Impact Factor ISSN: 1837-9664
J Cancer 2019; 10(13):2969-2981. doi:10.7150/jca.31011 This issue Cite
Research Paper
Toll-like receptor 6 expression, sequence variants, and their association with colorectal cancer risk
1. Groupe de Recherche en Écologie Buccale, Département de stomatologie, Faculté de Médecine Dentaire, Université Laval, Québec, Québec, Canada.
2. Department of Biochemistry, College of Science King Saud University, Riyadh, Kingdom of Saudi Arabia
3. Zoology Department, College of Science, King Saud University, Riyadh, Kingdom of Saudi Arabia.
4. College of Medicine, King Saud University, Riyadh, Kingdom of Saudi Arabia
5. Division of Gastroenterology, King Khalid University Hospital, King Saud University, Riyadh, Saudi Arabia.
6. Integrated Gulf Biosystems (IGB), Riyadh, Kingdom of Saudi Arabia
7. Pharmacology department, College of Medicine, Imam Mohammed Ibn Saud Islamic University (IMSIU), Riyadh, Kingdom of Saudi Arabia
Received 2018-10-28; Accepted 2019-4-25; Published 2019-6-2
Abstract
This is the first study to examine the potential correlation of the rs3796508 and rs5743810 SNPs of the TLR6 gene in patients with colorectal cancer (CRC) in a subset of the Saudi population. TLR6 gene expression was studied by real-time PCR assaysin 10 matching normal and cancer colon tissues. TLR6 expression at the protein level was determined by immunohistochemistry. A case-control search was conductedon 115 case patients and 102 controls. All samples were genotyped with the TaqMan assay for the TLR6 gene. Odds ratios and 95% confidence interval were computed from logistic regression models after adjusting for age, sex, and tumor localization. Our findings showed a decrease in TLR6 expression (p <0.001) in colon cancer tissues when compared to normal colon tissues. Global analysis revealed no significant association between the TLR6 rs3796508 and rs5743810 and CRC in this population. However, the Val/Met genotype of rs3796508 had a significantly higher frequency in the control group than in the cases for the male group (OR= 0.095, and p= 0.03385) or the volunteers aged more than 57 years OR= 0.152; and p= 0.04069, respectively). Two non-synonymous single nucleotide polymorphisms (SNP; S249P and V327M) were common in a few patients and were predicted as damaging by SIFT and Polyphen and were further analyzed for their protein stability and function using advanced bioinformatics tools. The results suggest that TLR6 rs3796508 has a crucial role as a protective factor against colorectal cancer in the older Saudi male population.
Keywords: Colorectal Cancer, SNP, TLR6, Saudi population, gene expression, Case, control
Introduction
Colorectal cancer (CRC) is an important public health concern worldwide and remains the second leading cause of cancer-associated mortality among all malignancies [1]. Its accounts for an estimated 500,000 cancer deaths every year, with one million new malignancy cases occurring annually; metastasis of this disease is still correlated with a high mortality rate and a poor prognosis [2]. To date, CRC can be diagnosed both by fecal occult blood screening tests and optical colonoscopy [3] but, unfortunately, patients with CRC are not normally diagnosed until the disease has reached an advanced stage [4]. Therefore, a clear need exists for identifying CRC disease at the early stage to reduce the CRC-related mortality and to improve the survival of patients with advanced CRC. In the Kingdom of Saudi Arabia (KSA), CRC accounts for 12% of all cancer types and 11% of all cancer new cases; the incidence of CRC has increased in recent years and further increases are predicted in the future [5]. In the Saudi population, CRC is the second most common malignancy among various cancers and is now the most common malignancy in males [5].
Evidence from previous studies indicates that cancer is closely associated with a deregulation of in immune function and that a low immune response has a direct and inverse association with many types of cancer. In humans, the immune system consists of two classes: innate immunity and adaptive immunity [6]. An association between the dysregulation of innate immunity and cancer development risk has been reported previously in our own and other studies [7-9]. Of the two immune system classes in humans [10,11], the innate system is responsible for nonspecific immune responses mediated by phagocytotic cells, such as dendritic cells, macrophages, and neutrophils, and represents a first line defensive mechanism for protecting the host from invading microbial pathogens, pollutants, and endogenous dangerous molecules [12,13].
The human innate immunity system is activated by specific pathogen-associated molecular patterns (PAMPs). PAMPs are detected by special receptors, the pattern recognition receptors (PRRs), which are located in both cellular membrane and cytosol of the host cells [14,15]. A main form of PRRs are the Toll-like receptors (TLRs) [12,16], which function as the first line of defense against attacking pathogens. TLRs have the ability to initiate inflammatory responses and they also have roles in the proliferation, regulation, and survival of immune cells [17]. The genes for TLRs have vital roles in acquired and innate immunity and their expression profiles, which vary in different cells type, are correlated with various diseases and cancer susceptibility [18]. Dysregulation of TLR expression and activation has been reported in most inflammatory disorders, suggesting an involvement of TLR signaling in the pathogenesis of infectious diseases and autoimmune inflammatory diseases [19-21], and this dysregulation is correlated with tumor progression and carcinogenesis during chronic inflammation.
