Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(4):1011-1022. doi:10.7150/jca.48310 This issue Cite
Research Paper
EHMT2 inhibitor BIX-01294 induces endoplasmic reticulum stress mediated apoptosis and autophagy in diffuse large B-cell lymphoma cells
1. Blood Diseases Institute, Xuzhou Medical University, Xuzhou, Jiangsu, China.
2. Department of Hematology, the Affiliated Hospital of Xuzhou Medical University, Xuzhou, Jiangsu, China.
3. Key Laboratory of Bone Marrow Stem Cell, Xuzhou, Jiangsu, China.
4. Department of Hematology, Luoyang Central Hospital Affiliated to Zhengzhou University, Luoyang, Henan, China.
*Equal contributors to this work.
Received 2020-5-17; Accepted 2020-11-15; Published 2021-1-1
Abstract
Despite advancement in the treatment of diffuse large B-cell lymphoma (DLBCL), many patients tend to relapse or become refractory after initial therapy. Therefore, it is essential to identify novel therapeutic targets and drugs, understand the molecular pathogenesis mechanism of DLBCL, and find ways to prevent and treat relapsed or refractory DLBCL. BIX-01294 is a small molecule compound that specifically inhibits EHMT2 activity. In this study, we demonstrate that BIX-01294 triggered the inhibition of human DLBCL cell proliferation, lead to G1 phase arrest via increasing P21 level and reducing cyclin E level. BIX-01294 also induced apoptosis via endogenous and exogenous apoptotic pathways. Moreover, BIX-01294 triggered autophagy and activated ER stress in human DLBCL cells. Furthermore, we showed that both key components of ER stress, ATF3, and ATF4, are required for BIX-01294-induced apoptosis and autophagy. Hence, this study provides new evidence that EHMT2 may be a new therapeutic target, and BIX-01294 may be a potential therapeutic drug for treating DLBCL.
Keywords: apoptosis, autophagy, BIX-01294, diffuse large B cell lymphoma, endoplasmic reticulum stress
Introduction
Diffuse large B-cell lymphoma (DLBCL) is the most common type of lymphoma and represents 30-40% of non-Hodgkin lymphoma (NHL) cases worldwide. At present, the standard therapy R-CHOP (rituximab, cyclophosphamide, doxorubicin, vincristine and prednisone) regimen may lead to complete remission. However, 30-40% of patients are refractory to therapy or relapse after treatment indicating standard cytotoxic therapy has its limitations [1-4]. Factors that influence the cure of DLBCL include patient age, molecular subtypes, gene expression, and others [5]. Therefore, it is essential to identify novel therapeutic targets and drugs, understand the molecular pathogenesis mechanism of DLBCL and find ways to prevent and treat relapsed/refractory DLBCL.
Epigenetic regulation plays an important role in gene expression and also influences tumor progression in many types of cancers, of which histone methylation is a vital posttranslational modification [6-8]. Histone lysine N-methyltransferase EHMT2 (G9a) is a methyltransferase that is mainly responsible for catalyzing mono- and di-methylation of histone H3 lysine 9 (H3K9) in euchromatic regions, and inhibits transcription [9]. Recently, abnormal expression of EHMT2 has been discovered in various types of malignant cells, including acute myeloid leukemia, bladder cancer, lung cancer, squamous cell carcinoma, ovarian cancer, etc. [10-12]. Cancers with elevated expression levels of EHMT2 often show lower expression of some important tumor suppressor genes and epithelial-to-mesenchymal transition (EMT) related genes, so its higher expression may contribute to poor prognosis [13-16]. Studies have proven that inhibiting EHMT2 expression suppressed cell growth and inhibited cancer cell migration and invasion [17]. So EHMT2 is an important methyltransferase and may be an excellent target for anticancer therapy.
BIX-01294 is a small molecule compound that specifically inhibits EHMT2 activity and induces demethylation of H3K9 [18]. Studies have shown that BIX-01294 inhibited cell proliferation and induced apoptosis in many types of cancers including T lymphoblastic leukemia cells, bladder cancer, glioma tumor, renal cancer, and others [10, 19-21]. However, the role of BIX-01294 and the involvement of EHMT2 in DLBCL is not well understood.
In this study, we investigated the effect of BIX-01294 on the proliferation and cell cycle of DLBCL cells, confirmed BIX-01294 induced apoptosis both in extrinsic and intrinsic pathways. Meanwhile, we explored the possible molecular mechanism of this anti-tumor effect.
Materials and Methods
Cell culture
The human DLBCL cell lines U2932, SUDHL2, OCI-LY10, WSU-DLCL2, and DB were obtained from the American Type Culture Collection (Manassas, VA, USA). Cells were grown in IMDM medium and cultured with 10% fetal bovine serum at 37 °C in a humidified atmosphere consisting of 5% CO2. Peripheral blood samples were obtained from two healthy donors. Peripheral blood mononuclear cells (PBMC) were separated using Lymphoprep™ (STEMCELL Technologies, Canada) by centrifugation and cultured with 10% fetal bovine serum.
Antibodies and reagents
BIX-01294 in powder form was purchased from Sigma-Aldrich (Saint Louis, Missouri, USA). EHMT2, DR4, DR5, caspase 3, PARP, Bax, Mcl-1, GRP78, ATF3, CHOP, LC3B, ATF4, β-actin and GAPDH antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA). The c-FLIP antibody was purchased from EZNO Life Sciences (Farmington, NY, USA).
Cell viability assay
Human DLBCL cell lines were seeded in 96-well plates with 2×104 cells per well and treated with BIX-01294 with the indicated concentrations for the indicated times. Then cell viability was measured using Cell Counting Kit-8 (CCK-8) according to the manufacturer's instructions (Dojindo Laboratories, Kumamoto, Japan).
Western blot
Human DLBCL cell lines were seeded in 6-well plates with 2×106 cells per well and treated with BIX-01294 with the indicated concentrations for the indicated times. Cells were harvested and lysed with cell lysis buffer containing protease inhibitor. BCA assay was used to detect protein concentrations and 50 µg protein was used for sample preparation. The sample was electrophoresed and transferred to PVDF membranes and then blocked with 5% skim milk and incubated with primary antibodies overnight. Next day, the membranes were washed with PBST and incubated with HRP-conjugated secondary antibodies, and protein bands were subsequently captured. The relative density of bands was quantified using Image J software.
Apoptosis assay
Human DLBCL cell lines were seeded in 6-well plates with 2×106 cells per well and treated with BIX-01294 with the indicated concentrations for indicated times. Then cells were harvested, and apoptosis was detected using Annexin V-PE/7-AAD Apoptosis Detection Kit (BD Biosciences, San Jose, CA, USA) following the manufacture's protocol.
