Impact Factor ISSN: 1837-9664
J Cancer 2019; 10(24):5955-5963. doi:10.7150/jca.35716 This issue Cite
Research Paper
HIF-1α rs11549465 C>T polymorphism contributes to increased cancer susceptibility: Evidence from 49 studies
1. Emergency and Critical Care Center, Renmin Hospital, Hubei University of Medicine, Shiyan 442000, Hubei, China.
2. Department of Neurology, Renmin Hospital, Hubei University of Medicine, Shiyan 442000, Hubei, China.
3. Animal Experimental Management Center, Public Technology Service Platform, Shenzhen Institutes of Advanced Technology, Chinese Academy of Sciences, Shenzhen 518055, Guangdong, China.
*These authors contributed equally to this work.
Received 2019-4-13; Accepted 2019-8-20; Published 2019-10-15
Abstract
HIF-1α (hypoxia-inducible factor-1α) is a transcriptional factor that participates in the regulation of oxygen homeostasis. Despites numbers of case-control studies working on this area, the actual relationship of HIF-1α gene generic variant rs11549465 C>T imposing on cancer susceptibility remains unveiled. To get a better understanding of such relationship, this meta-analysis was carried out by incorporating all eligible case-control studies. Qualified articles were acquired from PubMed, CNKI, EMBASE, PMC, and Wanfang database update to April 2019. Odds ratios (ORs) and their corresponding 95% confidence intervals (CIs) were employed to estimate the relationship of interest. Heterogeneity tests, sensitivity analyses and publication bias assessments were also carried out to ensure the strength of our conclusion. A total of 46 articles with 49 studies including 12920 cases and 13363 controls were included. The results indicated that HIF-1α rs11549465 C>T was significantly related to the increased risk of overall cancer under four genetic models (TT vs. CC: OR=2.06, 95% CI=1.34-3.16; TT vs. CC/CT: OR=2.42, 95% CI=1.60-3.65; CT/TT vs. CC: OR=1.21, 95% CI=1.04-1.40; T vs. C: OR=1.29, 95% CI=1.12-1.48). Furthermore, enhanced cancer risk was detected after stratification by cancer type, ethnicity, the source of controls and HWE. These results suggest that HIF-1α rs11549465 C>T polymorphism may predispose to cancer susceptibility.
Keywords: HIF-1α, rs11549465 C>T, polymorphism, cancer, susceptibility
Introduction
Cancer ranks itself the leading causes of death around the world. In 2019, 1,762,450 new cancer cases and 606,880 cancer deaths are projected to occur in the United States. It has become a universal public health issue [1]. The most distinguished feature of cancer, un-controlled cell proliferation being one of them, is that it can assault the other vicinal parts of the body and diffuse to other organs. We refer this process to metastases, and this process could later evolve into a major cause of death from cancer. The exact etiology of carcinogenesis has not been fully verified [2]. More and more evidence point to genetic variation in contributing to the initiation and progression of cancer [3, 4]. However, due to cancer's complexity in nature, with heterogeneity being one of is feature, identification of this susceptibility is still a puzzle for us and most correlation has not been ascertained. On the other hand, during the decades, it has become universally agreed that single nucleotide polymorphisms (SNPs) are a common type of genetic variations that is the most frequently studied in connection with cancer susceptibility and that it consequently can act as the markers of many cancers [5].
Hypoxia possesses a vital role in the maintenance of tumor microenvironments. Hypoxic tumor microenvironment triggers extensive cellular responses, such as angiogenesis, proliferation and invasion [6]. By adjusting the oxygen pressure that results in gene alteration, hypoxia may control tumor cell phenotypes [6]. Hypoxia-inducible factor 1 (HIF-1) is a major transcriptional regulator implicated in homeostasis of oxygen. Koshiji et al. illustrated that HIF-1 leads to genetic instability by restraining the DNA mismatching repair system (MSH2 and MSH6) [7]. HIF-1 is a dimeric protein complex that possesses two components known as α and β subunits [8]. Studies have demonstrated that HIF-1α plays a vital role in activating various genes that is significantly involved with cell adhesion, erythropoiesis, angiogenesis and glucose transportation in the process of cancer development and progress [9].
Mounting evidence provided that featuring a high tumor grade, HIF-1α is over-stated in numbers of human cancers, indicating that HIF-1α functions as an independent element of cancer prognosis [10]. HIF-1α has been a research hot spot and numerous SNPs in HIF-1α were identified, whose polymorphism known as 1772 C>T (rs11549465 C>T, Pro582Ser), having been the most widely investigation polymorphism. rs11549465 C>T is a nonsynonymous SNP. Compared to the wild type, this polymorphic variant can tremendously enhance transcriptional activity in both normoxic and hypoxic environment in in-vitro studies [11]. Moreover, HIF-1α rs11549465 C>T is linked to increased tumor microvessel density which makes contribution to the cancer progression. HIF-1α rs11549465 C>T polymorphism was previously investigated in various types of cancer. Nevertheless, the conclusions obtained from previous epidemiological studies are inconsistent and contradictory. Thus, the relationship between HIF-1α rs11549465 C>T polymorphism and cancer risk requires further exploration. Herein, we performed this more comprehensive meta-analysis on selected case-control studies in the aim of giving a more thorough demonstration of the association of HIF-1α rs11549465 C>T polymorphism with cancer risk.
Materials and Methods
Publication search
We systematically searched EMBASE, PubMed, PMC, Wanfang and CNKI to retrieve relatively pertinent publications based on case-control studies (update to March 18, 2019). No language restriction is made for this analysis. The search terminology involved were as listed: 1) hypoxia-inducible factor-1 or HIF-1α or rs11549465 or 1772 C>T; 2) SNPs or polymorphisms or polymorphism or variants; 3) cancer or carcinoma or neoplasm or tumor. To acquire all qualified publications, we also reviewed the references of the selected studies.
Eligibility criteria
Impertinent and irrelevant studies were excluded on primary stage. Elimination criteria were: if 1) the study population was not mapped out; 2) it is not case-control study; 3) lack of information in allele frequency. Other than that, editorials, reviews and meta-analysis were ruled out. Only case-control studies with detailed number of different genotypes for estimating odds ratios (ORs) with 95% confidence intervals (CIs) were taken into the final analysis.
Data extraction
Two authors (Hu-Nian Li and Ting He) were arranged to extract information of all the articles respectively. Items listed below were extracted from every single study: 1) authors name; 2) publication year; 3) ethnicity of the study subject; 4) cancer type; 5) allelic frequency; 6) quality score. Studies with scores ≤9 were of low quality, whereas those with scores >9 were of high quality [12, 13]. All the disputable parts were compromised by discussion before consensus was made finally.