TLRs are type I single-path transmembrane proteins that contain extracellular leucine-rich repeats (which are responsible for recognition of particular pathogens), a single transmembrane helix, and a cytoplasmic Toll/interleukin-1 receptor (TIR) domain responsible for downstream signaling [22]. The human TLR family includes at least ten TLRs, named TLR1 to TLR10. Human TLR6 has a single exon and is located on chromosome 4p13 [6]. The majority of TLRs seem to function as homodimers, except for TLR6, which forms heterodimers with TLR2 [23].
Genomic instability initiated by DNA damage resulting from tobacco smoking, alcohol consumption, fat metabolism, and chemical agents, can lead to the development of CRC [24]. One common genetic variation that could be used as a biomarker to detect the development of CRC risk are single nucleotide polymorphisms (SNPs) [25], which have a recognized association with the susceptibility to inflammatory disease and other infections in different human populations [26]. Genetic variations caused by SNPs in TLR may lead to alterations in the immune system and variations have been identified in different types of tumors, such as prostate [27], gastric [28,29], bladder [30], cervical [31,32], breast [33], endometrial [34] and esophageal [32] cancers. The presence of SNPs has also been correlated with the following malignancies: acute lymphoblastic leukemia [35], hepatocellular carcinoma (HCC) [36], Hodgkin's lymphoma [37], and Burkett's lymphoma [38]. However, the TLR 2, 3, 4, and 9 polymorphism genes are not associated with increased breast cancer risk [7,39].
CRC tissues show higher expression of the TLR1, TLR2, TLR4, TLR8, and TLR9 genes [8,9,40], andTLR6 polymorphisms are linked to a risk of pancreatic cancer [41]. We hypothesized that genetic polymorphisms in TLR6 may be correlated with the risk of CRC. Consequently, the aim of the present study was to determine the potential effect of the SNPs rs3796508 at Valine 327 and rs5743810 at Serine 249 of the TLR6 gene on the CRC risk in the Saudi population when compared with healthy subjects. A second aim was to identify differences in genotype and allele distributions between the two groups according to age, sex, and tumor location. The expression profiles of TLR6 were therefore compared between patients with CRC and healthy subjects.
Materials and Methods
Patients selected for the current study
Population-based case and control groups were selected in the Kingdom of Saudi Arabia (KSA). This study included a total of 115 patients with histologically confirmed CRC and 102 healthy individuals. Blood and tissue samples were obtained from the endoscopy service of the King Khalid University Hospital (KKUH) in Riyadh, KSA. This study was approved by the local institutional review board with IRB number CAMS 13/3536. At the time of the initial recruitment of groups, all clinical data and blood and tissue samples were collected from the Saudi patients with CRC, together with matching normal colorectal tissues. All participants were asked to complete a self-administered written consent form and questionnaire regarding their socio-demographic characteristics, including age, lifestyle, smoking habits, family history of cancer, stage of CRC cancer, medications, presence of other diseases, personal medical history, and date of diagnosis (Table 1). The cases and control subjects were frequency-matched by age and sex. The study population median ages were 56.04±14.37 years for CRC cancer cases and 52.84±15.88 years for the control group (p= 077). Tissue samples were used for both RNA isolation and for immunohistochemistry, while blood samples were used for genomic DNA isolation.
General clinical parameters of the study participants
| Variables | Cancer N (%) | Control N (%) | P*- Value |
|---|---|---|---|
| Participants | 115(100%) | 102(100%) | --- |
| Gender | |||
| Males | 66(57.39%) | 60(58.82%) | 0.59 |
| Females | 49(42.61%) | 42(41.18%) | 0.46 |
| Age | |||
| Below 57 | 57(49.57%) | 62(60.78%) | --- |
| Above 57 | 58(50.43%) | 40(39.22%) | --- |
| Median Age | 56.04± 14.37 | 52.84 ± 15.88 | 0.77 |
| Tumor Localization | |||
| Colon | 76(66.09%) | --- | --- |
| Rectum | 39(33.91%) | --- | --- |
| Smoking Status | |||
| Smokers | 7(6.09%) | 5(4.91%) | 0.56 |
| Nonsmokers | 108(93.91%) | 97(95.09%) | 0.44 |
| Tumor Treatment | |||
| Under Chemotherapy | 3(2.61%) | --- | --- |
| No Chemotherapy | 112(97.39%) | --- | --- |
| Under Radiotherapy | 5(4.35%) | --- | --- |
| No Radiotherapy | 110(95.65%) | --- | --- |
DNA isolation
Sample collection has been described in detail elsewhere [8]. Genomic DNA was extracted from peripheral blood leukocytes using the Blood DNA extraction kit (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions. Briefly, 200 to 300 μl blood samples stored in EDTA tubes at -80°C were equilibrated at room temperature and then mixed with protease K digestion reagent. The reaction mixtures were incubated at 56 °C for 10 minutes, then 100% ethanol was added and the mixture was spun through the extraction column. The column membrane was washed and then the DNA was eluted with 100 μl elution buffer (AE). The isolated DNA concentration was calculated with a Nano-Drop 8000 spectrophotometer (Thermo Scientific). The DNA purity was determined by calculating the A260/A280 and A260/A230 ratios.