Cell cycle analysis
Human DLBCL cell lines were seeded in 6-well plates with 2×106 cells per well and treated with indicated concentrations of BIX-01294 for 36 h. Cells were harvested and fixed with 70% ice-cold ethanol at 20 °C overnight. Then cells were washed with PBS and resuspended with 0.25 mL staining solution (50 µg/mL RNase A and 50 µg/mL propidium iodide) for 30 min in the dark, followed by cytometric analysis using FACS Calibur system (BD Biosciences, Franklin Lakes, NJ, USA) and analyzed using ModFit LT 3.3 software (Verity Software House Inc., Topsham, ME, USA).
RNA extraction and real-time qPCR
The cells were harvested, and RNA was extracted with TRIzol™ reagent according to the manufacturer's instruction. The concentration was measured at 260 nm and then reverse-transcripted to cDNA (Roche, Basel, Switzerland). Q-PCR was performed with SYBR® Green Supermix (Roche) using LightCycler480 according to the manufacturer's instruction. The primer sequences are shown in Table 1. The PCR reaction conditions were as follows: 94 °C for 2 min, followed by 35 cycles of 94 °C for 30 s, 60 °C for 30 s, 72 °C for 35 s, and a final extension at 72 °C for 2 min. The 2-ΔΔCT method was used to calculate the relative mRNA expression of target genes.
Sequences of primers for RT-qPCR
| Name | Sequences |
|---|---|
| EHMT2 | 5′-AGTGTGACCCTGACCTCTGT--3′ (forward) |
| 5′-AGATGGTGCCAGCAATAGAT-3′ (reverse) | |
| P21 | 5′-TGAGCCGCGACTGTGATG-3′ (forward) |
| 5′-GTCTCGGTGACAAAGTCGAAGTT-3′ (reverse) | |
| CyclinE | 5′-TTCTTGAGCAACACCCTCTTCTGCAGCC-3′ (forward) |
| 5′-TCGCCATATACCGGTCAAAGAAATCTTGTGCC-3′ (reverse) | |
| DR4 | 5′-TCCAGCAAATGGTGCTGAC-3′ (forward) |
| 5′-GAGTCAAAGGGCACGATGTT-3′ (reverse) | |
| DR5 | 5′-CCAGCAAATGAAGGTGATCC-3′ (forward) |
| 5′-GCACCAAGTCTGCAAAGTCA-3′ (reverse) | |
| ATF4 | 5′-GCTAAGGCGGGCTCCTCCGA-3′ (forward) |
| 5′-ACCCAACAGGGCATCCAAGTCG-3′ (reverse) | |
| ATF3 | 5′-TGATGCTTCAACACCCAGGCC-3′ (forward) |
| 5′-AGGGGACGATGGCAGAAGCA-3′ (reverse) | |
| CHOP | 5′-CATCACCACACCTGAAAGCA-3′ (forward) |
| 5′-TCAGCTGCCATCTCTGCA-3′ (reverse) | |
| REDD1 | 5′-ACGCACTTGTCTTAGCAGTT-3′ (forward) |
| 5′-TAAGCCGTGTCTTCCTCC-3′ (reverse) | |
| LC3B | 5′-CGCACCTTCGAACAAAGAG-3′ (forward) |
| 5′-CTCACCCTTGTATCGTTCTATTATCA-3′ (reverse) | |
| Beclin1 | 5′-GGTTGAGAAAGGCGAGACAC-3′ (forward) |
| 5′-GATGGAATAGGAGCCGCCAC-3′ (reverse) | |
| ATG7 | 5′-ACCCAGAAGAAGCTGAACGA-3′ (forward) |
| 5′-CTCATTTGCTGCTTGTTCCA-3′ (reverse) | |
| ATG5 | 5′-TTTGAATATGAAGGCACACC-3′ (forward) |
| 5′-TGCAATCCCATCCAGAGTTG-3′ (reverse) | |
| β-Actin | 5′-CTCCATCCTGGCCTCGCTGT-3′ (forward) |
| 5′-GCTGTCACCTTCACCGTTCC-3′ (reverse) |
ShRNA design and generation of lentiviruses and construction of stable cell lines
The ATF4 small-hairpin RNA (shRNA) primer sequences were designed as 5′-GCCTAGGTCTCTTAGATGATT-3′ (1), 5′-GCCAAGCACTTCAAACCTCAT-3′ (2). ATF3 shRNA sequences were designed as 5′-GCTGAACTGAAGGCTCAGATT -3′ (1), 5′-GCACCTCTGCCACCGGATG-3′ (2). The shRNA sequences were constructed into a pLVshRNA-eGFP vector. HEK-293T cells were transfected with 10 µg desired plasmid, 7.5 µg of the packaging plasmid psPAX2 and 2.5 µg of the envelope plasmid pMD2.G using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instruction. After 12 h, the culture media was exchanged, and the lentivirus supernatant was collected at 48 h and 72 h for concentration and stored at -80 °C. U2932, WSU-DLCL2, and SUDHL2 cell lines were transduced with lentivirus, screened the stable knockdown cell lines by puromycin, and identified using flow cytometry and western blotting.
Statistical analysis
Data were shown as means ± standard deviation (SD). Two-tailed unpaired Student's t-test was used to analyze the statistical significance between two treatment groups. One-way ANOVAs was used to analyze the statistical significance of multiple treatment groups. Statistical significance was established at *P<0.05, **P<0.01, ***P<0.001. All statistical analyses were performed using the GraphPad Prism software ver. 6.0 (GraphPad Software Inc., La Jolla, CA, USA).
Results
High EHMT2 expression predicts poor survival in DLBCL patients and BIX-01294 inhibits EHMT2 expression in human DLBCL cells
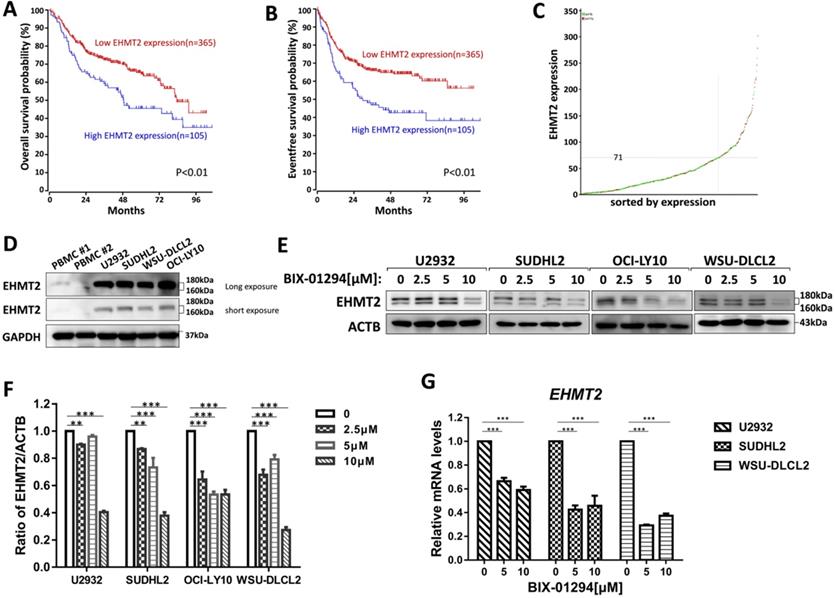
To investigate the prognostic significance of the EHMT2 expression, we analyzed the microarray-based gene expression and clinical outcome data from the R2 genomics database. The Kaplan-Meier analysis was conducted online, and auto scan mode was used to determine the cutoff values for separating the high and low expression groups. As shown in Fig. 1A-1C, patients with high EHMT2 expression had a worse Overall survival (OS) and Event-free survival (EFS) compared with patients with low EHMT2 expression (P<0.01), indicating that high expression of EHMT2 had a prognostic value for predicting EFS and OS. Next, we assessed the expression of EHMT2 in DLBCL cells and peripheral blood mononuclear cells (PBMC) from healthy donors, and found PBMC minimally expressed EHMT2, while DLBCL cells highly expressed EHMT2 (Fig. 1D). Several DLBCL cells were treated with BIX-01294 at different concentrations for 48 h, and western blot showed that BIX-01294 inhibited EHMT2 expression (Fig. 1E and 1F). Using qPCR assay, the mRNA levels of EHMT2 were down-regulated in U2932, SUDHL2, and WSU-DLCL2 cells (Fig. 1G).