Statistical methods
We first performed Hardy-Weinberg equilibrium (HWE) for the controls utilizing the goodness-of-fit test. Homozygous model, heterozygous model, recessive model, dominant model, and allele model were employed to determine the relationship between HIF-1α rs11549465 C>T polymorphism and cancer risk by calculating ORs with the corresponding 95% CIs. Moreover, we conducted the stratification analysis by ethnicity, cancer type, source of control, and HWE in controls. We also used Chi square-base Q-test to gauge the presence of heterogeneity. The fixed-effect model was used to compute the pooled OR, given the studies were confirmed to be homogeneous (P>0.10 for the Q test). Or the random-effect model should be used instead. Sensitivity analysis was undertaken on the base of re-calculation of the ORs and 95% CIs by excluding each study individually. In order to detect the presence of publication bias, Begg's funnel plot and Egger's linear regression were adopted simultaneously. We also performed the trial sequential analysis (TSA) to avoid the random errors caused by repeated significance testing and dispersed data [13]. Version 11.0 STATA (Stata Corporation, College Station, TX) was selected to generate all statistical analysis. All the statistics were two-sided with P value <0.05 as a baseline significant finding.
Results
Study characteristics
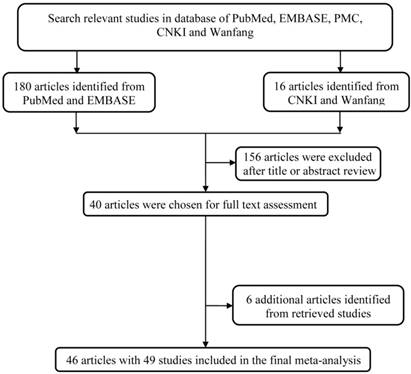
The study workflow was graphically displayed in Figure 1. We first collected 196 articles of the interest by a comprehensive search in the above-mentioned databases. After a basic check-up on articles relevance and abstracts conciseness, 156 articles were ruled out, which left us a total of 40 articles for full text assessment. To expand its sample size to ensure statistical representativeness, we identified another 6 articles from retrieve studies, quantity adding up to 46 articles in total [14-59]. Ultimately, 46 articles with 49 studies were contained in this analysis. A total of 12920 cases and 13363 controls was enrolled into this study for analyzing (Table 1).
The main flowchart of this work.
Main characteristics of included studies in the meta-analysis
| Surname | Year | Cancer type | Country | Ethnicity | Control source | Genotype method | Case | Control | HWE | Score | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CC | CT | TT | All | CC | CT | TT | All | |||||||||
| Clifford | 2001 | RCC | UK | Caucasian | PB | PCR | 30 | 5 | 0 | 35 | 110 | 27 | 6 | 143 | 0.018 | 6 |
| Tanimoto | 2003 | HNSCC | Japan | Asian | PB | PCR-Sequencing | 45 | 10 | 0 | 55 | 98 | 12 | 0 | 110 | 0.545 | 5 |
| Ollerenshaw | 2004 | RCC | UK | Caucasian | PB | PCR | 16 | 54 | 90 | 160 | 1 | 90 | 71 | 162 | <0.001 | 6 |
| Kuwai | 2004 | Colorectal cancer | Japan | Asian | PB | PCR-Sequencing | 100 | 0 | 0 | 100 | 89 | 11 | 0 | 100 | 0.561 | 7 |
| Chau | 2005 | Prostate cancer | USA | Mixed | PB | PCR | 161 | 29 | 6 | 196 | 179 | 14 | 3 | 196 | 0.002 | 6 |
| Ling | 2005 | ESCC | China | Asian | PB | PCR-RFLP | 84 | 11 | 0 | 95 | 93 | 11 | 0 | 104 | 0.569 | 6 |
| Fransen | 2006 | Colorectal cancer | Sweden | Caucasian | PB | PCR-RFLP | 167 | 28 | 3 | 198 | 213 | 43 | 2 | 258 | 0.916 | 8 |
| Konac | 2007 | Cervical cancer | Turkey | Caucasian | HB | PCR-RFLP | 10 | 14 | 8 | 32 | 68 | 37 | 2 | 107 | 0.229 | 7 |
| Konac | 2007 | Ovarian cancer | Turkey | Caucasian | HB | PCR-RFLP | 34 | 14 | 1 | 49 | 68 | 37 | 2 | 107 | 0.229 | 5 |
| Konac | 2007 | Endometrial cancer | Turkey | Caucasian | HB | PCR-RFLP | 4 | 12 | 5 | 21 | 68 | 37 | 2 | 107 | 0.229 | 5 |
| Orr-Urtreger | 2007 | Prostate cancer | Israel | Caucasian | PB | PCR-RFLP | 287 | 99 | 16 | 402 | 217 | 80 | 3 | 300 | 0.137 | 10 |
| Li | 2007 | Prostate cancer | USA | Mixed | PB | PCR-RFLP | 818 | 209 | 14 | 1041 | 175 | 13 | 0 | 188 | 0.623 | 10 |
| Horre´e | 2008 | Endometrial cancer | Netherlands | Caucasian | PB | PCR | 50 | 5 | 3 | 58 | 463 | 84 | 12 | 559 | 0.001 | 10 |
| Apaydin | 2008 | Breast cancer | Turkey | Caucasian | PB | PCR-RFLP | 79 | 21 | 2 | 102 | 68 | 29 | 5 | 102 | 0.415 | 6 |
| Jacobs | 2008 | Prostate cancer | USA | Mixed | HB | MassARRAY | 1156 | 252 | 12 | 1420 | 1138 | 284 | 28 | 1450 | 0.040 | 11 |
| Kim | 2008 | Breast cancer | Korea | Asian | HB | PCR-Sequencing | 81 | 8 | 1 | 90 | 93 | 9 | 0 | 102 | 0.641 | 9 |
| Lee | 2008 | Breast cancer | Korea | Asian | PB | SNP-ITTM | 1207 | 119 | 6 | 1332 | 1245 | 123 | 1 | 1369 | 0.250 | 11 |
| Nadaoka | 2008 | Bladder cancer | Japan | Asian | PB | PCR-RFLP | 197 | 21 | 1 | 219 | 419 | 42 | 0 | 461 | 0.350 | 10 |
| Chen | 2009 | Oral cancer | China | Asian | PB | PCR-RFLP | 163 | 10 | 1 | 174 | 334 | 13 | 0 | 347 | 0.722 | 9 |
| Li | 2009 | Gastric cancer | China | Asian | PB | PCR-LDR | 83 | 4 | 0 | 87 | 93 | 13 | 0 | 106 | 0.501 | 6 |
| Naidu | 2009 | Breast cancer | Malaysia | Asian | PB | PCR-RFLP | 294 | 100 | 16 | 410 | 222 | 50 | 3 | 275 | 0.922 | 10 |