SNP selection and genotyping
The two selected SNPs, rs3796508 and rs5743810, were genotyped and were previously identified within the TLR6 gene (www.ncbi.nlm.nih.gov/snp/). Genotyping was carried out using a TaqMan allelic discrimination assay, as previously described [42]. The positional information about the TLR6 SNPs is presented in Table S1. Approximately 10 to 20 ng DNA from each sample was used per reaction in the genotyping assay. The tube reaction also contained 200 nM primers and 5.6 μL of 2X Universal Master Mix. All genotypes in this study were determined by endpoint reading on an ABI 7500 real-time PCR machine. The primer and probe mixtures were obtained from the assays-on-demand service of Applied Biosystems.
Total RNA isolation
Total RNA was extracted from 10 CRC tissues and 10 matching normal colon tissues using the RNA Mini kit from Qiagen (Qiagen, Hilden, Germany). The isolated RNA purity, concentration, and quality were all measured with a NanoDrop 8000 (Thermo Scientific, Paisley, UK). The extracted RNA was stored in nuclease-free collection tubes at -80°C until further use.
Complementary DNA (cDNA) synthesis
As described by Semlali et al [43,44], 1μg of total RNA from each sample was reverse transcribed into cDNA using random primers and the High-capacity cDNA Reverse Transcription kit according to the manufacturer's protocol (Applied Biosystems, USA). The conditions for the preparation of the cDNA templates for PCR analysis were 10 min at 25°C, 2 h at 37°C, and 5 min at 85°C. Afterwards, the synthesized cDNA was kept at -20 °C or 4°C in nuclease-free collection tubes for the subsequent analysis.
Real-Time quantitative reverse transcription PCR (qRT-PCR)
Real time PCR was carried out by using the QuantStudio 7Flex detection system from Applied Biosystems and was performed using the Taq™ Universal SYBR® Green Supermix kit from BIO-RAD Laboratories (San Diego, USA) according to the manufacturer's protocol. Five μL of each cDNA sample was added to a 20 μL PCR mixture containing 12.5 μL of Taq™ Universal SYBR Green SuperMix (Bio-Rad) and 0.5 μL of specific primers for TLR6 and GAPDH from Eurofins MWG operon (Canada) and 7 μL of RNase/DNase-free water. The PCR tube contents were then reacted in a PCR cycling machine to amplify the samples with the following program: a 5-minutepre-denaturation step at 95°C, followed by 40 cycles of 10-second denaturation steps at 95°C, then a 30-second annealing step at 60°C, and a final 10-second extension stage at 95°C. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as a positive control for the qRT-PCR results. The PCR results were analyzed on a7500 Real-Time PCR System (Applied Biosystems) and evaluated via the 2-∆∆Ct (Livak) relative expression method. The TLR6 and GAPDH primer sequences and amplicon sizes are described in Table 2.
Primers used in Real-Time PCR
| Gene | Primer sequence (5' to 3') | Size (bp) |
|---|---|---|
| TLR-6- sense | 5'-CATCCTATTGTGAGTTTCAGGCAT-3' | 121 |
| TLR-6- antisense | 5'-GCTTCATAGCACTACATCCCAAG-3' | |
| GAPDH- sense | 5'-GGTATCGTCGAAGGACTCATGAC-3' | 180 |
| GAPDH- antisense | 5'-ATGCCAGTGAGCTTCCCGTTCAGC-3' |
Immunohistochemistry (IHC) assay
The CRC tissues and matching normal colon tissue specimens were embedded in paraffin wax and sectioned at about 5-μm thickness with a microtome (Leica Microsystems GmbH, Wetzlar, Germany). The tumorareas in IHC slides were defined by staining each section with hematoxylin and eosin stains. The array tissue blocks were obtained as described elsewhere [45] and each slide was incubated in a hot air oven at 60°C for 15-20 minutes, followed by incubation with a primary TLR-6 antibody (Santa Cruz Biotechnology, CA, USA) at a 1/100 dilution. The other steps and analyses for IHC were conducted as described by Semlali et al. [9]. The immuno-stained slides were analyzed with an Olympus BX51 light microscope (Olympus America Inc., Center Valley, PA, USA) equipped with aDP72 Olympus digital camera with powers 400×.
