Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(6):1604-1615. doi:10.7150/jca.52183 This issue Cite
Research Paper
Machine learning classifiers for predicting 3-year progression-free survival and overall survival in patients with gliomas after surgery
1. Department of Radiology, The First Affiliated Hospital of Jinan University, Guangzhou, Guangdong, China.
2. Department of MRI, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, China.
3. Big Data Decision Institute, Jinan University, Guangzhou, Guangdong, China.
4. School of management, Jinan University. Department of Catheterization Lab, Guangdong Cardiovascular Institute, Guangdong, Provincial Key Laboratory of South China
5. Structural Heart Disease, Guangdong Provincial; People's Hospital/Guangdong Academy of Medical Sciences, Guangzhou, Guangdong, P.R. China.
6. Department of Neurosurgery, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, China.
7. Department of Pathology, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, China.
*These authors contributed equally to this work.
Abstract
Background: To develop machine-learning based models to predict the progression-free survival (PFS) and overall survival (OS) in patients with gliomas and explore the effect of different feature selection methods on the prediction.
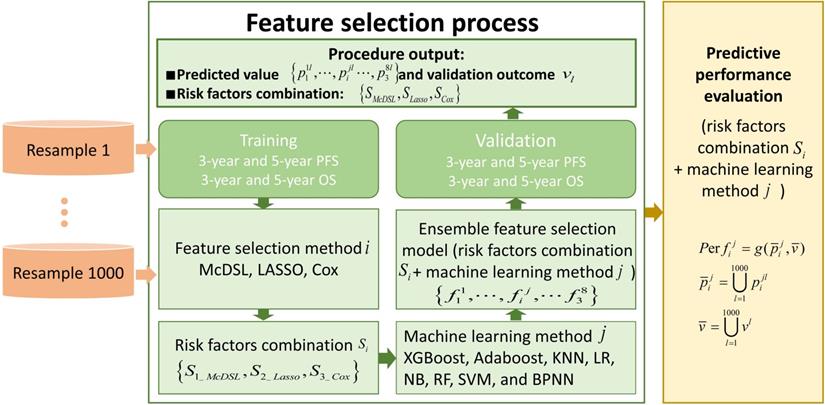
Methods: We included 505 patients (training cohort, n = 354; validation cohort, n = 151) with gliomas between January 1, 2011 and December 31, 2016. The clinical, neuroimaging, and molecular genetic data of patients were retrospectively collected. The multi-causes discovering with structure learning (McDSL) algorithm, least absolute shrinkage and selection operator regression (LASSO), and Cox proportional hazards regression model were employed to discover the predictors for 3-year PFS and OS, respectively. Eight machine learning classifiers with 5-fold cross-validation were developed to predict 3-year PFS and OS. The area under the curve (AUC) was used to evaluate the prognostic performance of classifiers.
Results: McDSL identified four causal factors (tumor location, WHO grade, histologic type, and molecular genetic group) for 3-year PFS and OS, whereas LASSO and Cox identified wide-range number of factors associated with 3-year PFS and OS. The performance of each machine learning classifier based on McDSL, LASSO, and Cox was not significantly different. Logistic regression yielded the optimal performance in predicting 3-year PFS based on the McDSL (AUC, 0.872, 95% confidence interval [CI]: 0.828-0.916) and 3-year OS based on the LASSO (AUC, 0.901, 95% CI: 0.861-0.940).
Conclusions: McDSL is more reproducible than LASSO and Cox model in the feature selection process. Logistic regression model may have the highest performance in predicting 3-year PFS and OS of gliomas.
Keywords: gliomas, molecular biomarkers, machine learning, progression-free survival, overall survival

 Global reach, higher impact
Global reach, higher impact