Impact Factor
ISSN: 1837-9664
J Cancer 2021; 12(12):3530-3538. doi:10.7150/jca.51493 This issue Cite
Research Paper
Development and validation of nomogram with tumor microenvironment-related genes and clinical factors for predicting overall survival of endometrial cancer
Department of Gynecology and Obstetrics, Peking Union Medical College Hospital (PUMCH), Chinese Academy of Medical Sciences (CAMS) & Peking Union Medical College, National Clinical Research Center for Obstetric & Gynecologic Diseases. Beijing, P.R, China
Abstract
Background: Tumor microenvironment (TME) has attracted lots of attention with its important role in the tumor development. This study aimed to explore TME- related genes of prognostic value in patients with endometrial cancer (EC) and establish a prediction model for EC.
Methods: The RNA-Seq data and clinicopathological characteristics of 521 subjects were obtained from The Cancer Genome Atlas (TCGA) database. Differentially expressed genes (DEGs) were identified based on the immune and stromal scores, which were calculated by the ESTIMATE algorithm. Hub genes were initially screened using the Cytoscape and further selected through Cox regression. Gene correlation analysis was performed in TIMER database. A nomogram was constructed integrating prognosis-related hub genes and clinical factors and validated in the validation group. Risk stratification was performed based on the nomogram.
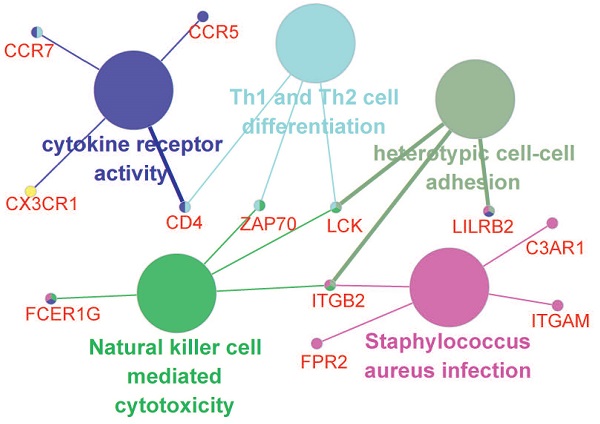
Results: Three TME-related hub genes (CCR5, FCER1G, and ITGB2) were found with significant prognostic value for EC patients. The expression of CCR5, FCER1G, and ITGB2 were significantly correlated with various immune cells infiltration. Based on the Cox regression, a nomogram was constructed by integrating five predictors (stage, grade, immune score, expression of FCER1G, and ITGB2), with a C-index of 0.765. Discrimination of the model was confirmed in the validation group (C-index: 0.716). The calibration curves for the 3- and 5- year survival indicated good calibration. Patients in high- and low- risk groups presented significantly different survival outcomes (P<0.001) in both discovery and validation group.
Conclusion: TME-related hub genes of prognostic value identified in our study may provide references for the mechanisms underlying EC development and the immunotherapy for EC. The prediction model may help assess the prognosis of EC patients.
Keywords: endometrial cancer, tumor microenvironment, nomogram, prognosis, immune infiltration.

 Global reach, higher impact
Global reach, higher impact