BIX-01294 inhibits EHMT2 expression in human DLBCL cells. A, Kaplan-Meier analysis of OS from R2 genomics database. EHMT2 was high expression in 105 out of 470 cases of DLBCL and with poor patient survival. B, Kaplan-Meier analysis of EFS from R2 genomics database. C, The expression level of EHMT2 gene from 470 cases of DLBCL patients. D, U2932, SUDHL2, WSU-DLCL2, and OCI-LY10 cells and human PBMC were harvested and subjected to western blot analysis using EHMT2 antibody. E, U2932, SUDHL2, OCI-LY10, and WSU-DLCL2 cells were incubated with the indicated concentrations (0, 2.5 µm, 5 µm, 10 µm) of BIX-01294 for 48 h. Then whole cells were harvested and subjected to western blot analysis using EHMT2 antibody. F, Gray value analysis of corresponding protein of E. G, U2932, SUDHL2, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 36 h. The expression of EHMT2 mRNA was assessed by real-time PCR. Error bars mean ± SD. **P < 0.01; ***P < 0.001.
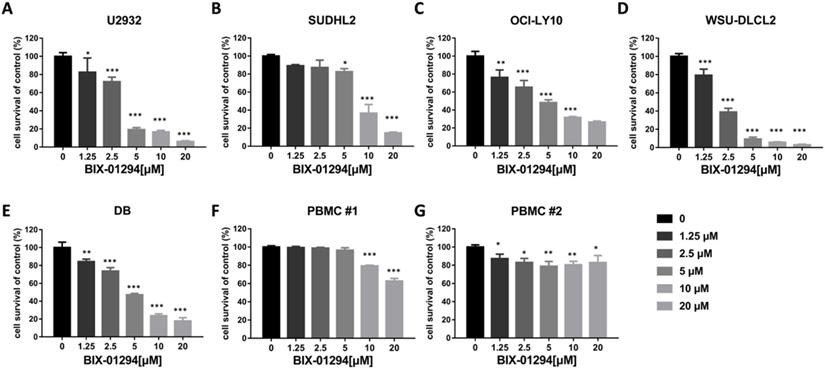
BIX-01294 inhibits DLBCL proliferation and induces G1 phase arrest in human DLBCL cells
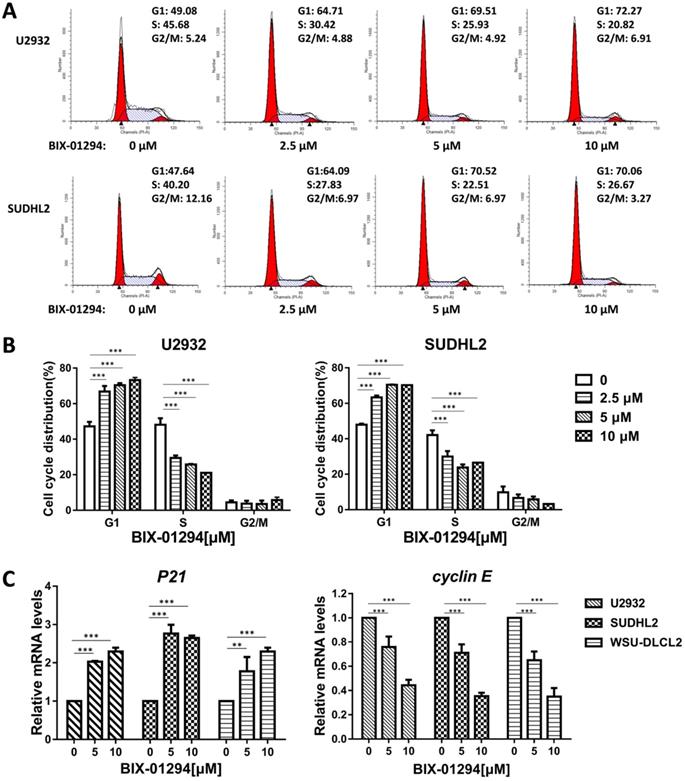
To determine the effect of BIX-01294 on DLBCL cells growth status, CCK-8 assay was performed to detect the viability. As shown in Fig. 2A-E, BIX-01294 exhibited notable proliferation suppression in a dose-dependent manner in five DLBCL cells, no matter the ABC type or GCB type cells. At the same time, we examined the inhibitory effect of BIX-01294 on PBMC, and there was a slight inhibition effect compared to that in DLBCL cells (Fig. 2F and 2G), indicating BIX-01294 exerted a relatively specific inhibition effect for tumor cells. To further study the underlying mechanism that BIX-01294 inhibited DLBCL cells proliferation, flow cytometry analysis was carried out to investigate the cell cycle distribution when treated with BIX-01294 (Fig. 3A and 3B). The cell population in G1 phase was increased both in U2932 and SUDHL2 cells, accompanied with the population in S phase decreasing from 45.68% to 20.82% in U2932 cells and 40.20% to 26.67% in SUDHL2 cells (Fig. 3A and 3B). Moreover, we studied the mechanism of G1 phase arrest, and found the mRNA levels of P21 was increased, and accompanied by cyclin E level was decreased (Fig. 3C). Taken together, these results indicate that BIX-01294 caused G1 phase arrest via increasing P21 level and reducing cyclin E level and then inhibits proliferation in DLBCL cells.
BIX-01294 inhibits human DLBCL cells proliferation. A-G. U2932 (A), SUDHL2 (B), OCI-LY10 (C), WSU-DLCL2 (D), DB (E) and human PBMC (F and G) were incubated with the indicated concentrations of BIX-01294 for 48 h, then CCK-8 assay was performed to detect the viability. Error bars, mean ± SD. Compare with the control group, *P < 0.05; **P < 0.01; ***P < 0.001.
