Impact Factor ISSN: 1837-9664
J Cancer 2020; 11(14):4037-4046. doi:10.7150/jca.30360 This issue Cite
Research Paper
Plasma exo-hsa_circRNA_0056616: A potential biomarker for lymph node metastasis in lung adenocarcinoma
1. Department of Epidemiology and Health Statistics, School of Public Health, Fujian Medical University, Fuzhou, 350108, China; Fujian Provincial Key Laboratory of Tumor Microbiology, Fujian Medical University, Fuzhou, 350108, China; Key Laboratory of Ministry of Education for Gastrointestinal Cancer, Fujian Medical University, Fuzhou, 350108, China
2. Department of Science and Education, The Affiliated Longyan First Hospital of Fujian Medical University, Longyan, 364000, China
3. Department of Chest Surgery, The First Affiliated Hospital of Fujian Medical University, Fuzhou, 350001, China
#The first two authors contributed equally to this paper.
Received 2018-10-2; Accepted 2020-1-4; Published 2020-4-6
Abstract
Background: To investigate the relationship between CXCR4-related circular RNAs (circRNAs) in exosomes and lymph node metastasis of lung adenocarcinoma.
Methods: Totally 41 lung adenocarcinoma tissues (21 with lymph node metastasis and 20 without) were collected. Expression of CXCR4 protein was detected by western blotting analysis. A stable PC9/CXCR4-shRNA and PC14/CXCR4-shRNA knockdown lung adenocarcinoma cell lines were established and subjected to functional assays (cell proliferation, colony formation, migration and invasion) for phenotype changes. Exo-hsa-circRNAs (has-circRNAs in exosomes) were detected in vivo and in vitro. The diagnostic value of differentially expressed exo-has-circRNAs was evaluated.
Results: Expression levels of CXCR4 were higher in patients with lymph node metastasis than in those without (P = 0.001). Silencing CXCR4 expression in PC9 and PC14 cell lines with short hairpin RNA could effectively abolish colony formation frequency, proliferation rate, migration rate, and the number of invasive cells (all P < 0.001). Exo_circRNA_0056616 was detected in both PC-9/CXCR4-shRNA cells and lung adenocarcinoma plasma at significantly higher levels than in the corresponding control (P < 0.001). When a receiver operating characteristic (ROC) curve for plasma exo-hsa_circRNA_0056616 levels and diagnosis of lymph node metastasis of lung adenocarcinoma was generated, a cutoff value of 0.394 was identified with an area under the curve of 0.812 (95% confidence interval 0.720-0.903), a sensitivity of 0.792, and specificity of 0.810.
Conclusions: Taken together, our findings suggested that CXCR4 was higher in the lung adenocarcinoma tissues with lymph node metastasis. Higher plasma levels of exo-hsa_circRNA_0056616 in these patients also suggest that this circRNA represents a potential biomarker for lymph node metastasis predictor in lung adenocarcinoma.
Keywords: Lung adenocarcinoma, CXCR4, exosomes, circRNA, lymph node metastasis
Background
Lung cancer is one of the most frequent malignancies and leads cause of cancer-related deaths worldwide. The number of new cases of lung cancer is estimated to be 1.82 million (representing 12.9% of the total number of cancer cases diagnosed), while the number of deaths due to lung cancer is estimated to be 1.59 million (19.4% of all deaths due to cancer) (1). The high incident areas include China, which the number of new cases and deaths due to lung cancer being 73 million and 61 million, respectively, in 2015 (2). 80 % of lung cancers are non-small cell lung cancer (NSCLC), with the primary pathology being lung adenocarcinoma. Due to the aggressive and metastatic phenotype of lung adenocarcinoma, the prognosis is extremely poor.
Tumor metastasis is a critical step in the malignant progression of tumors. Many studies have demonstrated that the chemokine receptor, CXCR4, is overexpressed in tumor tissues to regulate autocrine and paracrine signaling, angiogenesis, proliferation immune responses and other processes (3). It has been demonstrated that CXCR4 can bind to SDF-1, which activates ERK/AKT signaling or PI-3K signaling to regulate tumor cell proliferation (4). Taken together, these observations support the proposal that CXCR4 may be a useful biomarker for predicting tumor progression (5).
Exosomes secreted by the cells is a kind of extracellular vesicles with lipid bilayer structure, containing a variety of cell specific proteins, lipid, and nucleic acids, involved in intercellular communication, plays an important role in different physiological and pathological process (6). Exosomes can also carry non-coding RNAs (ncRNA) in the blood to achieve long-distance cell communication (6) and can promote microenvironments that facilitate the formation of lung cancer, as well as its invasive and metastatic phenotypes (7-10).
Circular RNAs (circRNAs) are a class of endogenous non-coding RNAs molecules with closed loops and high stability. Previously, circRNAs were considered miRNA sponges, and can bind to single or multiple miRNAs and regulate the expression of their downstream genes. However, a large number of studies have revealed the transcriptional and translational regulatory functions of circRNAs in various diseases, including cancers (11). CircRNAs are abundant and stable in exosomes and can continually play their roles after the exosomes are taken up by neighboring cells. When an expression profile of circRNAs was determined for exosomes obtained from serum samples from patients with lung cancer, this profile significantly differed from the profile obtained in parallel from healthy individuals (12). Thus, circRNAs may as biomarkers in lung cancer.
In this study, the aims were to: i) confirm whether expression of CXCR4 is associated with lymph node metastasis of lung adenocarcinoma in vivo; and ii) screen and identify circRNAs present in exosomes (exo-hsa_circRNA) that are associated with CXCR4 expression and have potential to serve as predictors of lymph node metastasis in lung adenocarcinoma.