| Foley | 2009 | Prostate cancer | Ireland | Caucasian | PB | PCR-Sequencing | 65 | 30 | 0 | 95 | 175 | 13 | 0 | 188 | 0.623 | 9 |
| Muñoz-Guerra | 2009 | Oral cancer | Spain | Caucasian | PB | PCR | 57 | 6 | 7 | 70 | 113 | 27 | 8 | 148 | 0.001 | 7 |
| Morris | 2009 | RCC | UK | Caucasian | PB | Taqman | 290 | 39 | 3 | 332 | 262 | 46 | 5 | 313 | 0.080 | 10 |
| Konac | 2009 | Lung cancer | Turkey | Caucasian | HB | PCR-RFLP | 110 | 31 | 0 | 141 | 111 | 43 | 2 | 156 | 0.335 | 8 |
| Shieh | 2010 | OSCC | China | Asian | HB | PCR-Sequencing | 282 | 23 | 0 | 305 | 89 | 7 | 0 | 96 | 0.710 | 8 |
| Shieh | 2010 | Oral cancer | China | Asian | HB | PCR | 187 | 12 | 0 | 199 | 89 | 7 | 0 | 96 | 0.710 | 8 |
| Chai | 2010 | Cervical cancer | China | Asian | HB | PCR | 65 | 25 | 7 | 97 | 94 | 21 | 2 | 117 | 0.520 | 8 |
| Hsiao | 2010 | HCC | China | Asian | HB | PCR-RFLP | 94 | 8 | 0 | 102 | 334 | 13 | 0 | 347 | 0.722 | 9 |
| Kim | 2011 | Cervical cancer | Korea | Asian | HB | SNaPShot | 177 | 22 | 0 | 199 | 187 | 27 | 0 | 214 | 0.325 | 9 |
| Putra | 2011 | Lung cancer | Japan | Asian | HB | PCR-Sequencing | 74 | 9 | 0 | 83 | 98 | 12 | 0 | 110 | 0.545 | 9 |
| Wang | 2011 | Pancreatic cancer | China | Asian | HB | PCR-Sequencing | 209 | 54 | 0 | 263 | 242 | 29 | 0 | 271 | 0.352 | 10 |
| Xu | 2011 | Glioma cancer | China | Asian | HB | PCR-RFLP | 121 | 27 | 2 | 150 | 135 | 14 | 1 | 150 | 0.354 | 8 |
| Li | 2012 | Prostate cancer | China | Asian | HB | Taqman | 612 | 48 | 2 | 662 | 659 | 57 | 0 | 716 | 0.267 | 10 |
| Ruiz-Tovar | 2012 | Pancreatic cancer | Spain | Caucasian | PB | PCR | 47 | 1 | 11 | 59 | 116 | 28 | 8 | 152 | 0.0016 | 9 |
| Kuo | 2012 | Lung cancer | China | Asian | HB | PCR-RFLP | 153 | 94 | 38 | 285 | 216 | 73 | 11 | 300 | 0.132 | 10 |
| Alves | 2012 | Oral cancer | Brazil | Mixed | PB | PCR | 0 | 1 | 39 | 40 | 0 | 85 | 3 | 88 | <0.001 | 9 |
| Zagouri | 2012 | Breast cancer | Greece | Caucasian | HB | PCR-RFLP | 98 | 15 | 0 | 113 | 107 | 17 | 0 | 124 | 0.413 | 5 |
| Qin | 2012 | RCC | China | Asian | HB | Taqman | 572 | 46 | 2 | 620 | 578 | 43 | 2 | 623 | 0.220 | 10 |
| Rebeiro | 2013 | Breast cancer | Portugal | Caucasian | PB | PCR-RFLP | 74 | 21 | 1 | 96 | 61 | 7 | 4 | 72 | 0.001 | 8 |
| Mera-Menendez | 2013 | Glottic cancer | Spain | Caucasian | HB | PCR | 85 | 18 | 15 | 118 | 114 | 27 | 8 | 149 | 0.001 | 10 |
| Fu | 2014 | Cervical cancer | China | Asian | HB | PCR | 467 | 49 | 2 | 518 | 492 | 60 | 1 | 553 | 0.550 | 11 |
| Fraga | 2014 | Prostate cancer | Portugal | Caucasian | HB | Taqman | 566 | 156 | 14 | 736 | 579 | 164 | 11 | 754 | 0.400 | 11 |
| Liu | 2014 | HCC | China | Asian | HB | PCR-RFLP | 152 | 4 | 1 | 157 | 162 | 11 | 0 | 173 | 0.6658 | 9 |
| Ni | 2015 | Digestive tract cancers | China | Asian | HB | PCR-RFLP | 219 | 44 | 4 | 267 | 241 | 34 | 0 | 275 | 0.2745 | 10 |
| Meka | 2015 | Breast cancer | India | Asian | HB | PCR | 245 | 94 | 9 | 348 | 229 | 89 | 2 | 320 | 0.0322 | 10 |
| Yamamoto | 2016 | Lung cancer | Japan | Asian | HB | TaqMan | 405 | 55 | 2 | 462 | 341 | 37 | 1 | 379 | 0.9972 | 10 |
| Demirel | 2017 | Colorectal cancer | Turkey | Caucasian | HB | ARMS-PCR | 62 | 27 | 3 | 92 | 81 | 16 | 4 | 101 | 0.0144 | 8 |
| Uslu | 2018 | Laryngeal Cancer | Turkey | Caucasian | HB | PCR | 28 | 7 | 0 | 35 | 28 | 7 | 0 | 35 | 0.5109 | 5 |
HWE, Hardy-Weinberg equilibrium; PB, population based; HB, hospital based; RCC, renal cell carcinoma; HNSCC, head and neck squamous cell carcinoma; ESCC, esophageal squamous cell carcinoma; OSCC, oral squamous cell carcinoma; HCC, hepatocellular cancer; PCR-RFLP, polymerase chain reaction-restriction fragment length polymorphism.
Quantitative analysis
The quantitative results of the meta-analysis were displayed in Table 2 and Figure 2. The results concluded that the rs11549465 C>T polymorphism was significantly related to the increased risk of overall cancer under homozygous model (TT vs. CC: OR=2.06, 95% CI=1.34-3.16), recessive model (TT vs. CC/CT: OR=2.42, 95% CI=1.60-3.65); dominant model (CT/TT vs. CC: OR=1.21, 95% CI=1.04-1.40), and allele model (T vs. C: OR=1.29,95% CI =1.12-1.48). We failed to detect any distinguished relationship between rs11549465 C>T and renal cell carcinoma (RCC), endometrial cancer, colorectal cancer, lung cancer, breast cancer, hepatocellular cancer (HCC) under all the five genetic models. However, we observed that the rs11549465 C>T polymorphism could confer to increased risk in subgroups of prostate cancer (CT vs. CC/CT: OR=1.51, 95% CI=1.01-2.26; CT/TT vs. CC: OR=1.56, 95% CI=1.04-2.34; T vs. C: OR=1.54, 95% CI =1.05-2.25), cervical cancer (TT vs. CC: OR=7.63, 95% CI=1.83-31.8; TT vs. CC/CT: OR=6.60, 95% CI=2.07-21.0), oral cancer (TT vs. CC: OR=2.61, 95% CI=1.19-5.72; TT vs. CC/CT: OR=13.2, 95% CI=1.08-162), pancreatic cancer (TT vs. CC: OR=3.39, 95% CI=1.28-8.97; TT vs. CC/CT: OR=2.42, 95% CI=1.60-3.65) and other cancers (TT vs. CC: OR=2.62, 95% CI=1.24-5.55; TT vs. CC/CT: OR=2.64, 95% CI=1.26-5.56; T vs. C: OR=1.28, 95% CI=1.00-1.62).