IHC analysis was performed by using points (0 to 4) according to the level and range of the color in each condition: 0 point corresponds to no positive color; 1 point corresponds to <20% positive staining; 2 points means a positive staining between 21 50%; 3 points corresponds to , 51-75% positive staining; and 4 points means >75% positive staining.
Predicting the Functionality of the Non-synonymous Variants
The possible associations between genetic mutation and a varying degree of TLR6 gene mutations in patients with CRC were explored by changing the amino acid stability through mutation, analyzing using SIFT and Polyphen. Both native and mutant protein secondary structures were analyzed for solvent accessibility. A better understanding of the disease association and the specific development of inhibitors or rational drugs were obtained by constructing three dimensional structures of the mutated protein. These 3D structures were further exposed to external and internal forces using MD simulations, CHARMM and GROMACS 4.0.5, along with the Molecular Operating Environment (MOE). All the associated atoms were predicted in every single discrete step using classical equations of motion, in their simplest form of Newton's law, in millions of steps and for up to a few femtoseconds.
The possible impact of the two SNPs (S249P and V327M) on the structure/ function of the TLR6 protein was predicted using SIFT and PolyPhen. SIFT (http://sift.jcvi.org/) is a sequence homology-based tool that can predict variants as neutral or deleterious based on the normalized probability score. Variants at specific positions are considered “predicted to be deleterious” when the normalized probability score is less than 0.05, whereas variants are “predicted to be neutral” when the score greater is than 0.05 [46]. PolyPhen 2.0 (http://genetics.bwh.havard.edu/pph2/) utilizes a combination of sequence and structure-based attributes and uses naïve Bayesian classifiers for the identification of an amino acid substitution and the effect of a mutation. The output levels of “probably damaging” and “possibly damaging” were classified as deleterious (=0.5) and the benign level was classified as tolerated (=0.51) (Table 7).
SIFT and PolyPhen scores for the studies SNPs
| SIFT | PolyPhen | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| SNP | Allele | Transcript | protein | AA_pos | Wild_AA | mutant_AA | Score | Prediction | Confidence | Score | Prediction |
| rs3796508 | C|T | ENST00000381950 | ENSP00000371376 | 327 | V | M | 0.02 | DAMAGING | HIGH | 0.028 | BENIGN |
| rs5743810 | A|G | ENST00000381950 | ENSP00000371376 | 249 | S | P | 0.38 | TOLERATED | HIGH | 0 | BENIGN |
Statistical analysis
Statistical analyses were carried out as follows:
1) For gene expression, cycle threshold (Ct) values were calculated and presented as arithmetical mean ± standard deviation (SD) for TLR6 gene expression, and transformed as fold changes. Statistical significance was evaluated by Student's t-test for comparisons between groups. A p-value of < 0.05 was considered statistically significant. Graphs were constructed using Microsoft Excel 2010. All graph values represent mean values.
2) For genotyping, the Hardy-Weinberg equilibrium test (HWE) (http://ihg2.helmholtz-muenchen.de/cgi-bin/hw/hwa1.pl) was used to evaluate the frequencies of the analyzed polymorphisms of the TLR6 gene (genotypes and alleles) for the CRC patients and healthy controls. Genetic comparisons between the CRC cases and controls according to age, gender, and tumor location were achieved using the chi-square test. The odds ratios (OR) and 95% confidence intervals (CIs) were designed with Fisher's exact probability test (two-tailed). For statistical analysis, SPSS (Statistical Package for the Social Sciences) 16.0 software was used and p values <0.05 were considered statistically significant.
Results
Basic analysis distribution of clinical data parameters
Table 1 shows the distribution of the key clinical parameters of the study subjects by CRC case or healthy control status, including the total number of participants in each class, gender, age at diagnosis, median age, and tumor localization. The total number of the current study population was 217, consisting of 115 CRC cases and 102 healthy controls. Our analysis of the general demographic characteristics of the participants determined a median age ± standard deviation (SD) of 56.04±14.37 years for the CRC patients and 52.84±15.88 years for the controls. The ratio of males to females was 66 (57.39%)/49 (42.61%) for the CRC cases and 60 (58.82%)/42 (41.18%) for the healthy controls. The differences in the mean age (p=077), male ratio (p=0.59) or female ratio (p=0.46) were not statistically significant. Among the cases with CRC, 76 (66.09%) of these patients had tumors localized in the colon and 39 (33.91%)) had CRC malignancies localized in the rectum.