Methods
Subjects and samples
A total of 90 lung adenocarcinomas samples, including 42 with lymph node metastasis and 48 without lymph node metastasis, were obtained from the Fujian Provincial Cancer Hospital (Fujian, China). Both groups of patients were diagnosed with lung cancer after pathological diagnosis, and the pathological type was adenocarcinoma. No chemotherapy or radiotherapy was performed before surgery; informed consent was signed; clinical data and tissue specimens were complete. Metastasis was determined based on postoperative pathological reports according to the following criteria: the number of lymph node dissection is ≥5, the number of metastasis is ≥1 for lymph node metastasis; the number of lymph node dissection is ≥5, and the number of metastasis=0 is no lymph node metastasis. Exclusion of severe cardiac and brain disease, secondary adenocarcinoma and recurrence of lung adenocarcinoma.
Peripheral venous blood (5 mL) was collected from each patient prior to surgery into EDTA anticoagulant tubes. After a centrifugation step at 2000 rpm for 10 min, the samples were transferred to RNase-free centrifuge tubes and stored at -80 °C. Meanwhile, 41 tumor tissue specimens were immediately preserved in liquid nitrogen after their resection and were stored at -80 °C until analyzed.
Information regarding patients and their samples were obtained with permission from the Institutional Medical Ethics Committee of Fujian Medical University. Written informed consent forms were obtained from all the participating subjects.
Cell culture
PC9 cells and PC14 cells were obtained from YUBO Biological Company (Shanghai, China) and 293T cells were obtained from ATCC (Shanghai, China). Cell lines were maintained in Dulbecco's Modified Eagle Medium (DMEM, Gibco, NY, USA) supplemented with 10% fetal bovine serum (FBS, Gibco) and 1% penicillin-streptomycin (Gibco) at 37 °C in a 5% CO2 humidified incubator.
Western blotting analysis
Cells were lysed in whole-cell lysate buffer (Solarbio, Beijing, China) and then resolved by 10% or 12% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). The separated proteins were then transferred to polyvinylidene difluoride (PVDF) membranes (Pall, NY, USA) and blocked with 5 % skim milk. The membranes were incubated with primary antibodies overnight at 4 °C and with appropriate secondary antibodies at room temperature for 1 h. The primary antibodies recognized CXCR4 and actin (Abcam, MA, USA), glyceraldehyde 3-phosphate dehydrogenase (GAPDH, Santa Cruz Biotechnology, TX, USA), CD81 and CD63 (System Biosciences, CA, USA). Protein bands were visualized with reagents from an enhanced chemiluminescence kit (Pierce, IL, USA) and band intensities were analyzed with ImageJ software (NIH, MD, USA). Detection of actin was performed as a loading control.
Generation and characterization of a stable cell line expressing low levels of CXCR4
Short hairpin RNAs (shRNAs) targeting CXCR4 were designed with online RNAI design software (http://www.protocol-online.org/prot/Research_Tools/Online_Tools/SiRNA_Design/) and synthesized by Shanghai Sangon Ltd. (Shanghai, China). These shRNAs were each cloned into the PLVX-shRNA-PURO lentivirus vector (Takara Biomedical Technology, Beijing, China) to generate fusion proteins with ZsGreen fluorescent protein (Takara Biomedical Technology). The corresponding plasmids were then transfected into 293T cells with Xfect Transfection reagent (Takara Biomedical Technology), according to the manufacturer's directions. Puromycin selection and screening of the transfected cells based on GFP expression resulted in the identification of cells to produce lentivirus. Infections of the PC9 and PC14 cells were conducted with an Endo-free Plasmid Maxi Kit (OMEGA, Norcross, GA) and Lenti-X™ Lentiviral Expression Systems (Takara Biomedical Technology, Beijing, China), according to the manufacturers' protocols. The infected cells were named PC9/CXCR4-shRNA and PC9/shCtrl cells, PC14/CXCR4-shRNA and PC14/shCtrl cells, respectively. GFP expressions in these two sets of infected cells were confirmed with a fluorescence inverted microscope and the infected cells were subsequently cultured as single cell colonies. After expressions of the appropriate shRNAs were confirmed, these cell lines were used for experimentation.
Soft agar colony assay
PC9/CXCR4-shRNA, PC9/shCtrl, and PC9, PC14/CXCR4-shRNA, PC14/shCtrl, and PC14 cells were digested and counted to adjust the cell density of each to 1 × 103 cells/mL. Approximately 500 cells (0.5 ml) of each cell suspension were then added to volumes of 9.4 ml DMEM. Twenty-four well plates containing 5% agar at 37 °C had 40 cells added per well, with triplicate wells prepared in duplicate for each sample. Clone formation rates (%) were calculated as follows: [number of clones observed × 5 / number of inoculated cells] x 100%.
CCK8 assay
For the CCK8 assays, PC9/CXCR4-shRNA, PC9/shCtrl, PC9, and PC14/CXCR4-shRNA, PC14/shCtrl, and PC14 cells were plated in 96-well plates (4 × 103 cells/well). After 24 h at 37 °C and 5% CO2, 10 mL of CCK8 buffer was added to each well. After 1-2 h, optical density values were measured at 450 nm with a Multiskan™FC microplate photometer (ThermoFisher Scientific, MA, USA). The cell viability values determined for each group were subsequently normalized to the cell viability value of the untreated control cells. This assay was repeated at least three times.
Wound healing assay
PC9/CXCR4-shRNA, PC9/shCtrl, PC9, and and PC14/CXCR4-shRNA, PC14/shCtrl, and PC14 cells suspension were adjusted to 3 × 106 cells/mL. Then, 200 μL of each cell suspension was added to 2 mL of complete medium in 6-well plates. When the cells reached 80% confluency, a pipette was used to draw a vertical line in each of the wells. The scratched wells were incubated for 48 h and then each well was imaged to evaluate cell migration.
