Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(3):874-884. doi:10.7150/jca.49392 This issue Cite
Research Paper
Identifying Key Genes for Nasopharyngeal Carcinoma by Prioritized Consensus Differentially Expressed Genes Caused by Aberrant Methylation
1. School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, 800 Dong Chuan Road, Shanghai 200240, China.
2. Center for Allergic and Inflammatory Diseases & Department of Otolaryngology, Head and Neck Surgery, Affiliated Eye, Ear, Nose and Throat Hospital, Fudan University, Shanghai 200031, China.
3. CAS Key Laboratory of Computational Biology, CAS-MPG Partner Institute for Computational Biology, Shanghai Institute of Nutrition and Health, Shanghai Institutes for Biological Sciences, University of Chinese Academy of Sciences, Chinese Academy of Sciences, Shanghai 200031, China.
Abstract
Background: Nasopharyngeal carcinoma (NPC) is an Epstein-Barr virus (EBV)-associated epithelial malignancy. Large-scale genetics or epigenetics studies of NPC have been relatively scarce and sporadic, and there are no effective targeted drugs for NPC. Integrative analysis of multiple different omics profiles has been proved to be an effective approach to shed new light on cancer.
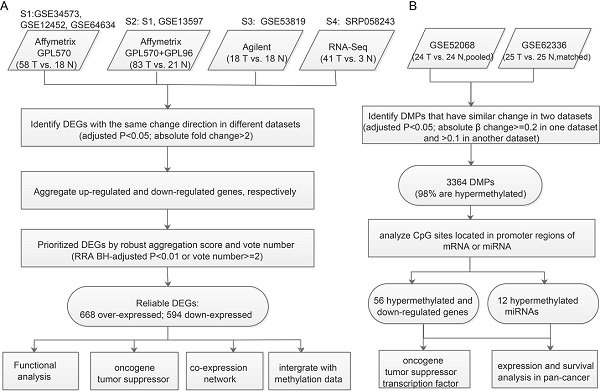
Methods: We developed a pipeline to aggregate consensus differentially expressed genes (DEGs) from multiple expression datasets from different platforms. Integrated bioinformatics analysis of DNA methylation and gene expression was used to prioritize key genes in NPC. We explored the biological and clinical importance of key genes, combining differential co-expression analysis, network analysis of protein-protein and microRNA (miRNA)-target interactions, and pan-cancer survival analysis.
Results: We obtained 668 upregulated and 594 downregulated consensus DEGs, which enriched in the PI3K-AKT, NF-κB and immune-related pathways. In NPC, 98% of 3364 differentially methylated sites were hypermethylated. Actively expressed EBV gene EBNA1 was positively correlated with over-expressed genes coding DNA methyltransferase and Polycomb group proteins, suggesting that EBV infection may have an important role in the hypermethylation of NPC. Through integrated analysis of DNA methylation and mRNA and miRNA expression profiles, we prioritized 56 hypermethylated downregulated genes, including 7 tumor suppressor genes, and constructed a miRNA-target regulation network consisting of 12 hypermethylated miRNAs and 25 upregulated oncogenes. The promoter hypermethylation of PRKCB causing its downregulation was validated by experimental results and higher PRKCB expression was associated with longer overall survival in head-neck squamous cell carcinoma, suggesting the potential of PRKCB as a promising disease biomarker for NPC.
Conclusions: Our integrative analysis provides reliable key genes for candidate biomarkers for diagnosis and prognosis in NPC. Based on the combined evidence of promoter hypermethylation, expression up-regulation, and association with overall survival, genes such as SCUBE2, PRKCB, IKZF1, MAP4K1, and GATA6 could be promising novel diagnostic biomarkers, and miRNAs including MIR150, MIR152, and MIR34 could be candidate prognosis biomarkers.
Keywords: integrative analysis, differently expressed genes, EBV infection, aberrant methylation, disease biomarkers

 Global reach, higher impact
Global reach, higher impact