TLR6 gene expression profiles in colon cancer tissues and matching normal colon tissues
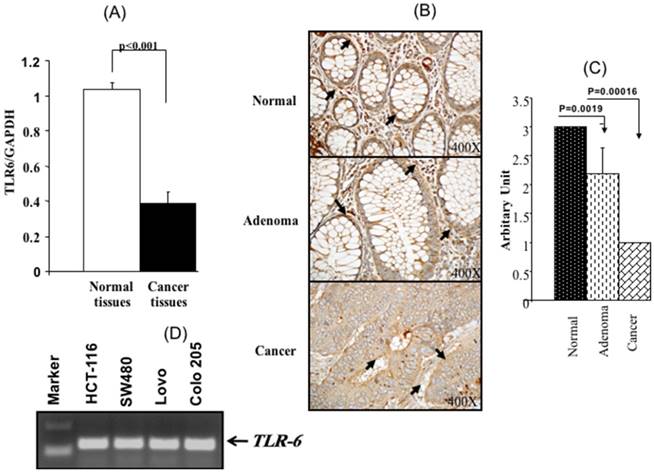
For RNA analysis, 10 matching tissue samples isolated from either normal or cancer tissues were immediately stored in RNA-later solution to protect and stabilize cellular RNA. These tissues were used later for qRT-PCR to compare the TLR6 mRNA expressions between colon cancer tissues and matching normal colon tissues. Figure 1A shows a decrease in TLR6 expression in colon cancer tissues (1.03 ± 0.04) when compared to normal colon tissues (0.38 ± 0.7, p<0.0001). The real-time PCR results for TLR9 mRNA were confirmed at the protein level by IHC, which showed a less intense positive immuno-staining for TLR6 in the cancer tissues than in the normal colon tissues, indicating a pronounced decrease in TLR6 protein in colon cancer tissues (Figure 1B).
Real time quantitative and immunohistochemistry analyses for TLR6 in matching normal and colon cancer tissues. Panel (A) shows mRNA expression of TLR6 in normal tissues and colon cancer tissues (n=10). However, protein levels were shown in Panel (B) by IHC assay using specific TLR6 antibody (n=10) and analysis by Score points (C). The panel (D) represents the TLR6 expression on colon cancer cells.
Global analysis of the correlation between TLR6 gene polymorphisms and susceptibility to CRC development
DNA was extracted from the whole blood of the 115 patients diagnosed with CRC and 102 healthy controls. The DNA was genotyped by HWE for the presence of the rs3796508 C/T and rs5743810 C/T TLR6 SNPs. The allele and genotype frequencies and the correlation analysis are presented in Table 3. Both SNPs were located in the exon region of TLR6 gene; however, the global analysis of TLR6 for two SNPs did not display any relationship with increased CRC risk in the study population. The prevalences of the Val/Val, Val/Met, and Met/Met genotypes in the TLR-6 rs3796508 SNP were 96%, 2%, and 2%, respectively, in the CRC case group and 94%, 6%, and 0%, respectively, in the control group (Table 3). The prevalences of the Ser/Ser, Ser/Pro, and Pro /Pro genotypes in the rs5743810 SNP were 2%, 25%, and 73%, respectively, in the CRC case group and 3%, 19%, and 78%, respectively, in the control group. The prevalence of the “Val” and “Met” frequencies was similar for rs3796508 in both groups. The “Val” distribution was 79% in the CRC samples and 3% in the normal control. Similarly, the “Ser” and “Pro” allele frequencies were similar for the CRC Saudi population and healthy controls from the same ethnic population. The “Ser” allele frequencies were 14% and 12%, respectively, in the CRC cases and healthy controls, whereas the prevalence of the “Pro” allele was 86% and 88%, respectively (Table 3).