Transwell assay
For the transwell assays, the bottom membrane of each well (12 mm, Corning, MA, USA) was coated with 50 mg/L matrigel (diluted 1:8 in DMEM) then air dried at 4 °C. PC9/CXCR4-shRNA, PC9/shCtrl, PC9, and PC14/CXCR4-shRNA, PC14/shCtrl, and PC14 cells suspension densities were adjusted to 5 × 105 cells/mL, and 100 μL of each cell suspension was added to the top of each Transwell chamber. One mL of DMEM/10% FBS was added to the lower chamber. After 24 h, cotton swabs were used to wipe each upper membrane and then the Transwells were placed in 70% alcohol to fix the cells on the lower membrane. Subsequently, these cells were stained with 100 μL Giemsa stain for 5 min before the membranes were imaged and the stained cells were counted.
Analysis of plasma exosomes
Plasma exosomes were extracted from lung adenocarcinoma samples and PC9/CXCR4-shRNA, PC9/shCtrl, and PC9 cells by using an ExoQuick Isolation kit (System Biosciences, USA).
Total RNA was subsequently extracted from these samples with a QIAGEN RNeasy Mini kit (Qiagen, Duesseldorf, Germany), according to the manufacturer's instructions. Real-time quantitative reverse transcription-polymerase chain reactions (RT-qPCR) were performed with SYBR® Premix Ex TaqTM II (Tli RNaseH Plus, TaKaRa, Dalian, China), according to the manufacturer's instructions. Divergent primers for exo-hsa_circRNA_0056616 and primers for CXCR4 and GAPDH, which was to be the reference gene, were designed by Biocan (Shenzhen, China) and synthesized by TaKaRa. The normalization of qPCR in the expression of exo-hsa circRNA 0056616 was referring to Wang's methods (13). All RT-qPCR primer sets are listed in Table S1 and S2.
Statistical analysis
Statistical analysis was performed with SPSS 22.0 (IBM, NY, USA). Differences in the values for exo-hsa_circRNA_0056616 and CXCR4 between groups were determined by using the t-test and analysis of variance (ANOVA). Wilcoxon rank sum test and Pearson correlation analysis were used to determine correlations between expression levels of exo-hsa_circRNA_0056616 and CXCR4 and clinical indexes. A receiver operating characteristics (ROC) curve was generated for exo-hsa_circRNA_0056616 for lymph node metastasis detection. A P value less than 0.05 was considered statistically significant.
Results
CXCR4 was upregulated and associated with lymph node metastasis in lung adenocarcinoma
Expression of CXCR4 in 41 lung adenocarcinoma tissues (21 lymph node metastasis tissues and 20 without) was detected by western blotting analysis, Actin was used as a loading control (Fig. S1A). Results showed that the median expression value of CXCR4 was 0.576 (interquartile range, 0.371-0.757) in 41 lung adenocarcinoma tissues. The median levels of CXCR4 expression was upregulated in metastasis tissues 0.624 (interquartile range: 0.564-0.998) than in those of without metastasis tissues 0.432 (interquartile range: 0.294-0.577) (Z = -3.417, P = 0.001) (Fig. S1B). Together, these results suggest that overexpression of CXCR4 in lung adenocarcinoma is related to lymph node metastasis.
Establishment of stable CXCR4 knockdown cell lines
To determine whether shRNAs affects the expression of CXCR4 in a lung adenocarcinoma cell line, three shRNAs (shRNA-1, shRNA-2, and shRNA-3) specifically targeting CXCR4 were generated with lentivirus system, respectively. As shown in Figure S2A, electrophoresis results showed that shRNA clones 3-8, and a representative Sanger sequencing read for one of those clones is shown in Figure S2B. three shRNAs specifically targeting CXCR4 were transfected into PC9 cell line, as well as a scrambled control shRNA (shRNA-c), respectively. In RT-qPCR assays revealed that three shRNAs could specifically down-regulate the levels of CXCR4 compared with the controls (P < 0.05) (Fig. S2C). However, shRNA1, subsequently referred to as CXCR4-shRNA1 (or PC9-KD in the figures) was selected to perform subsequent experiments. The CXCR4-shRNA1, PC9/shCtrl (or PC9-C in the figures) and untransfected PC9 cells were detected with microscopy and fluorescence microscopy (Fig. S2D). The silencing effect was checked by western blotting and RT-qPCR analysis, and results showed that shRNA1 could specifically down-regulate expression of CXCR4 (Fig. S2E and S2F).
The levels of CXCR4 protein detected in the PC9/CXCR4-shRNA1 cells versus the PC-9/shCtrl cells were 0.239 ± 0.016 versus 1.016 ± 0.020, respectively (P < 0.001) (Fig. S2E). In the RT-qPCR assays, the relative expression levels of CXCR4 mRNA were 0.288 ± 0.013 for the PC9/CXCR4-shRNA1 cells, and 1.047 ± 0.056 and 1.001 ± 0.055 for the PC9/shCtrl and un-transfected PC9 cells, respectively (P < 0.001) (Fig. S2F). Therefore, a significant 3.6-fold decrease in CXCR4 mRNA levels was achieved by using CXCR4-shRNA1 to target CXCR4 (P < 0.001).