BIX-01294 induces G1 phase arrest in human DLBCL cells. A, U2932 and SUDHL2 cells were incubated with the indicated concentrations (0, 2.5 µm, 5 µm, 10 µm) of BIX-01294 for 36 h, and then the cells were harvested and prepared for cell cycle analysis. B, Percentages of subpopulation of cells at different cell cycle phases based on three independent experiments. C, U2932, SUDHL2, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 36 h. The expression of P21 and cyclin E mRNA was assessed by real-time PCR. Error bars, mean ± SD. **P < 0.01; ***P < 0.001.
BIX-01294 induces apoptosis and activates apoptotic signaling pathway in human DLBCL cells
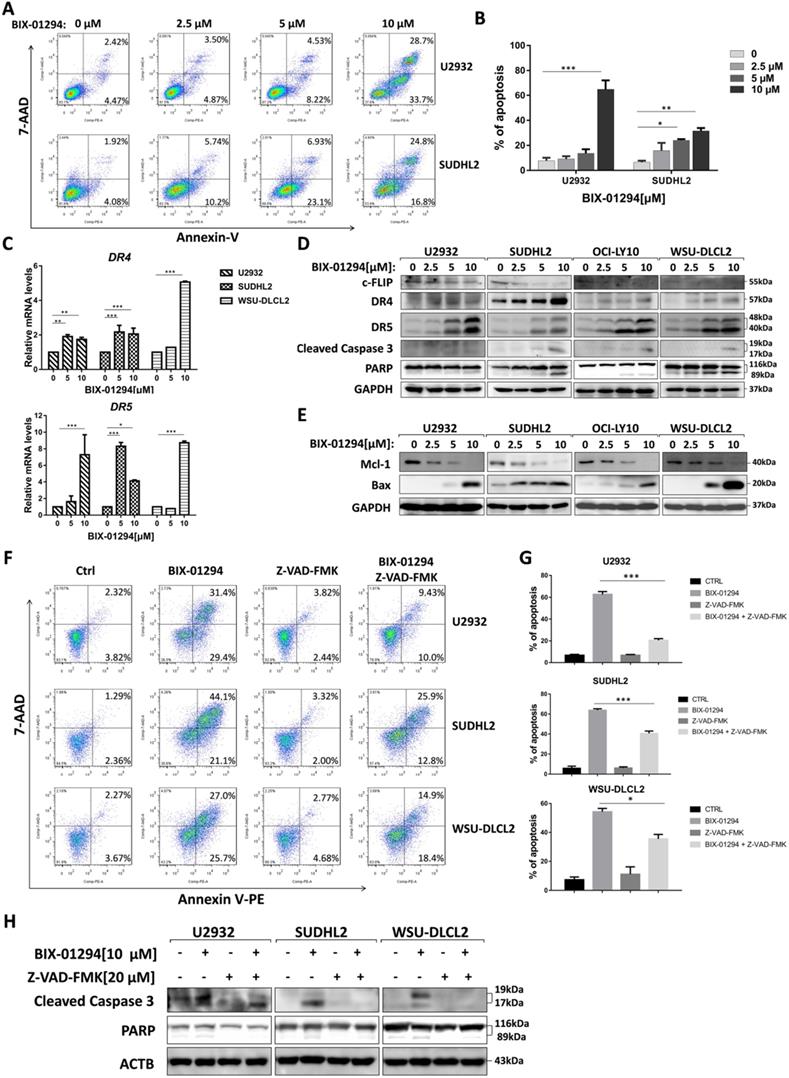
Furthermore, we performed flow cytometric assay to elucidate the apoptotic effect and found that BIX-01294 treatment induced U2932 and SUDHL2 apoptosis. As the concentration increases, the percentage of apoptosis was significantly increased (Fig. 4A and 4B). Particularly, 10 μM BIX-01294 induced about 62% of apoptotic cells in U2932. Meanwhile, we elucidated the apoptotic effect induced by BIX-01294 in Molt-4, Jurkat and PMBC cells as internal and external controls [21] (Supplementary Fig. 1). Then we examined whether the apoptotic signaling pathway was activated. First, we testified that mRNA levels of death receptors DR4 and DR5 were elevated in U2932, SUDHL2 and WSU-DLCL2 cells (Fig. 4C), suggesting BIX-01294 activated exogenous apoptosis pathway. By western blot, we also showed that the expression of anti-apoptotic protein c-FLIP was decreased, and the level of DR4 and DR5 was up-regulated in U2932, OCI-LY10, SUDHL2, WSU-DLCL2 cells (Fig. 4D). Conformably, BIX-01294 increased the cleaved forms of caspase 3 and PARP, further proved that BIX-01294 induces exogenous apoptosis pathway. Meanwhile, the experiments also showed that BIX-01294 down-regulated anti-apoptotic protein Mcl-1 expression and up-regulated pro-apoptotic protein Bax level (Fig. 4E), indicating BIX-01294 also activates endogenous apoptotic pathways. Pretreatment with the caspase inhibitor Z-VAD-FMK for 1 h decreased the cleaved forms of caspase 3 and PARP in U2932, SUDHL2, and WSU-DLCL2 cells (Fig. 4H). The percentage of BIX-01294 induced apoptosis was also significantly reduced with pretreatment with Z-VAD-FMK (Fig. 4F and 4G). Taken together, BIX-01294 activates exogenous and endogenous apoptotic signaling pathways in human DLBCL cells.
BIX-01294 induces apoptosis and activates exogenous and endogenous apoptotic signaling pathway in human DLBCL cells. A, U2932 and SUDHL2 cells were incubated with the indicated concentrations of BIX-01294 for 36 h, cells were harvested and subsequently stained with annexin-V-PE and 7-AAD and analyzed by flow cytometry for apoptosis. B, Percentages of apoptotic cells were determined from three independent experiments. C, U2932, SUDHL2, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 36 h. The expression of DR4 and DR5 mRNA was assessed by real-time PCR. D and E, U2932, SUDHL2, OCI-LY10, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 48 h. Then whole cells were harvested and subjected to western blot using c-FLIP, DR4, DR5, caspase 3, PARP (D) and Mcl-1, Bax (G) antibodies. F-H, U2932, SUDHL2, and WSU-DLCL2 cells were pretreated with Z-VAD-FMK for 1 h and then incubated with BIX-01294 for 48 h. Whole cells were harvested and subjected to analyze by flow cytometry for apoptosis (F) or western blot (H). Percentages of apoptotic cells were determined from three independent experiments (G). Error bars, mean ± SD. *P < 0.05; **P < 0.01; *** P < 0.001.