Genotype frequencies of TLR6 gene polymorphism in CRC cases and controls
| SNP ID | Genotype | CRC N (%) | Controls N (%) | OR | (95% CI) | χ2 -Value | p*- Value |
|---|---|---|---|---|---|---|---|
| rs3796508 | Val/Val | 110(0.96) | 94(0.94) | Ref | |||
| Val/Met | 2(0.02) | 6(0.06) | 0.285 | 0.056-1.44 | 2.58 | 0.10795 | |
| Met/Met | 2(0.02) | 0(0.00) | 4.276 | 0.203-90.18 | 1.70 | 0.19294 | |
| Val/Met+Met/Met | 4(0.04) | 6(0.06) | 0.570 | 0.156-2.079 | 0.74 | 0.38896 | |
| Val | 222(0.97) | 194(0.97) | Ref | ||||
| Met | 6(0.03) | 6(0.03) | 0.874 | 0.277-2.75 | 0.05 | 0.81781 | |
| rs5743810 | Ser/Ser | 2(0.02) | 3(0.03) | Ref | |||
| Ser/Pro | 29(0.25) | 19(0.19) | 2.289 | 0.349-15.010 | 0.78 | 0.37792 | |
| Pro/Pro | 83(0.73) | 79(0.78) | 1.576 | 0.256-9.683 | 0.24 | 0.62065 | |
| Ser/Pro+Pro/Pro | 112(0.98) | 98(0.97) | 1.714 | 0.281-10.471 | 0.35 | 0.55493 | |
| Ser | 33(0.14) | 25(0.12) | Ref | ||||
| Pro | 195(0.86) | 177(0.88) | 0.835 | 0.478-1.458 | 0.40 | 0.52513 |
* = p value < 0.05 is significant
OR= Odds Ratio, CI= confidence interval, χ2= chi-square value
Distribution of TLR6 gene polymorphisms according to gender
The possibility of a statistically significant relationship between the risk of CRC development in the Saudi Arabia population according to gender was determined based on comparison of the distribution of the TLR6 genotype and allele frequencies from male CRC patients versus male controls and female CRC cases versus female controls and the results are summarized in Table 4 A and Table 4B, respectively. In the male group (66 patients and 60 controls), the heterozygous Val/Met was a more significant genotype of SNP rs3796508 in the control group than in the CRC case group (OR= 0.095, 95% CI: 0.005-1.807, and p= 0.03385), suggesting that the Val/Met genotype was a protective factor against CRC. The “Val” and “Met” alleles were similar in male CRC patients and male normal controls, at 98% for “Val” in the CRC cases and 96% in the controls. By contrast, the distribution of “Met” alleles was 2% in the CRC cases and 4% in the controls (Table 4A).
Genotype frequencies of TLR6gene polymorphism in CRCcases and controls based on gender
| SNP ID | Genotype | CRC N (%) | Controls N (%) | OR | (95% CI) | χ2 -Value | p*- Value |
|---|---|---|---|---|---|---|---|
| (A) Male vs control | |||||||
| rs3796508 | Total | 63 | 57 | ||||
| Val/Val | 62 (0.98) | 53 (0.93) | Ref | ||||
| Val/Met | 0(0.00) | 4(0.07) | 0.095 | 0.005-1.807 | 4.50 | 0.03385* | |
| Met/Met | 1(0.02) | 0(0.00) | 2.568 | 0.102-64.36 | 0.85 | 0.35695 | |
| Val/Met +Met/Met | 1(0.02) | 4(0.07) | 0.214 | 0.023-1.971 | 2.21 | 0.13713 | |
| Val | 124(0.98) | 110(0.96) | Ref | ||||
| Met | 2(0.02) | 4(0.04) | 0.444 | 0.080-2.469 | 0.91 | 0.42870 | |
| rs5743810 | Total | 65 | 58 | ||||
| Ser/Ser | 2(0.03) | 2(0.03) | Ref | ||||
| Ser/Pro | 18(0.28) | 11(0.19) | 1.636 | 0.201-13.34 | 0.21 | 0.64330 | |
| Pro/Pro | 45(0.69) | 45(0.78) | 1 | 0.135-7.412 | 0.00 | 1.00000 | |
| Ser/Pro+Pro/Pro | 63(0.97) | 56(0.97) | 1.125 | 0.153-8.253 | 0.01 | 0.90773 | |
| Ser | 22(0.17) | 15(0.13) | Ref | ||||
| Pro | 108(0.83) | 101(0.87) | 0.729 | 0.358-1.483 | 0.76 | 0.38192 | |
| (B) Female vs control | |||||||
| rs3796508 | Total | 51 | 41 | ||||
| Val/Val | 48 (0.94) | 39 (0.95) | Ref | ||||
| Val/Met | 2(0.04) | 2(0.05) | 0.812 | 0.109-6.034 | 0.04 | 0.83890 | |
| Met/Met | 1(0.02) | 0(0.00) | 2.443 | 0.097-61.65 | 0.81 | 0.36958 | |
| Val/Met +Met/Met | 3(0.06) | 2(0.05) | 1.219 | 0.194-7.662 | 0.04 | 0.83273 | |
| Val | 98(0.96) | 80(0.98) | Ref | ||||
| Met | 4(0.04) | 2(0.02) | 1.633 | 0.292-9.144 | 0.32 | 0.69448 | |
| rs5743810 | Total | 49 | 41 | ||||