Knockdown of CXCR4 abolishes a typical metastasis phenotype
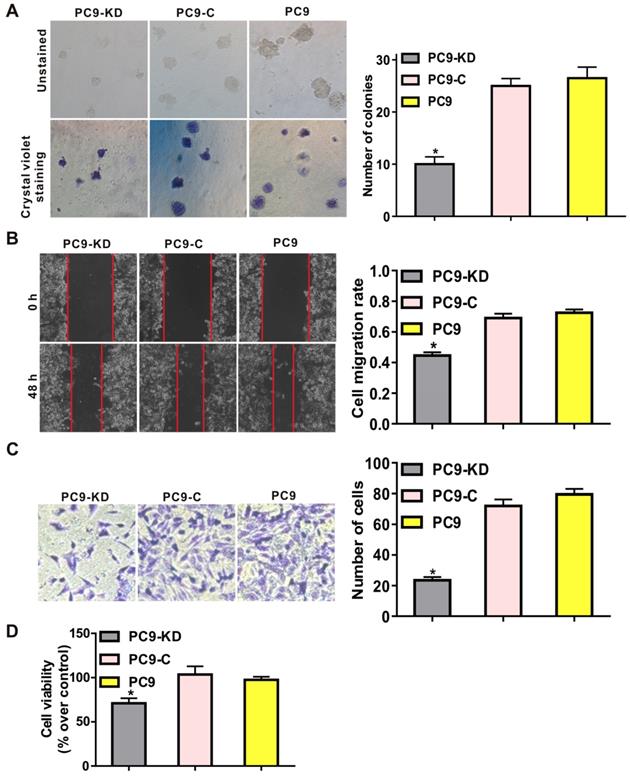
To characterize the function of CXCR4 silencing in a lung adenocarcinoma cell line, functional assays revealed that CXCR4 silencing could effectively inhibit the tumorigenic phenotype by reducing frequencies of colony formation in soft agar compared with cells treated with scramble shRNAs (P < 0.001) (Fig. 1A and Fig. S3A). To investigate the effects of CXCR4 silencing on tumor metastasis, both migration and invasion assays showed that migratory and invasive abilities were significantly decreased in PC9/CXCR4-shRNA1 and PC14/CXCR4-shRNA1 cells compared to the control cells (P < 0.001) (Fig. 1B and C; Fig. S3B and C). To evaluate cell proliferation, CCK8 assay demonstrates that CXCR4 silencing could decrease cell viability compared to the control cells (P < 0.01) (Fig. 1D). Taken together, all these results indicate that CXCR4 silencing regulates the hallmarks of the metastasis process by inhibiting colony formation, cell proliferation, migration and invasive in lung adenocarcinoma cell line.
Effect of CXCR4 silencing on various phenotypes of the PC9 cell line. Low levels of CXCR4 expression were compared with higher levels of CXCR4 expression in: (A) soft agar colony formation assays, (B) wound healing assays, (C) transwell assays, and (D) CCK8 assays. These assays were performed to evaluate colony formation, cell migration, cell invasion and cell viability, for the PC9 cell groups indicated, respectively. In panel C, crystal violet stained cells (at left) and the corresponding quantitation of these stained cells (at right) indicate the number of invasion cells for each group. PC9-KD: PC9/CXCR4-shRNA1 group; PC9-C: PC9/shCtrl group; PC9: untransfected PC9 group.
CXCR4-regulated PC9 phenotype changes associated with hsa_circRNA_0056616
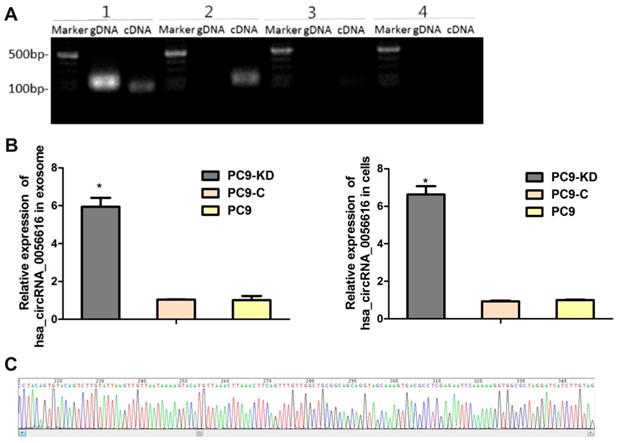
A search was performed of the circbase, circRNABase, and Circ2Traits databases to identify potential circRNAs that may be associated with CXCR4. Hsa_circRNA_0056615, hsa_circRNA_0056616, and hsa_circRNA_0117403 were identified (Table S3). RT-qPCR was performed to detect these circRNAs, as shown in Fig. 2A, the only of hsa_circRNA_0056616 was detected in the cells and exosomes of the PC9/CXCR4-shRNA1 cells. And the sequence of amplified hsa_circRNA_0056616 DNA was verified by sanger sequencing (Fig. 2B). Futhermore, the expression levels of hsa_circRNA_0056616 were detected in exosomes that were isolated from three PC9 groups by RT-qPCR assays, our data have demonstrated that the levels of hsa_circRNA_0056616 were found to be 5.940 ± 0.479 for the PC-9/CXCR4-shRNA1 groups, and 1.034 ± 0.016 and 1.015 ± 0.219 for the negative control PC-9/shCtrl and blank control PC9 groups, respectively (P < 0.001) (Fig. 2C, left panel). Meanwhile, the mean expression level of hsa_circRNA_0056616 in the PC9/CXCR4-shRNA1 cells was 6.631 ± 0.439, compared with levels of 0.931 ± 0.045 and 1.000 ± 0.030 in the negative control PC9/shCtrl and blank control PC9 cells, respectively (P < 0.001) (Fig. 2C, right panel). Thus, our findings indicate that the hsa_circRNA_0056616 is remarkably upregulated in PC9/CXCR4-shRNA1 cells and PC9/CXCR4-shRNA1 cells-derived exosomes.