BIX-01294 triggers autophagy in human DLBCL cells
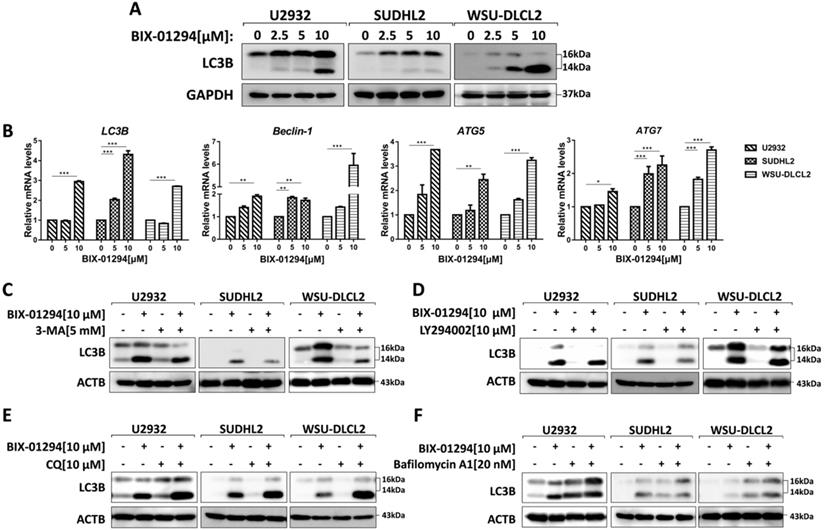
To determine whether BIX-01294 induced autophagy, we treated three human DLBCL cell lines with BIX-01294. LC3B is the hallmark of autophagy, so we examined the expression of LC3B using western blot and qPCR. After treatment with BIX-01294 at the indicated concentration for 48 h, both protein and mRNA levels of LC3B increased in DLBCL cells (Fig. 5A and 5B). We also examined other autophagy markers such as Beclin-1, ATG5, and ATG7, and their mRNA levels were elevated in our tested cells (Fig. 5B). Furthermore, we co-treated DLBCL cells with BIX-01294 and autophagy inhibitors. 3-MA or LY294002, both block the upstream steps of autophagy, reduced LC3B expression when combined with BIX-01294 (Figs. 5C and 5D). While the combination of BIX-01294 and chloroquine (CQ) or bafilomycin A1, both block the downstream steps of autophagy, increased the expression of LC3B (Fig. 5E and 5F). Collectively, these results confirm that BIX-01294 triggers autophagy in human DLBCL cells.
BIX-01294 triggers autophagy in human DLBCL cells. A, U2932, SUDHL2 and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 48 h. Then whole cells were harvested and subjected to western blot analysis using LC3B antibody. B, U2932, SUDHL2, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 36 h. The expression of LC3B, Beclin-1, ATG5 and ATG7 mRNA was assessed by real-time PCR. Error bars, mean ± SD. *P < 0.05; **P < 0.01; *** P < 0.001. C- F, U2932, SUDHL2, and WSU-DLCL2 cells were pretreated with 3-MA (5 mM) (C), LY294002 (10 µM) (D), chloroquine (CQ, 10 µM) (E) or bafilomycin A1 (20 nM) (F) for 1 h and then incubated with BIX-01294 for 48 h. Afterward, the whole cells were harvested and subjected to western blot analysis using the LC3B antibody.
BIX-01294 activates ER stress
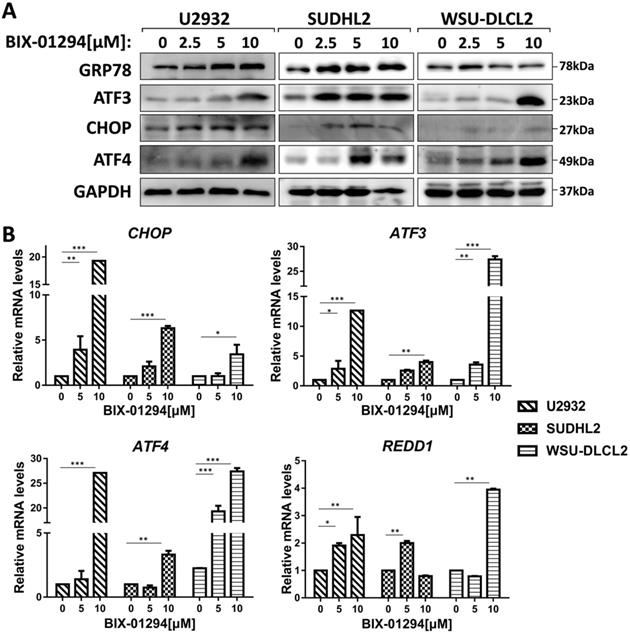
To elucidate the mechanism of BIX-01294-induced apoptosis and autophagy in DLBCL cells, we studied the active stage of endoplasmic reticulum (ER) stress. We examined the expression of GRP78, CHOP, ATF3, and ATF4, which are regarded as important protein markers of ER stress and found that their expression was enhanced when treated with BIX-01294 (Fig. 6A). At the same time, we found that the mRNA levels of CHOP, ATF3, ATF4, and REDD1 were markedly increased (Fig. 6B). Collectively, these results indicate BIX-01294 activates ER stress.
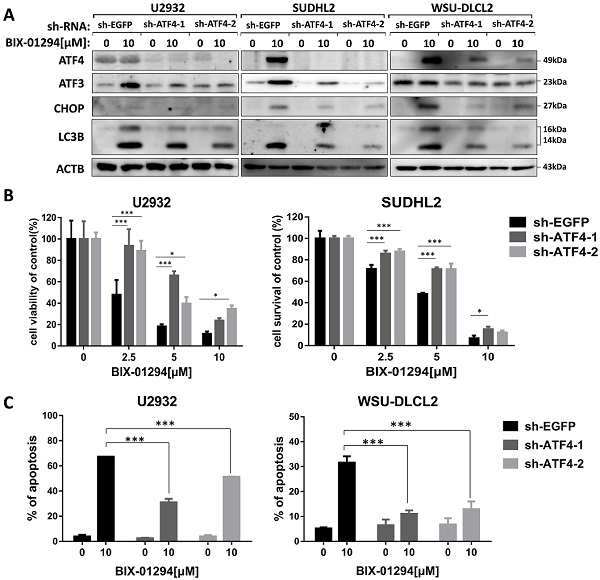
BIX-01294 activates ER stress. A, U2932, SUDHL2, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 48 h. Then whole cells were harvested and subjected to western blot analysis using GRP78, ATF3, CHOP, and ATF4 antibodies. B, U2932, SUDHL2, and WSU-DLCL2 cells were incubated with the indicated concentrations of BIX-01294 for 36 h. The expression of ATF3, CHOP, ATF4 and REDD1 mRNA was assessed by real-time PCR. Error bars, mean ± SD. *P < 0.05; **P < 0.01; ***P < 0.001.
