409 polymorphism in patatin-like phospholipase domain-containing protein 3 (PNPLA3) gene and hepatocellular carcinoma susceptibility: Evidence from case-control studies. Gene. 2019;685:143-8
68. Huang Z, Guo X, Zhang G, Liang L, Nong B. Correlation between rs738409 polymorphism and hepatocellular carcinoma: a meta-analysis of 10,330 subjects. Int J Biol Markers. 2019;34:117-22
69. Valenti L, Motta BM, Soardo G, Iavarone M, Donati B, Sangiovanni A. et al. PNPLA3 I148M polymorphism, clinical presentation, and survival in patients with hepatocellular carcinoma. PLoS One. 2013;8:e75982
70. Nischalke HD, Berger C, Luda C, Berg T, Muller T, Grunhage F. et al. The PNPLA3 rs738409 148M/M genotype is a risk factor for liver cancer in alcoholic cirrhosis but shows no or weak association in hepatitis C cirrhosis. PLoS One. 2011;6:e27087
71. Takeuchi Y, Ikeda F, Moritou Y, Hagihara H, Yasunaka T, Kuwaki K. et al. The impact of patatin-like phospholipase domain-containing protein 3 polymorphism on hepatocellular carcinoma prognosis. J Gastroenterol. 2013;48:405-12
72. Guyot E, Sutton A, Rufat P, Laguillier C, Mansouri A, Moreau R. et al. PNPLA3 rs738409, hepatocellular carcinoma occurrence and risk model prediction in patients with cirrhosis. J Hepatol. 2013;58:312-8
73. Burza MA, Pirazzi C, Maglio C, Sjoholm K, Mancina RM, Svensson PA. et al. PNPLA3 I148M (rs738409) genetic variant is associated with hepatocellular carcinoma in obese individuals. Dig Liver Dis. 2012;44:1037-41
74. Chen L, Magliano DJ, Zimmet PZ. The worldwide epidemiology of type 2 diabetes mellitus-present and future perspectives. Nat Rev Endocrinol. 2011;8:228-36
75. Wu Y, Ding Y, Tanaka Y, Zhang W. Risk factors contributing to type 2 diabetes and recent advances in the treatment and prevention. Int J Med Sci. 2014;11:1185-200
76. Zhao Y, Jiang Z, Guo C. New hope for type 2 diabetics: targeting insulin resistance through the immune modulation of stem cells. Autoimmun Rev. 2011;11:137-42
77. Bellan M, Colletta C, Barbaglia MN, Salmi L, Clerici R, Mallela VR. et al. Severity of Nonalcoholic Fatty Liver Disease in Type 2 Diabetes Mellitus: Relationship between Nongenetic Factors and PNPLA3/HSD17B13 Polymorphisms. Diabetes Metab J. 2019;43:700-10
78. Xia M-F, Lin H-D, Chen L-Y, Wu L, Ma H, Li Q. et al. The PNPLA3 rs738409 C>G variant interacts with changes in body weight over time to aggravate liver steatosis, but reduces the risk of incident type 2 diabetes. Diabetologia. 2019;62:644-54
79. Petit JM, Guiu B, Masson D, Duvillard L, Jooste V, Buffier P. et al. Specifically PNPLA3-mediated accumulation of liver fat in obese patients with type 2 diabetes. J Clin Endocrinol Metab. 2010;95:E430-6
80. Machado CM, Leite NC, Franca PH, Cardoso CR, Salles GF, Villela-Nogueira CA. PNPLA3 gene polymorphism in Brazilian patients with type 2 diabetes: A prognostic marker beyond liver disease? Nutr Metab Cardiovasc Dis. 2019;29:965-71
81. Cox AJ, Wing MR, Carr JJ, Hightower RC, Smith SC, Xu J. et al. Association of PNPLA3 SNP rs738409 with liver density in African Americans with type 2 diabetes mellitus. Diabetes Metab. 2011;37:452-5
82. Kantartzis K, Peter A, Machicao F, Machann J, Wagner S, Königsrainer I. et al. Dissociation between fatty liver and insulin resistance in humans carrying a variant of the patatin-like phospholipase 3 gene. Diabetes. 2009;58:2616-23
83. Lallukka S, Yki-Järvinen H. Non-alcoholic fatty liver disease and risk of type 2 diabetes. Best Pract Res Clin Endocrinol Metab. 2016;30:385-95
84. Barata L, Feitosa MF, Bielak LF, Halligan B, Baldridge AS, Guo X. et al. Insulin Resistance Exacerbates Genetic Predisposition to Nonalcoholic Fatty Liver Disease in Individuals Without Diabetes. Hepatol Commun. 2019;3:894-907
85. Go AS, Mozaffarian D, Roger VL, Benjamin EJ, Berry JD, Blaha MJ. et al. Heart disease and stroke statistics-2014 update: a report from the American Heart Association. Circulation. 2014;129:e28-e292
86. Targher G, Marra F, Marchesini G. Increased risk of cardiovascular disease in non-alcoholic fatty liver disease: causal effect or epiphenomenon? Diabetologia. 2008;51:1947-53