Identification and detection of hsa_circRNA_0056616 in lung adenocarcinoma cells and exosomes. (A) Based on a literature search, three circRNAs were previously reported to be related to CXCR4. These were detected in genomic DNA (gDNA) and complementary DNA (cDNA) samples by PCR. [1: GAPDH (as a control); 2: hsa_circRNA_0056616; 3: hsa_circRNA_0117403; 4: hsa_circRNA_0056615]. Marker: DNA ladder. (B) Hsa_circRNA_0056616 was detected by PCR in exosomes and cells from the cell groups indicated. *P < 0.001. PC9-KD: PC9/CXCR4-shRNA1 group; PC9-C: PC9/shCtrl group; PC9: blank control group. (C) Sequencing of hsa_circRNA_0056615.
Exo-hsa_circRNA_0056616 as a potential marker of lymph node metastasis of lung adenocarcinoma
For this study, 42 patients with lung adenocarcinoma metastasis, including 19 male patients and 23 female patients (mean age, 57.71 ± 8.41 y) were enrolled. In addition, 48 patients with non-metastatic lung adenocarcinoma, including 25 males and 23 females (mean age, 59.60 ± 7.47 y) were enrolled. The gender proportions and mean ages for the two groups have no significant difference (Table 1).
Plasma levels of exo-hsa_circRNA_0056616 and clinicopathological features of the cohort examined.
| Clinicopathological features | n | Exo-hsa_circRNA_0056616 | |
|---|---|---|---|
| M (P25, P75) | P-value | ||
| Gender | |||
| Male | 44 | 0.569 (0.254, 1.102) | 0.765 |
| Female | 46 | 0.370 (0.218, 1.237) | |
| Age | |||
| < 60.5 y | 45 | 0.467 (0.168, 1.344) | 0.869 |
| ≥ 60.5 y | 45 | 0.394 (0.274, 0.846) | |
| T stage | |||
| T1 / T2 | 61 | 0.658 (0.309, 1.476) | 0.002 |
| T3 / T4 | 27 | 0.282 (0.156, 0.547) | |
| M stage | |||
| M0 | 65 | 0.658 (0.306, 1.344) | 0.004 |
| M1 | 25 | 0.288 (0.165, 0.515) | |
| TNM classification | |||
| I-II | 50 | 0.830 (0.325, 1.730) | 0.001 |
| III-IV | 37 | 0.287 (0.164, 0.380) | |
| Tumor size* | |||
| (Maximum diameter of tumor) | 33 | 0.884 (0.423, 1.476) | 0.122 |
| < 2.40 cm | 33 | 0.347 (0.283, 1.243) | |
| Anatomical classification | |||
| Central | 0.438 (0.161, 1.132) | 0.205 | |
| Surrounding | 0.672 (0.309, 1.385) | ||
M: median; P25: 1st quartile (Q1); P75: 3rd quartile (Q3).
We examined exosomes morphology in plasma using transmission electron microscopy (Fig. 3A). Our results showed a rounded and oval-shaped vesicle with diameters ranging from 40-100 nm exhibited obvious heterogeneity. Furthermore, as shown in Figure 3C, western blotting analysis for exosomes marker proteins CD63 and CD81 were verified in plasma sample collected from a lung adenocarcinoma patient with metastasis.
Exo-hsa_circRNA_0056616 as a potential marker of lymph node metastasis of lung adenocarcinoma. (A) Plasma exosomes of lung adenocarcinoma patients were examined with transmission electron microscopy. Rounded and oval-shaped vesicles exhibiting obvious heterogeneity were observed. The diameters of these cells ranged from 40-100 nm. Scale bar: 100 nm. (B) Expression levels of exo-hsa_circRNA_0056616 in 90 plasma samples according to the metastasis status of the samples (meam±standard deviation). (C) A representative western blot of the detection of CD63 and CD81 in plasma samples from a lung adenocarcinoma patient with metastasis. (D) Plasma levels of exo-hsa_circRNA_0056616 expression was plotted in relation to levels of CXCR4 detected in the corresponding samples. (E) The ROC curve (in blue) generated from the detection of exo-has_circRNA_0056616 in 90 plasma samples in relation to the diagnosis of lymph node metastasis of lung adenocarcinoma for the samples. The green line represents non predictive values.
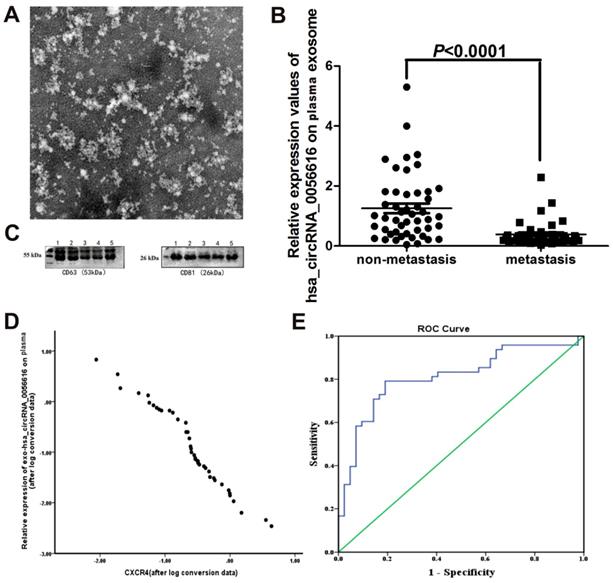
When expression of exo-hsa_circRNA_0056616 was detected in plasma samples collected from this cohort of lung adenocarcinoma patients, the patients with lymph node metastasis had significantly lower levels of exo-hsa_circRNA_0056616 than the patients without lymph node metastasis (Z = -5.079, P = 0.001) (Fig. 3B). Moreover, expression of exo-hsa_circRNA_0056616 was found to negatively correlate with CXCR4 expression (r = -0.979, P < 0.05) (Fig. 3D). Furthermore, higher expression levels of exo-hsa_circRNA_0056616 in plasma correlated with higher protein levels of CXCR4 in lung adenocarcinoma tissue.