ATF3 expression is required for BIX-01294-induced death
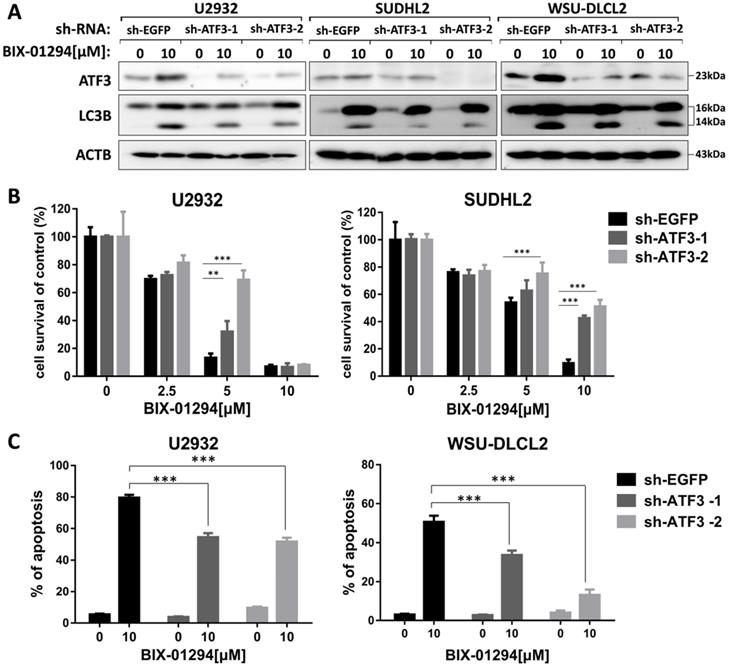
As ATF3 expression is upregulated in BIX-01294 induced cell death, we wondered whether ATF3 influenced apoptosis and autophagy induced by BIX-01294. We performed the shRNA knockdown assay to inhibit ATF3 expression (Fig. 7A) and found that inhibition of ATF3 upregulation decreased the expression of LC3B, indicating ATF3 contributed to BIX-01294 induced autophagy. CCK-8 assay showed that the viability was increased in ATF3 abrogated-U2932 and SUDHL2 cells after exposure to BIX-01294 (Fig. 7B). Consistently, the percentage of apoptosis was significantly decreased in ATF3 knockdown cells than that in the control cells by Annexin V/7-AAD (Fig. 7C). Taken together, we conclude that ATF3 expression is required for BIX-01294-induced autophagy and apoptosis.
ATF3 expression is required for BIX-01294-induced apoptosis and autophagy. U2932, SUDHL2, and WSU-DLCL2 cells were transduced with ATF3-shRNA and control-shRNA lentivirus and constructed stable knockdown cell lines. Then cells were incubated with BIX-01294 for 48 h. A, Whole cells were harvested and subjected to western blot analysis using ATF3 and LC3B antibodies. B, CCK-8 assay was performed to detect the viability. C, The apoptotic rates were detected with Annexin V/7-AAD by flow cytometry, and the percentages of cell apoptosis was determined from three independent experiments. Error bars, mean ± SD. ** P < 0.01;*** P < 0.001.
Inhibition of ATF4 expression decreases BIX-01294-induced death via ATF3
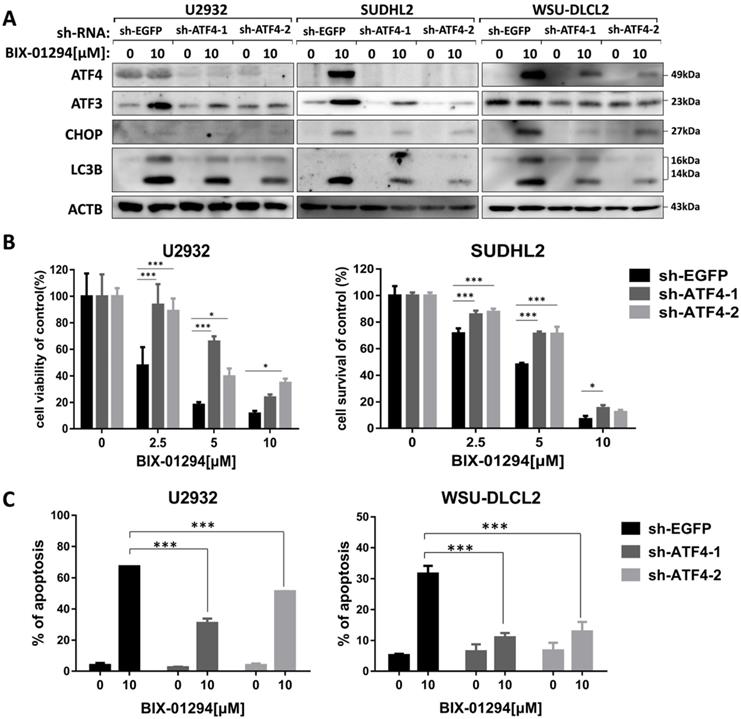
We then explored whether BIX-01294-induced ATF4 upregulation would contribute to LC3B expression. To do this, we successfully blocked BIX-01294-induced ATF4 expression using ATF4 shRNA in U2932, SUDHL2, and WSU-DLCL2 cells (Fig. 8A). We found that the inhibition of ATF4 expression decreased the expression of LC3B, indicating ATF4 expression contributed to autophagy. Besides, we noticed that after abolishing ATF4 expression, BIX-01294-induced cell survival inhibition was decreased in U2932 and SUDHL2 cells as determined using CCK-8 assay (Fig. 8B). Accordingly, the percentage of apoptotic cells was reduced in ATF4-shRNA transfected U2932 and WSU-DLCL2 cells than that in control cells (Fig. 8C). We also showed that suppressed ATF4 expression inhibited ATF3 and CHOP expression in DLBCL cells (Fig. 8A). Taken together, we speculate that BIX-01294 treatment induces ATF4 upregulation, and then promotes ATF3 and CHOP expression, subsequently contributing to BIX-01294-mediated autophagy and apoptosis.
Inhibition of ATF4 expression decreases BIX-01294-induced death via ATF3. U2932, SUDHL2, and WSU-DLCL2 cells were transduced with ATF4-shRNA and control-shRNA lentivirus and constructed stable knockdown cell lines. Then cells were incubated with BIX-01294 for 48 h. A, Whole cells were harvested and subjected to Western blot analysis using ATF4, CHOP, ATF3, and LC3B antibody. B, CCK-8 assay was performed to detect the viability. C, The apoptotic rates were detected with Annexin V/7-AAD by flow cytometry, and the percentages of cell apoptosis were determined from three independent experiments. Error bars, mean ± SD. * P < 0.05; *** P < 0.001.
