Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(1):181-189. doi:10.7150/jca.47883 This issue Cite
Research Paper
Loss of LIMCH1 predicts poor prognosis in patients with surgically resected Lung Adenocarcinoma: A study based on Immunohistochemical Analysis and Bioinformatics
1. Department of Respiratory Disease, Thoracic Disease Center, The First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou 310003, China.
2. Department of Pathology, The First Affiliated Hospital, College of Medicine, Zhejiang University, Hangzhou 310003, China.
Abstract
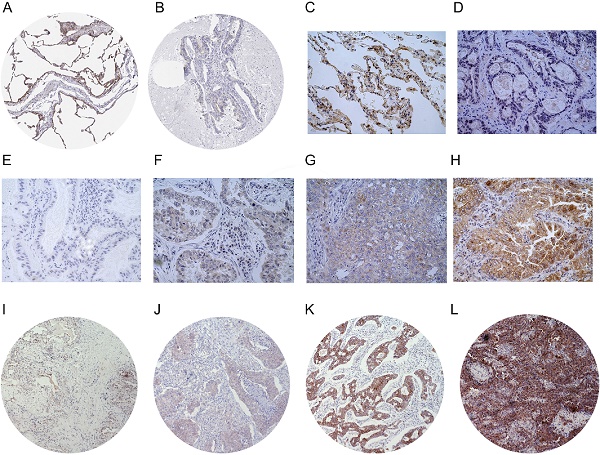
Background: LIMCH1, a novel actin-binding protein, is reported to correlate with tumorigenesis in multiple cancer types, but its clinical prognostic value in lung adenocarcinoma (LUAD) patients remains unclear.
Methods: A total of 196 patients with LUAD who underwent R0 resection were included for analysis. We integrated immunohistochemistry (IHC) and data mining analyses to determine LIMCH1 expression in tumor specimens; the chi-square test was used to explore the correlation between clinicopathologic factors and LIMCH1 expression in LUAD; Kaplan-Meier curves and the Cox proportional hazards model were used to investigate the clinical prognostic role of LIMCH1 expression in patients with LUAD; and DAVID enrichment and gene set enrichment analysis (GSEA) were used to determine the underlying molecular mechanism.
Results: LIMCH1 protein and mRNA expressions were significantly decreased in LUAD tissues. LIMCH1 mRNA expression was a potential diagnostic indicator in the TCGA cohort, and was associated with poor prognosis. IHC results in our LUAD cohort demonstrated that the LIMCH1 expression level was significantly associated with pleural invasion, tumor length, tumor differentiation grade, and clinical tumor stage. Patients with higher LIMCH1 expression had longer overall survival times. Cox multivariate survival analysis showed that LIMCH1 expression independently predicted the outcome. GO and KEGG clustering analyses showed that LIMCH1-related genes may be involved in 'cell adhesion', 'signal transduction', and several cancer-related pathways. GSEA showed 8 enriched hallmarks in the low LIMCH1 expression group, including mTOR signaling, MYC signaling, DNA repair, and G2M checkpoint.
Conclusions: Our findings suggest that LIMCH1 may serve as a promising biomarker to predict LUAD prognosis.
Keywords: lung adenocarcinoma, LIMCH1, TCGA, immunochemistry, prognosis

 Global reach, higher impact
Global reach, higher impact