Levels of exo-hsa_circRNA_0056616 in plasma were further examined in relation to T1/T2 lung adenocarcinoma versus T3/T4 lung adenocarcinoma. The T1/T2 stage patients had higher plasma levels of exo-hsa_circRNA_0056616 than the T3/T4 stage patients, and the difference was statistically significant (P = 0.002) (Table 1). In addition, the patients classified with lung adenocarcinoma M0 or lung adenocarcinoma I-II had higher levels of plasma exo-hsa_circRNA_0056616 than the patients with lung adenocarcinoma M1 (P = 0.004) or lung adenocarcinoma III-IV (P = 0.001), respectively (Table 1). In contrast, plasma levels of exo-hsa_circRNA_0056616 did not correlate with the age, gender, tumor size, or anatomical classification of lung adenocarcinoma in the patients examined (Table 1). In addition, a significant negative correlation was observed between the expression of exo-hsa_circRNA_0056616 and T stage, M stage, and TNM grade of lung adenocarcinoma (P < 0.05) (Table 2). Moreover, there was no significant correlation between tumor size and anatomical classification (P > 0.05) (Table 2).
Correlations between plasma levels of exo-hsa_circRNA_0056616 and lung adenocarcinoma features.
| Clinicopathological features | Plasma hsa_circRNA_0056616 | |
|---|---|---|
| rs | P-value | |
| T stage | -0.318 | 0.003 |
| M stage | -0.308 | 0.003 |
| TNM classification | -0.466 | 0.001 |
| Tumor size | -0.232 | 0.060 |
| Anatomical classification | 0.154 | 0.207 |
When a ROC curve for plasma exo-hsa_circRNA_0056616 levels and diagnosis of lymph node metastasis of lung adenocarcinoma was generated (n = 90) (Fig. 3E), the area under curve (AUC) was 0.812 (standard error: 0.047; 95% confidence interval, 0.720, 0.903), the maximum Jordan index value was 0.602, and the sensitivity and specificity were 0.792 and 0.810, respectively. As a result, a cutoff value of 0.394 was identified.
Discussion
It was previously reported that high levels of CXCR4 are expressed in lung adenocarcinoma (14, 15). In addition, high levels of CXCR4 have been closely related to lymph node metastasis, a high T stage, and patient prognosis (16). In the present study, our data demonstrate that CXCR4 expression is associated with lymph node metastasis of lung adenocarcinoma in the cohort examined. In addition, hsa_circRNA_0056616 was found to be present at higher levels in PC9 lung adenocarcinoma cells and their exosomes following the silencing of CXCR4 with shRNA. Detection of hsa_circRNA_0056616 in tumor tissues and plasma also distinguished lung adenocarcinoma patients with or without lymph node metastasis (cutoff value, 0.394). Thus, plasma levels of exo-hsa_circRNA_0056616 may represent a valuable biomarker for theranostics of lymph node metastasis lung adenocarcinoma.
The initial observation in the present study that CXCR4 was highly expressed in carcinoma tissues of lung adenocarcinoma metastasis patients is consistent with the hypothesis that CXCR4 plays an important role in many solid tumor metastases (17). For example, increased expression of CXCR4 has been associated with accelerated tumor progression and tumor cell proliferation (18). Similarly, our data demonstrate that silencing CXCR4 expression in PC9 and PC14 cell lines could effectively abolish cell proliferation. These results are also consistent with previous results (19, 20). It has also been reported that interactions between CXCR4 and SDF-1 lead to activation of G protein-mediated signaling pathways and downregulation of RAS and PI3 kinase to affect cell motility (21, 22). Furthermore, CXCR4/SDF-1 complexes have been shown to activate protein kinase B, mitogen-activated protein kinases, and nuclear factor kappa B in combination with phosphatidylinositol 3-kinases or Src family kinases and other downstream signaling proteins to promote the expression of heparanase and metalloproteinase-9 and degradation of extracellular matrix (23). As a result, tumor cells can gain access to adjacent tissues as part of tumor invasion and metastasis processes (23). In the present study, our data showed that CXCR4 silencing regulates the hallmarks of the metastasis process by inhibiting colony formation, cell proliferation, migration and invasive in PC9 and PC14 cell lines.
Currently, increasing evidence has revealed that cancer-related microRNAs (miRNAs) and regulation of circRNA-miRNA interactions in cancer signaling pathways are active. It has been demonstrated that circRNAs can act as stable endogenous RNAs, competitive endogenous RNAs, or miRNA sponges to regulate the expression of their downstream gene (24). Similarly, when circRNAs contain multiple tandem binding sites for miRNAs they can regulate miRNA activity by acting like an miRNA sponge (25). For example, the circRNA, CiRS-7, acts as a sponge of miR-7 since it contains more than 70 miR-7 conserved binding sites, and this allows CiRS-7 to negatively regulate miR-7 (26). Moreover, Chou (27) reported that overexpression of miR-7 is associated with poor prognosis in lung cancer, while inhibition of miR-7 expression may decrease cancer cell proliferation and increase apoptosis (28). CircRNAs that contain multiple MREs are more effective than traditional miRNA inhibitors, thereby providing an alternative to linear sponges. In malignant melanoma cell lines, a circRNA that acted similar to an miRNA sponge produced a longer lasting effect that resulted in a significant anti-cancer effect (29). CircRNAs have a closed loop structure that makes them less accessible to nucleases and more stable than linear RNAs. This characteristic has obvious advantages for the development and application of new markers. Thus, circRNAs may be the best miRNA inhibitors identified to date, and it is anticipated that they will be valuable targets in the diagnosis and treatment of tumors (30, 31).