Discussion
Overexpression of EHMT2 may be associated with a poor prognosis in patients with various types of malignant cancers including lung cancer, ovarian cancer, esophageal squamous cell carcinoma, and so on [14-16]. Several EHMT2 inhibitors have demonstrated good anti-tumor activity by preventing cancer cell proliferation and inducing apoptosis in many types of tumors, such as T lymphoblastic leukemia cells, bladder cancer, glioma tumor, renal cancer, and so on [10, 19-21]. However, the role of BIX-01294 and the involvement of EHMT2 in DLBCL are not well studied to date. In this study, we showed that with the increased concentration of BIX-01294, the cell survival rate of DLBCL was inhibited, and the cell cycle was arrested in G1 phase. Moreover, BIX-01294 induced DLBCL cells apoptosis, and both endogenous and exogenous apoptotic pathways were activated. Furthermore, we found activating the ER stress pathway is one of the underlying mechanisms of how BIX-01294 triggers apoptosis.
In the present study, we showed that BIX-01294 up-regulated the expression of death receptors DR4 and DR5 along with down-regulating anti-apoptotic protein c-FLIP expression, indicating that BIX-01294 induced exogenous apoptosis pathway. Meanwhile, anti-apoptotic protein Mcl-1 expression is decreased, and pro-apoptotic protein Bax expression is increased, suggesting that BIX-01294 also activated endogenous apoptotic pathways. These results were consistent with the results of Woo et al. that demonstrated BIX-01294 induced DR5 expression in renal cancer [19], and the results of Huang et al. that showed BIX-01294 increased Bax expression in leukemia cells [21].
Studies have shown that severe long-term ER stress could induce apoptosis, and DR5 expression is regulated by CHOP and ER stress [22, 23]. We observed that BIX-01294 elevated CHOP expression in protein and mRNA levels. Other important components of ER stress, including GRP78, ATF3, ATF4, and REDD1, were all increased when human DLBCL cells were treated with BIX-01294. These results implied BIX-01294 activated ER stress, which is consistent with other studies that BIX-01294 triggers ER stress response in bladder cancer cells [10]. Next, we analyzed the role of ATF3 and ATF4 in BIX-01294 induced apoptosis. We found that abolishing ATF3 expression effectively decreased the apoptosis induced by BIX-01294. Silencing ATF4 expression also suppressed BIX-01294-mediated cell proliferation inhibition and cell apoptosis. Meanwhile, we detected that inhibiting ATF4 expression obviously decreased ATF3 expression. So these results suggesting that ATF4 facilitated apoptosis through regulating ATF3, and both ATF3 and ATF4 are required for BIX-01294 induced apoptosis. Studies reported that BIX-01294 up-regulated ATF4/CHOP-dependent DR5 expression [24], however, whether ATF4 expression contributes to BIX-01294-induced CHOP and DR5 expression in DLBCL cells still needs to be investigated.
Previous studies reported that BIX-01294 induced autophagy by increasing Beclin-1 expression in breast cancer and human promyelocytic leukemia [13, 25]. Consistently, we proved that BIX-01294 improved LC3B expression both at protein and mRNA levels, and the mRNA levels of autophagy-related molecules Beclin-1, ATG5, and ATG7 were also elevated. In addition, we examined whether BIX-01294 could induce the autophagic flux in DLBCL cells. Combination of BIX-01294 and autophagy inhibitors, such as upstream steps of autophagy inhibitor, 3-MA, or LY294002 decreased LC3B-II formation. In contrast, co-treatment BIX-01294 with downstream steps of autophagy inhibitor, CQ, or bafilomycin A1 increased LC3B-II expression. These results indicated that BIX-01294 induces DLBCL cells autophagy. In-depth studies discovered that inhibition of ATF3 or ATF4 resulted in the decrement of LC3B expression, suggesting that ER stress is involved in regulating autophagy. This is consistent with other studies that ATF4 regulates autophagy [26, 27]. Studies have shown that ATF3 facilitates chemical compounds induced autophagy-associated cell death [28], while other studies have shown that ATF3 negatively regulates cellular autophagy [29]. Our study supported that ATF3 is conducive to autophagy.
Autophagy is also known as type II programmed cell death that is involved in the development of various diseases [30, 31]. It can also be activated by multiple stress stimuli, so the interaction regulation between autophagy and apoptosis is complex. In some cases, autophagy is the cell survival pathway and inhibits apoptosis; while in other cases, autophagy itself also induces cell death, cooperates with apoptosis [32-34]. The two pathways are related and regulate each other. So a comprehensive and in-depth study on the interaction between autophagy and apoptosis may lead to a breakthrough in the understanding and treatment of diseases such as tumors [35, 36]. In our study, we showed that BIX-01294 induced apoptosis and autophagy simultaneously, and ER stress regulated the two cellular physiological processes, so the specific mechanism still needs to be further studied.
In conclusion, we demonstrated that BIX-01294 inhibits the expression of EHMT2 in human DLBCL cells, and then inhibits cell proliferation by inducing G1 phase arrest and inducing apoptosis through both endogenous and exogenous apoptotic pathways. In addition, BIX-01294 triggers autophagy, and activates ER stress in human DLBCL cells. Both key components of ER stress, ATF3, and ATF4, are required for BIX-01294-induced apoptosis and autophagy.
Abbreviations
ABC: activated B-cell-like; ATF: activating transcription factor; CCK-8: cell counting kit-8; CHOP: C/EBP homologous protein; DR: death receptor; DLBCL: diffuse large B-cell lymphoma; EFS: event-free survival; EHMT2: euchromatic histone lysine methyltransferase 2; EMT: epithelial-mesenchymal transition; ER: endoplasmic reticulum; GCB: germinal center B cell-like; NHL: non-Hodgkin lymphoma; H3K9: histone H3 lysine 9; R-CHOP: rituximab, cyclophosphamide, doxorubicin, vincristine and prednisone; RT-qPCR: real-time quantitative polymerase chain reaction; PBMC: peripheral blood mononuclear cells; SD: standard deviation; OS: overall survival.
Supplementary Material
Supplementary figure S1.
Acknowledgements
Funding
The research was supported by National Natural Science Foundation of China (81700199, 81500121); Natural Science Foundation of Jiangsu Province (BK20170257, BK20171181); Jiangsu Provincial Key Research and Development Program (BE2017638); Natural Science Foundation of the Jiangsu Higher Education Institutions of China (17KJB320019); Project funded by China Postdoctoral Science Foundation (2017M611917, 2016M601891) and Jiangsu Planned Projects for Postdoctoral Research Funds (1601099C, 1701134C).