| Ser/Ser | 0(0.00) | 1(0.02) | Ref | ||||
| Ser/Pro | 11(0.22) | 8(0.20) | 4.059 | 0.15-112.4 | 1.29 | 0.25668 | |
| Pro/Pro | 38(0.78) | 32(0.78) | 3.554 | 0.14-90.243 | 1.17 | 0.27982 | |
| Ser/Pro+Pro/Pro | 49(1) | 40(0.98) | 3.667 | 0.145-92.45 | 1.21 | 0.27162 | |
| Ser | 11(0.11) | 10(0.12) | Ref | ||||
| Pro | 87(0.89) | 72(0.88) | 1.098 | 0.441-2.733 | 0.04 | 0.83990 | |
* = p value < 0.05 is significant
OR= Odds Ratio, CI= confidence interval, χ2= chi-square value
The distribution of the genotypes and alleles for TLR6 rs5743810 in males did not show any statistically significant differences either for genotype or allele frequencies between the CRC cases and healthy controls (Table 4A). The prevalence of Ser/Ser (the ancestral genotype) was 3% in the male CRC cases and the male controls (Table 4A), while the Ser/Pro genotype prevalence was 28% and 19% in the cases and controls, respectively and the Pro/Pro genotype prevalence was 69% and 78%, respectively (Table 4A). The Ser and Pro allele distributions were 17% and 83%, respectively, in the CRC study population and 13% and 87, respectively, for healthy controls from the Saudi Arabia population (Table 4A). By contrast, no genotype or allele frequencies differences were observed in female CRC cases or female healthy controls for either of the studied TLR6 SNPs (rs3796508 and rs5743810). The frequency distributions for the Val/Met and Met/Met rs3796508 mutations were similar in both study groups and no differences were noted between the female study groups in terms of the prevalence of Ser/Pro and Pro/Pro distribution for rs5743810 (Table 4B). These two study SNPs did not appear to have any association with CRC risk in females of the Saudi Arabian population (p > 0.05; Table 4B).
Distribution of TLR6 gene polymorphisms according to age at CRC diagnosis
The influence of the age at CRC diagnosis on the genetic frequency of TLR6 was examined by stratifying the study subjects by age as either less than 57 years of age or 57 years of age and over. All genotype prevalences and the statistical analysis for the individual TLR6 polymorphisms are presented in Tables 5A and 5B, respectively. No significant differences were detected for the genotypes or the allele prevalence for the subgroup analysis of age (p >0.05) for the rs5743810 SNP and the healthy control group (Tables 5A and 5B). However, the Val/Met heterozygote genotype of SNP rs3796508 provided significant protection against the risk of CRC in the patients diagnosed with CRC after 57 years of age. The frequency of the Val/Me genotype was 2% and 11%, respectively, for the CRC cases and healthy controls aged more than 57 years at diagnosis ( Table 5B); however, the frequency of this mutant Val/Met genotype was similar in both study populations aged less than 57 years at diagnosis (Table 5A). This age difference suggests that the presence of this genotype is an essential factor for protection against the risk of CRC development in the Saudi population (OR= 0.152; 95% CI: 0.016-1.415, and p= 0.04069; Table 5B); whereas the genotype frequency did not indicate a significant risk for the same SNPs in patients aged less than 57 years (Table 5A). By contrast, the TLR6 rs5743810 genotype and allele variation did not indicate any difference in CRC susceptibility in either sub-population (Tables 5A and 5B). The genotype and allele frequencies for TLR6 rs5743810 were similar in both populations regardless of age at diagnosis (Tables 5A and 5B).
Genotype frequencies of TLR6 gene polymorphisms in CRC cases and controls based on age.
| SNP ID | Genotype | CRC n (%) | Controls n (%) | OR | (95% CI) | χ2 -Value | p*- Value |
|---|---|---|---|---|---|---|---|