Meta-analysis of HIF-1α rs11549465 C>T polymorphism and cancer risk
| Variables | Homozygous | Heterozygous | Recessive | Dominant | Allele | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TT vs. CC | CT vs. CC | TT vs. CC/CT | CT/TT vs. CC | T vs. C | ||||||||||
| OR (95% CI) | P het | OR (95% CI) | P het | OR (95% CI) | P het | OR (95% CI) | P het | OR (95% CI) | P het | |||||
| All | 2.06 (1.34-3.16) | <0.001 | 1.14 (0.99-1.33) | <0.001 | 2.42 (1.60-3.65) | <0.001 | 1.21 (1.04-1.40) | <0.001 | 1.29 (1.12-1.48) | <0.001 | ||||
| Cancer type | ||||||||||||||
| RCC | 0.37 (0.12-1.12) | 0.282 | 0.64 (0.32-1.29) | 0.012 | 1.31 (0.77-2.24) | 0.350 | 0.66 (0.35-1.23) | 0.024 | 0.92 (0.70-1.19) | 0.252 | ||||
| Colorectal | 1.30 (0.40-4.17) | 0.579 | 0.83 (0.24-2.83) | 0.005 | 1.18 (0.37-3.78) | 0.465 | 0.86 (0.29-2.60) | 0.008 | 0.92 (0.37-2.26) | 0.019 | ||||
| Prostate | 1.67 (0.66-4.19) | 0.008 | 1.51 (1.01-2.26) | <0.001 | 1.62 (0.66-3.99) | 0.011 | 1.56 (1.04-2.34) | <0.001 | 1.54 (1.05-2.25) | <0.001 | ||||
| Cervical | 7.63 (1.83-31.8) | 0.170 | 1.22 (0.76-1.96) | 0.064 | 6.60 (2.07-21.0) | 0.289 | 1.46 (0.78-2.72) | 0.004 | 1.55 (0.80-3.02) | <0.001 | ||||
| Endometrial | 9.06 (0.53-156.2) | 0.014 | 1.69 (0.18-16.2) | 0.003 | 5.85 (0.93-36.9) | 0.086 | 2.29 (0.25-21.1) | 0.001 | 2.12 (0.46-9.78) | 0.002 | ||||
| Breast | 1.38 (0.33-5.74) | 0.045 | 0.99 (0.80-1.23) | 0.329 | 1.38 (0.33-5.75) | 0.044 | 1.02 (0.85-1.22) | 0.458 | 1.04 (0.88-1.23) | 0.434 | ||||
| Oral | 2.61 (1.19-5.72) | 0.514 | 1.06 (0.61-1.85) | 0.081 | 13.2 (1.08-162) | <0.001 | 1.24 (0.79-1.93) | 0.149 | 1.90 (0.88-4.07) | <0.001 | ||||
| Lung | 1.92 (0.35-10.5) | 0.103 | 1.19 (0.78-1.82) | 0.044 | 1.93 (0.43-8.66) | 0.154 | 1.23 (0.71-2.13) | 0.002 | 1.23 (0.69-2.20) | <0.001 | ||||
| HCC | 3.20 (0.13-79.1) | - | 0.96 (0.17-5.29) | 0.021 | 3.33 (0.14-82.2) | - | 1.06 (0.24-4.68) | 0.035 | 1.15 (0.33-4.06) | 0.061 | ||||
| Pancreatic | 3.39 (1.28-8.97) | - | 0.50 (0.02-14.0) | 0.001 | 2.42 (1.60-3.65) | - | 1.39 (0.54-3.56) | 0.032 | 1.75 (1.23-2.51) | 0.349 | ||||
| Others | 2.62 (1.24-5.55) | 0.784 | 1.13 (0.87-1.47) | 0.275 | 2.64 (1.26-5.56) | 0.810 | 1.22 (0.95-1.57) | 0.274 | 1.28 (1.00-1.62) | 0.239 | ||||
| Ethnicity | ||||||||||||||
| Caucasian | 1.54 (0.81-2.87) | <0.001 | 1.01 (0.75-1.35) | <0.001 | 1.82 (1.15-2.89) | 0.004 | 1.10 (0.84-1.44) | <0.001 | 1.21 (0.97-1.51) | <0.001 | ||||
| Asian | 4.07 (2.61-6.34) | 0.995 | 1.19 (1.02-1.38) | 0.010 | 3.67 (2.37-5.72) | 0.997 | 1.25 (1.06-1.47) | 0.001 | 1.28 (1.09-1.51) | <0.001 | ||||
| Mixed | 1.27 (0.26-6.15) | 0.028 | 1.85 (0.70-4.86) | <0.001 | 7.57 (0.31-184) | <0.001 | 1.86 (0.67-5.16) | <0.001 | 3.24 (1.02-10.3) | <0.001 | ||||
| Source of control | ||||||||||||||
| PB | 1.61 (0.90-2.89) | 0.014 | 1.03 (0.76-1.40) | <0.001 | 2.51 (1.33-4.74) | <0.001 | 1.12 (0.85-1.47) | <0.001 | 1.27 (0.99-1.62) | <0.001 | ||||
| HB | 2.61 (1.39-4.91) | <0.001 | 1.17 (1.00-1.36) | 0.001 | 2.36 (1.33-4.18) | <0.001 | 1.25 (1.05-1.48) | <0.001 | 1.30 (1.09-1.55) | <0.001 | ||||
| HWE | ||||||||||||||
| >0.05 | 2.92 (1.34-3.16) | 0.015 | 1.20 (1.02-1.41) | <0.001 | 2.71 (1.76-4.16) | 0.111 | 1.26 (1.06-1.50) | <0.001 | 1.30 (1.10-1.54) | <0.001 | ||||
| ≤0.05 | 1.18 (0.59-2.36) | <0.001 | 0.91 (0.62-1.33) | <0.001 | 2.10 (0.99-4.44) | <0.001 | 1.04 (0.78-1.38) | 0.002 | 1.24 (0.95-1.63) | <0.001 | ||||
| Score | ||||||||||||||
| >9 | 2.26 (1.32-3.85) | 0.001 | 1.13 (0.97-1.32) | <0.001 | 2.19 (1.32-3.63) | 0.004 | 1.21 (1.02-1.43) | <0.001 | 1.25 (1.05-1.49) | <0.001 | ||||
| ≤9 | 1.76 (0.84-3.67) | <0.001 | 1.10 (0.83-1.47) | <0.001 | 2.59 (1.31-5.14) | <0.001 | 1.18 (0.90-1.54) | <0.001 | 1.31 (1.03-1.67) | <0.001 | ||||
Het, heterogeneity; RCC, renal cell carcinoma; HB, hospital based; PB, population based.