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Authors' contributions
LYX, XG, and PY performed the experiments, WS and JJ analyzed the results, MSN and MDL made the Figures, YYQ, DMY and XGS wrote the manuscript. CS, YT, FZ and XSS analyzed the results. LYZ and ZYL reviewed the manuscript, KLX designed the experiment. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Sehn LH, Gascoyne RD. Diffuse large B-cell lymphoma: optimizing outcome in the context of clinical and biologic heterogeneity. Blood. 2015;125:22-32
2. Katia B, Riccardo DF. Germinal centres and B cell lymphomagenesis. Nat Rev Immunol. 2015;15:172-84
3. Clémentine S, Alexandra TG, Bertrand C. Double-hit and double-protein-expression lymphomas: aggressive and refractory lymphomas. Lancet Oncol. 2015;16:e555-e567
4. Zhang J, Medeiros LJ, Young KH. Cancer Immunotherapy in Diffuse Large B-Cell Lymphoma. Front Oncol. 2018;8:351
5. Riedell PA, Smith SM. Double hit and double expressors in lymphoma: Definition and treatment. Cancer. 2018;124:4622-32
6. Biswas S, Rao CM. Epigenetics in cancer: Fundamentals and Beyond. Pharmacol Therapeut. 2017;173:118-134
7. Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell. 2012;150:12-27
8. Kanwal R, Gupta S. Epigenetic modifications in cancer. Clin Genet. 2012;81:303-311
9. Chen WL, Sun HP, Li DD. et al. G9a - an Appealing Antineoplastic Target. Curr Cancer Drug Tar. 2017;17:555-568
10. Jing C, Wendong S, Xuexi H. et al. EHMT2 inhibitor BIX-01294 induces apoptosis through PMAIP1-USP9X-MCL1 axis in human bladder cancer cells. Cancer Cell Int. 2015;15:1-9
11. Huang T, Zhang P, Li W. et al. G9A promotes tumor cell growth and invasion by silencing CASP1 in non-small-cell lung cancer cells. Cell Death Dis. 2017;8:e2726
12. Bernhard L, Caroline P, Le S. et al. The methyltransferase G9a regulates HoxA9-dependent transcription in AML. Genes Dev. 2014;28:317-327
13. Sang EP, Yi HJ, Suh N. et al. Inhibition of EHMT2/G9a epigenetically increases the transcription of Beclin-1 via an increase in ROS and activation of NF-κB. Oncotarget. 2016;7:39796-808
14. Hyun-Soo C, Kelly JD, Shinya H. et al. Enhanced expression of EHMT2 is involved in the proliferation of cancer cells through negative regulation of SIAH1. Neoplasia. 2011;13:676-84
15. Xinwen Z, Xiaolong C, Xiaojiao G. et al. Overexpression of G9a and MCM7 in oesophageal squamous cell carcinoma is associated with poor prognosis. Histopathology. 2015;66:192-200
16. Hua KT, Wang MY, Chen MW. et al. The H3K9 methyltransferase G9a is a marker of aggressive ovarian cancer that promotes peritoneal metastasis. Mol Cancer. 2014;13:189
17. McGrath J, Trojer P. Targeting histone lysine methylation in cancer. Pharmacol Therapeut. 2015;150:1-22
18. Kubicek S, O'Sullivan RJ, August EM. et al. Reversal of H3K9 me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol Cell. 2007;25:473-81
19. Woo SM, Seo SU, Min KJ. et al. BIX-01294 sensitizes renal cancer Caki cells to TRAIL-induced apoptosis through downregulation of survivin expression and upregulation of DR5 expression. Cell Death Discov. 2018;4:29
20. Ciechomska IA, Przanowski P, Jackl J. et al. BIX01294, an inhibitor of histone methyltransferase, induces autophagy-dependent differentiation of glioma stem-like cells. Sci Rep-UK. 2016;6:38723
21. Huang Y, Zou Y, Lin L. et al. Effect of BIX-01294 on proliferation, apoptosis and histone methylation of acute T lymphoblastic leukemia cells. Leuk Res. 2017;62:34-9
22. Liu G, Su L, Hao X. et al. Salermide up-regulates death receptor 5 expression through the ATF4-ATF3-CHOP axis and leads to apoptosis in human cancer cells. J Cell Mol Med. 2012;16:1618-28
23. Xu LY, Su L, Liu XG. PKC delta Regulates Death Receptor 5 Expression Induced by PS-341 through ATF4-ATF3/CHOP Axis in Human Lung Cancer Cells. Mol Cancer Ther. 2012;11:2174-82
24. Kim SY, Hong M, Heo SH. et al. Inhibition of euchromatin histone-lysine N-methyltransferase 2 sensitizes breast cancer cells to tumor necrosis factor-related apoptosis-inducing ligand through reactive oxygen species-mediated activating transcription factor 4-C/EBP homologous protein-dea. Mol Carcinogen. 2018;57:1492-1506
25. Savickiene J, Treigyte G, Stirblyte I. et al. Euchromatic histone methyltransferase 2 inhibitor, BIX-01294, sensitizes human promyelocytic leukemia HL-60 and NB4 cells to growth inhibition and differentiation. Leuk Res. 2014;38:822-9
26. Matsumoto H, Miyazaki S, Matsuyama S. et al. Selection of autophagy or apoptosis in cells exposed to ER-stress depends on ATF4 expression pattern with or without CHOP expression. Biol Open. 2013;2:1084-90
27. Luo B, Lin Y, Jiang S. et al. Endoplasmic reticulum stress eIF2α-ATF4 pathway-mediated cyclooxygenase-2 induction regulates cadmium-induced autophagy in kidney. Cell Death Dis. 2016;7:e2251
28. Sood V, Sharma KB, Gupta V. et al. ATF3 negatively regulates cellular antiviral signaling and autophagy in the absence of type I interferons. Sci Rep-UK. 2017;7:8789
29. Ouyang L, Zhang L, Fu L. et al. A small-molecule activator induces ULK1-modulating autophagy-associated cell death in triple negative breast cancer. Autophagy. 2017;13:777-8
30. Bento CF, Renna M, Ghislat G. et al. Mammalian Autophagy: How Does It Work? Annu Rev Biochem. 2016;85:685-713
31. Yu L, Chen Y, Tooze SA. Autophagy pathway: cellular and molecular mechanisms. Autophagy. 2017;14:207-215
32. Huang K, Chen Y, Zhang R. et al. Honokiol induces apoptosis and autophagy via the ROS/ERK1/2 signaling pathway in human osteosarcoma cells in vitro and in vivo. Cell Death Dis. 2018;9:157
33. Jiang K, Zhang C, Yu B. et al. Autophagic degradation of FOXO3a represses the expression of PUMA to block cell apoptosis in cisplatin-resistant osteosarcoma cells. Am J Cancer Res. 2017;7:1407-22
34. Nikoletopoulou V, Markaki M, Palikaras K. et al. Crosstalk between apoptosis, necrosis and autophagy. Bba-Mol Cell Res. 2013;1833:3448-59
35. Guillermo MO, Mireia NS, Baehrecke EH. et al. Self-consumption: the interplay of autophagy and apoptosis. Nat Rev Mol Cell Biol. 2014;15:81-94
36. Su M, Mei Y, Sinha S. Role of the Crosstalk between Autophagy and Apoptosis in Cancer. J Oncol. 2013;2013:102735
Author contact
Corresponding author: Kailin Xu, 84 West Huaihai Road, Xuzhou, China. E-mail: lihmdcom; Tel.: 86-516-85802382; Fax: 86-8580-1527.

 Global reach, higher impact
Global reach, higher impact