87. Marchesini G, Brizi M, Bianchi G, Tomassetti S, Bugianesi E, Lenzi M. et al. Nonalcoholic fatty liver disease: a feature of the metabolic syndrome. Diabetes. 2001;50:1844-50
88. Posadas-Sánchez R, López-Uribe ÁR, Posadas-Romero C, Pérez-Hernández N, Rodríguez-Pérez JM, Ocampo-Arcos WA. et al. Association of the I148M/PNPLA3 (rs738409) polymorphism with premature coronary artery disease, fatty liver, and insulin resistance in type 2 diabetic patients and healthy controls. The GEA study. Immunobiology. 2017;222:960-6
89. Petta S, Valenti L, Marchesini G, Di Marco V, Licata A, Cammà C. et al. PNPLA3 GG genotype and carotid atherosclerosis in patients with non-alcoholic fatty liver disease. PloS one. 2013;8:e74089
90. Brouwers MCGJ, Simons N, Stehouwer CDA, Koek GH, Schaper NC, Isaacs A. Relationship Between Nonalcoholic Fatty Liver Disease Susceptibility Genes and Coronary Artery Disease. Hepatol Commun. 2019;3:587-96
91. Coresh J, Selvin E, Stevens LA, Manzi J, Kusek JW, Eggers P. et al. Prevalence of chronic kidney disease in the United States. JAMA. 2007;298:2038-47
92. Targher G, Byrne CD. Non-alcoholic fatty liver disease: an emerging driving force in chronic kidney disease. Nat Rev Nephrol. 2017;13:297-310
93. Mantovani A, Zaza G, Byrne CD, Lonardo A, Zoppini G, Bonora E. et al. Nonalcoholic fatty liver disease increases risk of incident chronic kidney disease: A systematic review and meta-analysis. Metab Clin Exp. 2018;79:64-76
94. Sun D-Q, Zheng KI, Xu G, Ma H-L, Zhang H-Y, Pan X-Y. et al. PNPLA3 rs738409 is associated with renal glomerular and tubular injury in NAFLD patients with persistently normal ALT levels. Liver international: official journal of the International Association for the Study of the Liver. 2020;40:107-19
95. Mantovani A, Zusi C, Sani E, Colecchia A, Lippi G, Zaza GL. et al. Association between PNPLA3rs738409 polymorphism decreased kidney function in postmenopausal type 2 diabetic women with or without non-alcoholic fatty liver disease. Diabetes Metab. 2019;45:480-7
96. Mantovani A, Taliento A, Zusi C, Baselli G, Prati D, Granata S. et al. PNPLA3 I148M gene variant and chronic kidney disease in type 2 diabetic patients with NAFLD: Clinical and experimental findings. Liver Int. 2020;40:1130-41
97. Oniki K, Saruwatari J, Izuka T, Kajiwara A, Morita K, Sakata M. et al. Influence of the PNPLA3 rs738409 Polymorphism on Non-Alcoholic Fatty Liver Disease and Renal Function among Normal Weight Subjects. PloS one. 2015;10:e0132640
98. Targher G, Mantovani A, Alisi A, Mosca A, Panera N, Byrne CD. et al. Relationship Between PNPLA3 rs738409 Polymorphism and Decreased Kidney Function in Children With NAFLD. Hepatology. 2019;70:142-53
99. Di Costanzo A, Pacifico L, D'Erasmo L, Polito L, Martino MD, Perla FM. et al. Nonalcoholic Fatty Liver Disease (NAFLD), But not Its Susceptibility Gene Variants, Influences the Decrease of Kidney Function in Overweight/Obese Children. International journal of molecular sciences. 2019 20
100. Perez-Matos MC, Sandhu B, Bonder A, Jiang ZG. Lipoprotein metabolism in liver diseases. Curr Opin Lipidol. 2019;30:30-6
101. Trepo E, Romeo S, Zucman-Rossi J, Nahon P. PNPLA3 gene in liver diseases. J Hepatol. 2016;65:399-412
102. Severson TJ, Besur S, Bonkovsky HL. Genetic factors that affect nonalcoholic fatty liver disease: A systematic clinical review. World journal of gastroenterology. 2016;22:6742-56
103. Colombo M, Pelusi S. Towards Precision Medicine in Nonalcoholic Fatty Liver Disease with PNPLA3 as a Therapeutic Target. Gastroenterology. 2019;157:1156-7
104. Lindén D, Ahnmark A, Pingitore P, Ciociola E, Ahlstedt I, Andréasson A-C. et al. Pnpla3 silencing with antisense oligonucleotides ameliorates nonalcoholic steatohepatitis and fibrosis in Pnpla3 I148M knock-in mice. Mol Metab. 2019;22:49-61
Author contact
Corresponding authors: Tao Wu, PhD, Professor, Institute of Interdisciplinary Integrative Medicine Research, Shanghai University of Traditional Chinese Medicine, Cailun Road 1200, Shanghai, China. Email: tw827edu.cn; or wutao001827com; Telephone: +86-21-51322748; Fax: +86-21-51322748.

 Global reach, higher impact
Global reach, higher impact