Forest plot for the correlation between the HIF-1α rs11549465 C>T polymorphism and cancer susceptibility under the allele comparison model. The horizontal lines represent the study-specific ORs and 95% CIs. The diamond represents the pooled results of OR and 95% CI.
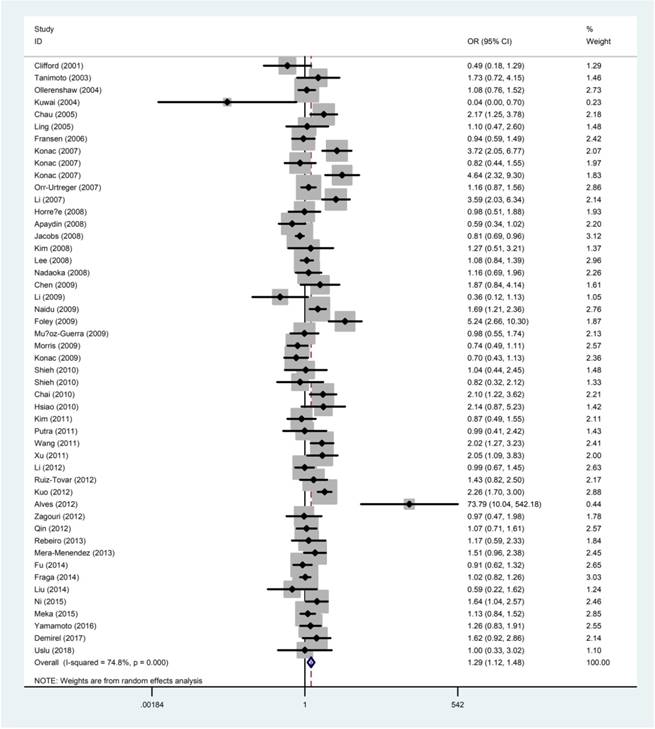
When it comes to the stratification analysis by the ethnicity, significant increased risk was detected in Asians, Caucasians and mixed population. In terms of source of controls, either population-based controls or hospital-based controls were associated with the increase risk of cancer. Further subgroup analysis by HWE in controls revealed that no significant correlation was observed in subgroup of HWE≤0.05. As regard to the quality of publications, significant increased risk was detected in high-quality and low-quality publications.
Heterogeneity and sensitivity analysis
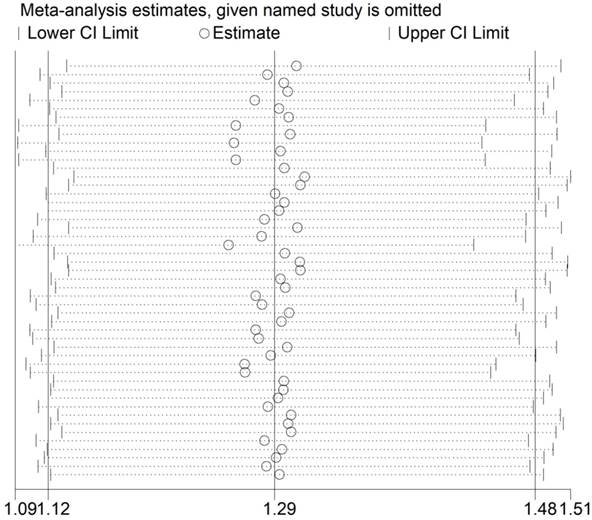
The Q test (P<0.001) implied an existence of heterogeneity under all the genetic models. Thus, we adopted a random-effect model to produce ORs and 95% CIs. In addition, the sequential sensitivity analysis was performed to give an evaluation of the impact of a sole study on the pooled estimation. Given the attempt of omitting in each study incurred no statistical fluctuation of the pooled ORs, we have reason to believe that the meta-analysis's data is of great reliability (Figure 3).
Sensitivity analysis of the association between HIF-1α rs11549465 C>T and cancer susceptibility. Each point represents the recalculated OR after deleting a separate study.
Publication bias
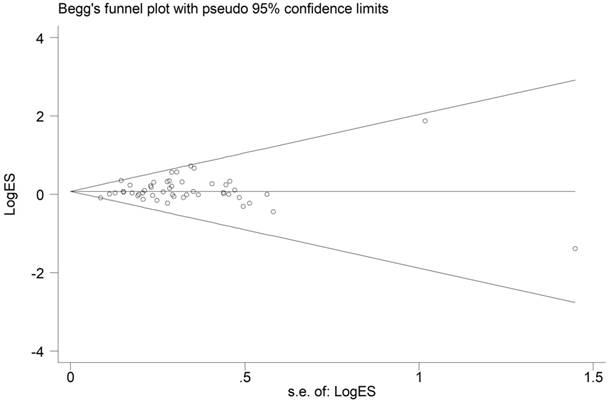
From the shape of the Begg's funnel plot shown in Figure 4, no evidence of asymmetry was found. Egger's test's statistics also gives no evidence of publication bias among the studies.
Funnel plot analysis to detect publication bias for HIF-1α rs11549465 C>T polymorphism under the allele comparison model. Each point represents a separate study.
Trial sequential analysis (TSA)
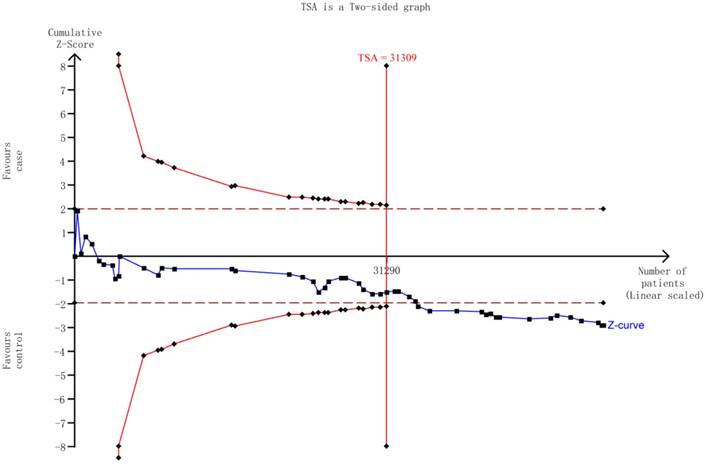
The TSA showed that the cumulative z-curve did not cross both the traditional threshold and the TSA threshold, yet the accumulated information was sufficient, indicating that no further evidence was needed to verify the conclusion (Figure 5).