To date, the potential for circRNAs to serve as tumor biomarkers has been evaluated in tumor tissues and cell lines (32-35). It remains for the clinical potential of circRNAs to be established. However, it has been observed by Li et al. (12) that the expression of circRNAs in exosomes was higher than in normal cells. Levels of circRNAs in serum exosomes of colon cancer patients were higher than in healthy individuals, and the expression levels in the exosomes correlated with the expression levels in cells (R = 0.43) [15]. The results of the present study regarding the detection of circRNAs in exosomes isolated from the peripheral blood of patients with lung adenocarcinoma are consistent with these results. In particular, exo-hsa_circRNA_0056616 was identified as a potential biomarker for lymph node metastasis. Furthermore, expression of exo-hsa_circRNA_0056616 was found to correlate with T, N, and M stages, as well as TNM grade, in patients with lung adenocarcinoma. With an area under the curve value of 0.812, this biomarker exhibited high accuracy and reliability for detecting lung adenocarcinoma lymph node metastasis versus non-metastatic lung adenocarcinoma, and this is an encouraging finding that needs to be confirmed in future studies.
There were limitations associated with the present study. First, due to the focus of this study on metastases, the PC-9 cell line was chosen. However, additional studies are needed to address whether the results obtained with PC-9 cells can be confirmed in other cell lines. Second, circRNAs contain multiple MREs that are more effective than traditional miRNA inhibitors and they can potentially serve as an alternative to linear sponges. A correlation between circRNA expression and mRNA expression was analyzed in relation to CXCR4, although, the miRNA and mRNA targets of hsa_circRNA_0056616 were not validated. Moreover, although this study quantitatively analyzed the expression of exo-hsa_circRNA_0056616 in plasma samples obtained from lung adenocarcinoma patients, a positioning analysis was not conducted.
Conclusions
In summary, the present study demonstrated that CXCR4 was upregulated and associated with lymph node metastasis, and exo-hsa_circRNA_0056616 in plasma to serve as a potential biomarker for theragnostic of lymph node metastasis in lung adenocarcinoma. However, it remains for these results to be validated in a larger number of samples, and further studies are warranted to investigate the molecular details that are responsible for the regulation and actions of hsa_circRNA_0056616. Nevertheless, the present findings are of value and they advance our understanding of lung adenocarcinoma metastasis.
Abbreviations
circRNAs: circular RNAs; exo-hsa-circRNAs: has-circRNAs in exosomes; SDS-PAGE: sodium dodecyl sulfate-polyacrylamide gel electrophoresis; PVDF: polyvinylidene difluoride; GAPDH: glyceraldehyde 3-phosphate dehydrogenase; shRNAs: short hairpin RNAs; ROC: receiver operating characteristics; AUC: area under curve; MREs: miRNA response elements.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
We thank all the staffs from Department of Thoracic Surgery, The first affiliated hospital of Fujian Medical University. And we also would like to express our appreciation to the patients participated in our study.
Ethics approval and consent to participate
This study was approved by the Institutional Review Board of Fujian Medical University (Fuzhou, China) and all participants signed informed consent forms.
Availability of data and material
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Funding
Financial support was received in the form of a grant from the National Key R&D Program of China (No.2017YFC0907100), National Natural Science Foundation of China (No.81402738), Fujian Program for Outstanding Young Researchers in University awarded by Education Department of Fujian (No. 2017B019) and Fujian Provincial Natural Science Foundation Project (No.2016J01355).
Author Contributions
HF and ZXJ carried out the experiments, participated in the drafted the manuscript. LZ, LJB and QML carried out experiments and collected samples. HF, ZXJ and HZJ participated in the design of the study and performed the statistical analysis. HZJ and LX conceived of the study, and participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. International journal of cancer. 2010;127(12):2893-917
2. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F. et al. Cancer statistics in China, 2015. Ca A Cancer Journal for Clinicians. 2016;66(2):115-32
3. Ogawa M, Watanabe M, Hasegawa T, Ichihara K, Yoshida K, Yanaga K. Expression of CXCR-4 and IDO in human colorectal cancer: An immunohistochemical approach. Molecular and Clinical Oncology. 2017;6(5):701-4
4. Huang WS, Hsieh MC, Huang CY, Kuo YH, Tung SY, Shen CH. et al. The Association of CXC Receptor 4 Mediated Signaling Pathway with Oxaliplatin-Resistant Human Colorectal Cancer Cells. PLoS One. 2016;11(9):e0159927
5. Liang Z, Brooks J, Willard M, Liang K, Yoon Y, Kang S. et al. CXCR4/CXCL12 axis promotes VEGF-mediated tumor angiogenesis through Akt signaling pathway. Biochemical & Biophysical Research Communications. 2007;359(3):716-22
6. Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biology. 2007;9(6):654
7. Cui H, Seubert B, Stahl E, Dietz H, Reuning U, Moreno-Leon L. et al. Tissue inhibitor of metalloproteinases-1 induces a pro-tumourigenic increase of miR-210 in lung adenocarcinoma cells and their exosomes. Oncogene. 2015;34(28):3640-50