The required information size to demonstrate the correlation between HIF-1α rs11549465 C>T polymorphism with cancer susceptibility. The solid blue line is the cumulative Z-curve. The dashed inward-sloping line to the left represents the trial sequential monitoring boundaries.
Discussion
In the current meta-analysis, we systematically evaluate the relationship between HIF-1α rs11549465 C>T polymorphism and cancer risk by using 49 case-control studies. Our analysis showed that HIF-1α rs11549465 C>T polymorphism could increase risk of overall cancer risk and specific cancer risk. Among all the epidemical studies on the rs11549465 C>T polymorphism and cancer risk, this could be by now the most comprehensive one.
The HIF-1α gene is located at chromosome 14q21-24. HIF-1α regulates the expression of hundreds of genes which moderates the vital cellular functions like proliferation, apoptosis, angiogenesis, glucose metabolism, erythropoiesis, and iron metabolism [60]. Due to the complex functional mechanism and regulatory roles of HIF-1a in hypoxic stress, the possible role of HIF-1a gene SNPs in cancer susceptibility has evoked intensive investigation. The most broadly studied HIF-1α polymorphism rs11549465 C>T (Pro582Ser) could induce proline-to-serine amino acid substitutions. However, the exact role of rs11549465 C>T polymorphism in cancer risk obtained from different studies remain inconclusive.
In 2001, Clifford et al. [14] carried out a first case-control study investigating the relationship between HIF-1α rs11549465 C>T and cancer risk. However, association analysis between rs11549465 C>T and RCC risk in panels of 20 cases and 44 non-neoplastic controls did not reveal allelic frequency differences. An investigation conducted by Konac et al. [21] using endometrial, ovarian, and cervical cancers in the Turkish population revealed that the rs11549465 C>T polymorphism of the HIF-1α may contribute to risk of endometrial and cervical cancers. In a meta-analysis performed by Zhao et al. [10] in 2009 using 5387 controls and 4131 cancer cases, the HIF-1α rs11549465 C>T polymorphism was reported to be related to increased cancer risk. In 2015, Li et al. [61] conducted an updated meta-analysis by enrolling 7807 cases and 8633 controls. They obtained a similar result that the HIF-1α rs11549465 C>T polymorphism predispose to higher overall cancer risk. To better illustrate the relationship of interest, we hereby conducted this updated meta-analysis by using all the qualified publications with a total of 12920 cases and 13363 controls. The results revealed that HIF-1α rs11549465 C>T polymorphism contributes to increased overall cancer risk. In a sense, this meta-analysis has succeeded in giving a clearer clue of the relationship between HIF-1α rs11549465 C>T polymorphism and cancer risk.
In the current meta-analysis, we undertaken many measurements to increase the credibility of our conclusion. First and foremost, we included as many as qualified studies to expand the analyzed sample size, by incorporating studies not only pressed in English but also in Chinese. Second, we adopted the sensitivity analysis and the publication bias. However, several limitations could not be settled down. First, between-study heterogeneity exists, which might weaken the persuasiveness of the conclusion. Second, the relationship strength was only assessed by use of unadjusted estimates. Lacking original data, such as environment factor, adjustment analysis was absent. Third, most of the included studies were conducted among Asians and Caucasians. The lack of other ethnicities, such as Africans, compromised the generalization of the conclusion.
In a word, our finding has come to a fruition that HIF-1α rs11549465 C>T polymorphism was significantly related to an increase in cancer risk. Our work no doubt will encourage more dedication into further elucidation of the etiology of cancer predisposition. However, with limited sample size of subgroup analysis, we must admit that this analysis is imperfect and thus in the future more case-control studies should be conducted with a larger size of samples.
Abbreviations
HIF-1: hypoxia-inducible factor 1; ORs: odds ratios; Cis: confidence intervals; HWE: Hardy-Weinberg equilibrium; TSA: trial sequential analysis.
Acknowledgements
This study was supported by grant from Hubei Provincial Microcirculation Society Personnel Training Special Fund Project [No: HBWXH2018(1)-4].
Author Contributions
H.L. and W.Y. conceived and designed the study; H.L. and T.H. collected articles and extracted information; H.L., Y.Z., F.D. and J.L. analyzed the data and prepared the tables and figures; H.L., T.H. and W.Y. wrote the manuscript. All authors read and approved the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67:7-30
2. Theodoratou E, Timofeeva M, Li X, Meng X, Ioannidis JPA. Nature, Nurture, and Cancer Risks: Genetic and Nutritional Contributions to Cancer. Annu Rev Nutr. 2017;37:293-320
3. Mroz EA, Rocco JW. The challenges of tumor genetic diversity. Cancer. 2017;123:917-27
4. Malhotra J. Molecular and genetic epidemiology of cancer in low- and medium-income countries. Ann Glob Health. 2014;80:418-25
5. Erichsen HC, Chanock SJ. SNPs in cancer research and treatment. Br J Cancer. 2004;90:747-51
6. Hill RP, Marie-Egyptienne DT, Hedley DW. Cancer stem cells, hypoxia and metastasis. Semin Radiat Oncol. 2009;19:106-11
7. Koshiji M, To KK, Hammer S, Kumamoto K, Harris AL, Modrich P. et al. HIF-1alpha induces genetic instability by transcriptionally downregulating MutSalpha expression. Mol Cell. 2005;17:793-803
8. Schmid T, Zhou J, Brune B. HIF-1 and p53: communication of transcription factors under hypoxia. J Cell Mol Med. 2004;8:423-31
9. Tsai YP, Wu KJ. Hypoxia-regulated target genes implicated in tumor metastasis. J Biomed Sci. 2012;19:102
10. Zhao T, Lv J, Zhao J, Nzekebaloudou M. Hypoxia-inducible factor-1alpha gene polymorphisms and cancer risk: a meta-analysis. J Exp Clin Cancer Res. 2009;28:159
11. Li D, Liu J, Zhang W, Ren J, Yan L, Liu H. et al. Association between HIF1A P582S and A588T polymorphisms and the risk of urinary cancers: a meta-analysis. PLoS One. 2013;8:e63445
12. He J, Liao XY, Zhu JH, Xue WQ, Shen GP, Huang SY. et al. Association of MTHFR C677T and A1298C polymorphisms with non-Hodgkin lymphoma susceptibility: evidence from a meta-analysis. Sci Rep. 2014;4:6159
13. Fu W, Zhuo ZJ, Chen YC, Zhu J, Zhao Z, Jia W. et al. NFKB1 -94insertion/deletion ATTG polymorphism and cancer risk: Evidence from 50 case-control studies. Oncotarget. 2017;8:9806-22
14. Clifford SC, Astuti D, Hooper L, Maxwell PH, Ratcliffe PJ, Maher ER. The pVHL-associated SCF ubiquitin ligase complex: molecular genetic analysis of elongin B and C, Rbx1 and HIF-1alpha in renal cell carcinoma. Oncogene. 2001;20:5067-74
15. Tanimoto K, Yoshiga K, Eguchi H, Kaneyasu M, Ukon K, Kumazaki T. et al. Hypoxia-inducible factor-1alpha polymorphisms associated with enhanced transactivation capacity, implying clinical significance. Carcinogenesis. 2003;24:1779-83
16. Kuwai T, Kitadai Y, Tanaka S, Kuroda T, Ochiumi T, Matsumura S. et al. Single nucleotide polymorphism in the hypoxia-inducible factor-1alpha gene in colorectal carcinoma. Oncol Rep. 2004;12:1033-7
17. Ollerenshaw M, Page T, Hammonds J, Demaine A. Polymorphisms in the hypoxia inducible factor-1alpha gene (HIF1A) are associated with the renal cell carcinoma phenotype. Cancer Genet Cytogenet. 2004;153:122-6
18. Chau CH, Permenter MG, Steinberg SM, Retter AS, Dahut WL, Price DK. et al. Polymorphism in the hypoxia-inducible factor 1alpha gene may confer susceptibility to androgen-independent prostate cancer. Cancer Biol Ther. 2005;4:1222-5
19. Ling TS, Shi RH, Zhang GX, Zhu H, Yu LZ, Ding XF. Common single nucleotide polymorphism of hypoxia-inducible factor-1alpha and its impact on the clinicopathological features of esophageal squamous cell carcinoma. Chin J Dig Dis. 2005;6:155-8
20. Fransen K, Fenech M, Fredrikson M, Dabrosin C, Soderkvist P. Association between ulcerative growth and hypoxia inducible factor-1alpha polymorphisms in colorectal cancer patients. Mol Carcinog. 2006;45:833-40
21. Konac E, Onen HI, Metindir J, Alp E, Biri AA, Ekmekci A. An investigation of relationships between hypoxia-inducible factor-1 alpha gene polymorphisms and ovarian, cervical and endometrial cancers. Cancer Detect Prev. 2007;31:102-9
22. Li H, Bubley GJ, Balk SP, Gaziano JM, Pollak M, Stampfer MJ. et al. Hypoxia-inducible factor-1alpha (HIF-1alpha) gene polymorphisms, circulating insulin-like growth factor binding protein (IGFBP)-3 levels and prostate cancer. Prostate. 2007;67:1354-61
23. Orr-Urtreger A, Bar-Shira A, Matzkin H, Mabjeesh NJ. The homozygous P582S mutation in the oxygen-dependent degradation domain of HIF-1 alpha is associated with increased risk for prostate cancer. Prostate. 2007;67:8-13
24. Apaydin I, Konac E, Onen HI, Akbaba M, Tekin E, Ekmekci A. Single nucleotide polymorphisms in the hypoxia-inducible factor-1alpha (HIF-1alpha) gene in human sporadic breast cancer. Arch Med Res. 2008;39:338-45
25. Horree N, Groot AJ, van Hattem WA, Heintz AP, Vooijs M, van Diest PJ. HIF-1A gene mutations associated with higher microvessel density in endometrial carcinomas. Histopathology. 2008;52:637-9
26. Jacobs EJ, Hsing AW, Bain EB, Stevens VL, Wang Y, Chen J. et al. Polymorphisms in angiogenesis-related genes and prostate cancer. Cancer Epidemiol Biomarkers Prev. 2008;17:972-7
27. Kim HO, Jo YH, Lee J, Lee SS, Yoon KS. The C1772T genetic polymorphism in human HIF-1alpha gene associates with expression of HIF-1alpha protein in breast cancer. Oncol Rep. 2008;20:1181-7
28. Lee JY, Choi JY, Lee KM, Park SK, Han SH, Noh DY. et al. Rare variant of hypoxia-inducible factor-1alpha (HIF-1A) and breast cancer risk in Korean women. Clin Chim Acta. 2008;389:167-70
29. Nadaoka J, Horikawa Y, Saito M, Kumazawa T, Inoue T, Narita S. et al. Prognostic significance of HIF-1 alpha polymorphisms in transitional cell carcinoma of the bladder. Int J Cancer. 2008;122:1297-302
30. Chen MK, Chiou HL, Su SC, Chung TT, Tseng HC, Tsai HT. et al. The association between hypoxia inducible factor-1alpha gene polymorphisms and increased susceptibility to oral cancer. Oral Oncol. 2009;45:e222-6
31. Foley R, Marignol L, Thomas AZ, Cullen IM, Perry AS, Tewari P. et al. The HIF-1alpha C1772T polymorphism may be associated with susceptibility to clinically localised prostate cancer but not with elevated expression of hypoxic biomarkers. Cancer Biol Ther. 2009;8:118-24
32. Konac E, Dogan I, Onen HI, Yurdakul AS, Ozturk C, Varol A. et al. Genetic variations in the hypoxia-inducible factor-1alpha gene and lung cancer. Exp Biol Med (Maywood). 2009;234:1109-16
33. Li K, Zhang Y, Dan Z, Wang Y, Ren ZC. Association of the hypoxia inducible factor-1alpha gene polymorphisms with gastric cancer in Tibetans. Biochem Genet. 2009;47:625-34
34. Morris MR, Hughes DJ, Tian YM, Ricketts CJ, Lau KW, Gentle D. et al. Mutation analysis of hypoxia-inducible factors HIF1A and HIF2A in renal cell carcinoma. Anticancer Res. 2009;29:4337-43
35. Munoz-Guerra MF, Fernandez-Contreras ME, Moreno AL, Martin ID, Herraez B, Gamallo C. Polymorphisms in the hypoxia inducible factor 1-alpha and the impact on the prognosis of early stages of oral cancer. Ann Surg Oncol. 2009;16:2351-8
36. Naidu R, Har YC, Taib NA. Associations between hypoxia-inducible factor-1alpha (HIF-1alpha) gene polymorphisms and risk of developing breast cancer. Neoplasma. 2009;56:441-7