8. Fujita Yu, Kosaka Nobuyoshi, Araya Jun. et al. Extracellular vesicles in lung microenvironment and pathogenesis. Trends in Molecular Medicine. 2015;21(9):533
9. Sun Y, Li L, Liu J, Wang L. Body Fluids-Derived Exosomes: Paving the Novel Road to Lung Cancer Diagnosis and Therapy. Anti-Cancer Agents in Medicinal Chemistry. 2017;17(13):1734-1740
10. Wang Y, Yi J, Chen X, Zhang Y, Xu M, Yang Z. The regulation of cancer cell migration by lung cancer cell-derived exosomes through TGF-β and IL-10. Oncology Letters. 2016;11(2):1527-30
11. Huang G, Li S, Yang N, Zou Y, Zheng D, Xiao T. Recent progress in circular RNAs in human cancers. Cancer Letter. 2017;404:8-18
12. Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Research. 2015;25(8):981
13. Wang X, Liu X, Liu C, Ren M, Gao S, Zhao G. et al. Validation of reference genes for the normalization of RT-qPCR expression studies in human tongue carcinoma cell lines and tissue. Oncology letters. 2017;13(5):3951-7
14. Otsuka S, Klimowicz AC, Kopciuk K, Petrillo SK, Konno M, Hao D. et al. CXCR4 overexpression is associated with poor outcome in females diagnosed with stage IV non-small cell lung cancer. Journal of Thoracic Oncology. 2011;6(7):1169-78
15. Spaks A. Role of CXC group chemokines in lung cancer development and progression. Journal of Thoracic Diseases. 2017;9(Suppl 3):S164-S71
16. Wagner PL, Hyjek E, Vazquez MF, Meherally D, Liu YF, Chadwick PA. et al. CXCL12 and CXCR4 in adenocarcinoma of the lung: association with metastasis and survival. Journal of Thoracic & Cardiovascular Surgery. 2009;137(3):615-21
17. Kryczek I, Wei S, Keller E, Liu R, Zou W. Stroma-derived factor (SDF-1/CXCL12) and human tumor pathogenesis. American Journal of Physiology-Cell Physiology. 2007;292(3):C987-C95
18. Franco R, Pirozzi G, Scala S, Cantile M, Scognamiglio G, Camerlingo R. et al. CXCL12-binding receptors expression in non-small cell lung cancer relates to tumoral microvascular density and CXCR4 positive circulating tumoral cells in lung draining venous blood. European journal of cardio-thoracic surgery. 2012;41(2):368-75
19. Saka B, Ekinci O, Dursun A, Akyurek N. Clinicopathologic and prognostic significance of immunohistochemical expression of HIF-1α, CXCR4 and CA9 in colorectal carcinoma. Pathology Research & Practice. 2017;213(7):783-92
20. Liu YL, Yu JM, Song XR, Wang XW, Xing LG, Gao BB. Regulation of the chemokine receptor CXCR4 and metastasis by hypoxia-inducible factor in non small cell lung cancer cell lines. Cancer Biology & Therapy. 2006;5(10):1320-6
21. Hatzistergos KE, Saur D, Seidler B, Balkan W, Breton M, Valasaki K. et al. Stimulatory Effects of MSCs on cKit+ Cardiac Stem Cells Are Mediated by SDF1/CXCR4 and SCF/cKit Signaling Pathways. Circulation Research. 2016;119(8):CIRCRESAHA.116.309281
22. Ohtsuka H, Iguchi T, Hayashi M, Kaneda M, Iida K, Shimonaka M. et al. SDF-1α/CXCR4 Signaling in Lipid Rafts Induces Platelet Aggregation via PI3 Kinase-Dependent Akt Phosphorylation. Plos One. 2017;12(1):e0169609
23. Sleightholm RL, Neilsen BK, Li J, Steele MM, Singh RK, Hollingsworth MA. et al. Emerging roles of the CXCL12/CXCR4 axis in pancreatic cancer progression and therapy. Pharmacology & Therapeutics. 2017;179:158-170
24. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA Hypothesis: The Rosetta Stone of a Hidden RNA Language? Cell. 2011;146(3):353-8
25. Ebert MS, Neilson JR, Sharp PA. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nature Methods. 2007;4(9):721
26. Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK. et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495(7441):384
27. Chou Y-T, Lin H-H, Lien Y-C, Wang Y-H, Hong C-F, Kao Y-R. et al. EGFR promotes lung tumorigenesis by activating miR-7 through a Ras/ERK/Myc pathway that targets the Ets2 transcriptional repressor ERF. Cancer research. 2010;70(21):8822 -
28. Cheng AM, Byrom MW, Shelton J, Ford LP. Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Research. 2005;33(4):1290-7
29. Tay FC, Lim JK, Zhu H, Hin LC, Wang S. Using artificial microRNA sponges to achieve microRNA loss-of-function in cancer cells. Advanced Drug Delivery Reviews. 2015;81:117-27
30. Peifei L, Shengcan C, Huilin C, Xiaoyan M, Tianwen L, Yongfu S. et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clinica Chimica Acta. 2015;444:132-6
31. Qin M, Liu G, Huo X, Tao X, Sun X, Ge Z. et al. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016;16(1):161-9
32. Wang X, Zhang Y, Huang L, Zhang J, Pan F, Li B. et al. Decreased expression of hsa_circ_001988 in colorectal cancer and its clinical significances. International Journal of Clinical & Experimental Pathology. 2015;8(12):16020
33. Shang X, Li G, Liu H, Li T, Liu J, Zhao Q. et al. Comprehensive Circular RNA Profiling Reveals That hsa_circ_0005075, a New Circular RNA Biomarker, Is Involved in Hepatocellular Crcinoma Development. Medicine. 2016;95(22):e3811
34. Xuan L, Qu L, Han Z, Peng W, Yu H, Wu T. et al. Circular RNA: a novel biomarker for progressive laryngeal cancer. American Journal of Translational Research. 2016;8(2):932
35. Nair A, Niu N, Tang X, Thompson K, Wang L, Kocher J. et al. Circular RNAs and their associations with breast cancer subtypes. Oncotarget. 2016;7(49):80967-79
Author contact
Corresponding author: Zhijian Hu, Email: hzj99955888com, TEL: +86 591 22862023, FAX number: +86 591 22862023

 Global reach, higher impact
Global reach, higher impact