37. Chai D, Chen YL, Zheng A, Liu YY, Chu YX, Han L. [Relationship between polymorphism of hypoxia inducible factor-1alpha and cervical cancer in Han population in Sichuan Province of China]. Sichuan Da Xue Xue Bao Yi Xue Ban. 2010;41:674-7
38. Hsiao PC, Chen MK, Su SC, Ueng KC, Chen YC, Hsieh YH. et al. Hypoxia inducible factor-1alpha gene polymorphism G1790A and its interaction with tobacco and alcohol consumptions increase susceptibility to hepatocellular carcinoma. J Surg Oncol. 2010;102:163-9
39. Shieh TM, Chang KW, Tu HF, Shih YH, Ko SY, Chen YC. et al. Association between the polymorphisms in exon 12 of hypoxia-inducible factor-1alpha and the clinicopathological features of oral squamous cell carcinoma. Oral Oncol. 2010;46:e47-53
40. Kang MJ, Jung SA, Jung JM, Kim SE, Jung HK, Kim TH. et al. Associations between single nucleotide polymorphisms of MMP2, VEGF, and HIF1A genes and the risk of developing colorectal cancer. Anticancer Res. 2011;31:575-84
41. Kim YH, Park IA, Park WY, Kim JW, Kim SC, Park NH. et al. Hypoxia-inducible factor 1alpha polymorphisms and early-stage cervical cancer. Int J Gynecol Cancer. 2011;21:2-7
42. Putra AC, Tanimoto K, Arifin M, Hiyama K. Hypoxia-inducible factor-1alpha polymorphisms are associated with genetic aberrations in lung cancer. Respirology. 2011;16:796-802
43. Xu G, Wang M, Xie W, Bai X. Hypoxia-inducible factor-1 alpha C1772T gene polymorphism and glioma risk: a hospital-based case-control study from China. Genet Test Mol Biomarkers. 2011;15:461-4
44. Alves LR FC, Oliveira MVM, Sousa AA, Jorge ASB, Marques-Silva L, Santos SHS, Jones KM, de Paula AMB, Guimarães ALS. High HIF-1α expression genotypes increase odds ratio of oral cancer. Head Neck Oncol. 2012;4:87
45. Kuo WH, Shih CM, Lin CW, Cheng WE, Chen SC, Chen W. et al. Association of hypoxia inducible factor-1alpha polymorphisms with susceptibility to non-small-cell lung cancer. Transl Res. 2012;159:42-50
46. Li P, Cao Q, Shao PF, Cai HZ, Zhou H, Chen JW. et al. Genetic polymorphisms in HIF1A are associated with prostate cancer risk in a Chinese population. Asian J Androl. 2012;14:864-9
47. Qin C, Cao Q, Ju X, Wang M, Meng X, Zhu J. et al. The polymorphisms in the VHL and HIF1A genes are associated with the prognosis but not the development of renal cell carcinoma. Ann Oncol. 2012;23:981-9
48. Ruiz-Tovar J, Fernandez-Contreras ME, Martin-Perez E, Gamallo C. Association of thymidylate synthase and hypoxia inducible factor-1alpha DNA polymorphisms with pancreatic cancer. Tumori. 2012;98:364-9
49. Zagouri F, Sergentanis TN, Gazouli M, Tsigginou A, Dimitrakakis C, Papaspyrou I. et al. HSP90, HSPA8, HIF-1 alpha and HSP70-2 polymorphisms in breast cancer: a case-control study. Mol Biol Rep. 2012;39:10873-9
50. Mera-Menendez F, Hinojar-Gutierrez A, Guijarro Rojas M, de Gregorio JG, Mera-Menendez E, Sanchez JJ. et al. Polymorphisms in HIF-1alpha affect presence of lymph node metastasis and can influence tumor size in squamous-cell carcinoma of the glottic larynx. Clin Transl Oncol. 2013;15:358-63
51. Ribeiro AL, Gaspar JF, Pereira T, Ribeiro V. Lack of relevance of HIF-1alpha polymorphisms in breast cancer in a Portuguese population. Anticancer Res. 2013;33:2549-55
52. Fraga A, Ribeiro R, Principe P, Lobato C, Pina F, Mauricio J. et al. The HIF1A functional genetic polymorphism at locus +1772 associates with progression to metastatic prostate cancer and refractoriness to hormonal castration. Eur J Cancer. 2014;50:359-65
53. Fu SL, Miao J, Ding B, Wang XL, Cheng WJ, Dai HH. et al. A polymorphism in the 3' untranslated region of Hypoxia-Inducible Factor-1 alpha confers an increased risk of cervical cancer in a Chinese population. Neoplasma. 2014;61:63-9
54. Liu Y, Sui J, Zhai L, Yang S, Huang L, Huang L. et al. Genetic polymorphisms in hypoxia-inducible factor-1a gene and its association with HBV-related hepatocellular carcinoma in a Chinese population. Med Oncol. 2014;31:200
55. Meka PB, Cingeetham A, Nanchari SR, Damineni S, Tipirisetti N, Gorre M. et al. HIF-1alpha (1772C>T) polymorphism as marker for breast cancer development. Tumour Biol. 2015;36:3215-20
56. Ni Zhi-Hai LX-J, Mo Jing-Gang, Zhang Yi, Liang Jian-Hua, Yang Yu-Sha, Zhou Yong, Li Zhao-Hua, Zhang Jian-Liang, Ding Yin-Lu, Zhang Peng, Wang Jin-Qing. Associations of hypoxia inducible factor-1α gene polymorphisms with susceptibility to digestive tract cancers: a case-control study and meta-analysis. Genes & Genomics. 2015;37:931-8
57. Demirel HS TP, Cetinkaya S, Cinar I, Kucukkartallar K, Dursun G. Colorectal Cancer Risk in Relation to Hypoxia Inducible Factor-1α (Hif-1 α) and Von Hippel-Lindau (Vhl) Gene Polymorphisms. International Journal of Hematology and Oncology. 2017;27:13-20
58. Yamamoto Y, Kiyohara C, Ogata-Suetsugu S, Hamada N, Nakanishi Y. Association between genetic polymorphisms involved in the hypoxia-inducible factor pathway and lung cancer risk: a case-control study in Japan. Asia Pac J Clin Oncol. 2017;13:234-42
59. Uslu C, Tuz M, Yasan H, Okur E. Investigation of GLUT1, HIF1alpha and TBX21 Gene Polymorphisms in Laryngeal Cancer. Turk Arch Otorhinolaryngol. 2018;56:70-4
60. Balamurugan K. HIF-1 at the crossroads of hypoxia, inflammation, and cancer. Int J Cancer. 2016;138:1058-66
61. Li Y, Li C, Shi H, Lou L, Liu P. The association between the rs11549465 polymorphism in the hif-1alpha gene and cancer risk: a meta-analysis. Int J Clin Exp Med. 2015;8:1561-74
Author contact
Corresponding authors: Hui-Ran Lin, Animal Experimental Management Center, Public Technology Service Platform, Shenzhen Institutes of Advanced Technology, Chinese Academy of Sciences, Shenzhen 518055, Guangdong, China. Email: hr.linac.cn. Wen-Zi Yang, Emergency and Critical Care Center, Renmin Hospital, Hubei University of Medicine, 39 Chaoyang Middle Road, Shiyan 442000, Hubei, China. Email: yangwenzisycom.

 Global reach, higher impact
Global reach, higher impact