| (A) Below 57 years | |||||||
| rs3796508 | Total | 56 | 62 | ||||
| Val/Val | 54 (0.96) | 60(0.97) | Ref | ||||
| Val/Met | 1(0.02) | 2(0.03) | 0.556 | 0.049-6.301 | 0.23 | 0.63067 | |
| Met/Met | 1(0.02) | 0(0.00) | 3.330 | 0.133-83.473 | 1.10 | 0.29416 | |
| Val/Met +Met/Met | 2(0.04) | 2(0.03) | 1.111 | 0.151-8.162 | 0.01 | 0.91749 | |
| Val | 109(0.97) | 122(0.98) | Ref | ||||
| Met | 3(0.03) | 2(0.02) | 1.679 | 0.275-10.24 | 0.32 | 0.67815 | |
| rs5743810 | Total | 58 | 62 | ||||
| Ser/Ser | 1(0.02) | 2(0.03) | Ref | ||||
| Ser/Pro | 11(0.19) | 13(0.21) | 1.692 | 0.135-21.270 | 0.17 | 0.68123 | |
| Pro/Pro | 46(0.79) | 47(0.76) | 1.957 | 0.172-22.338 | 0.30 | 0.58229 | |
| Ser/Pro+Pro/Pro | 57(0.98) | 60(0.97) | 1.900 | 0.168-21.532 | 0.28 | 0.59852 | |
| Ser | 13(0.11) | 17(0.14) | Ref | ||||
| Pro | 103(0.89) | 107(0.86) | 1.259 | 0.582-2.722 | 0.34 | 0.55797 | |
| (B) Above 57 years | |||||||
| rs3796508 | Total | 58 | 38 | ||||
| Val/Val | 56 (0.96) | 34 (0.89) | Ref | ||||
| Val/Met | 1(0.02) | 4(0.11) | 0.152 | 0.016-1.415 | 3.52 | 0.04069* | |
| Met/Met | 1(0.02) | 0(0.00) | 1.832 | 0.073-46.24 | 0.60 | 0.43739 | |
| Val/Met +Met/Met | 2(0.04) | 4(0.11) | 0.304 | 0.053-1.747 | 1.96 | 0.16120 | |
| Val | 113(0.97)) | 72(0.95) | Ref | ||||
| Met | 3(0.03) | 4(0.05) | 0.478 | 0.104-2.198 | 0.94 | 0.43795 | |
| rs5743810 | Total | 56 | 39 | ||||
| Ser/Ser | 1(0.02) | 1(0.03) | Ref | ||||
| Ser/Pro | 18(0.32) | 6(0.15) | 3.000 | 0.162-55.721 | 0.59 | 0.44379 | |
| Pro/Pro | 37(0.66) | 32(0.82) | 1.156 | 0.069-19.242 | 0.01 | 0.91933 | |
| Ser/Pro+Pro/Pro | 55(0.98) | 38(0.97) | 1.447 | 0.088-23.860 | 0.07 | 0.79488 | |
| Ser | 20(0.18) | 8(0.10) | Ref | ||||
| Pro | 92(0.82) | 70(0.90) | 0.526 | 0.219-1.263 | 2.11 | 0.14596 | |
* = p value < 0.05 is significant
OR= Odds Ratio, CI= confidence interval, χ2= chi-square value
Correlation between TLR6 gene polymorphisms variation and tumor location
CRC malignancy was detected in either the colon or the rectum. Table 6 shows the genotype and allele prevalence between colon and rectum cancers for the two SNPs of the TLR6 gene (Tables 6A and 6B). The genotype and allele frequencies of the TLR6 gene polymorphisms were similar for the CRC cases and healthy controls regardless of the cancer location.
Genotype frequencies of TLR6gene polymorphism in CRC cases and controls based on tumor location
| SNP ID | Genotype | CRC n (%) | Controls n (%) | OR | (95% CI) | χ2 -Value | p*- Value |
|---|---|---|---|---|---|---|---|
| (A) Colon area vs control | |||||||
| rs3796508 | Total | 67 | 100 | ||||
| Val/Val | 65 (0.97) | 94 (0.94) | Ref | ||||
| Val/Met | 2 (0.03) | 6 (0.06) | 0.482 | 0.094-2.463 | 0.80 | 0.37122 | |
| Met/Met | 0 (0.00) | 0 (0.00) | 1.443 | 0.028-73.63 | -- | 1.00000 | |
| Val/Met +Met/Met | 2 (0.03) | 6 (0.06) | 0.482 | 0.094-2.463 | 0.80 | 0.37122 | |
| Val | 132(0.99) | 194 (0.97) | Ref | ||||
| Met | 2 (0.01) | 6 (0.03) | 0.490 | 0.097-2.464 | 0.78 | 0.48592 | |
| rs5743810 | Total | 67 | 101 | ||||
| Ser/Ser | 1 (0.01) | 3 (0.03) | Ref | ||||
| Ser/Pro | 18 (0.27) | 19 (0.19) | 2.842 | 0.27-29.897 | 0.81 | 0.36757 | |
| Pro/Pro | 48 (0.72) | 79 (0.78) | 1.823 | 0.184-18.02 | 0.27 | 0.60256 | |
| Ser/Pro+Pro/Pro | 66 (0.99) | 98 (0.97) | 2.020 | 0.206-19.84 | 0.38 | 0.53843 | |
| Ser | 20 (0.15) | 25 (0.12) | Ref | ||||
| Pro | 114 (0.85) | 177 (0.88) | 0.805 | 0.427-1.517 | 0.45 | 0.50171 | |
| (B) Rectum area vs control | |||||||
| rs3796508 | Total | 34 | 100 | ||||
| Val/Val | 33 (0.97) | 94 (0.94) | Ref | ||||
| Val/Met | 0 (0.00) | 6 (0.06) | 0.217 | 0.012-3.957 | 2.07 | 0.14987 | |
| Met/Met | 1 (0.03) | 0 (0.00) | 8.463 | 0.34-212.82 | 2.79 | 0.09506 | |
| Val/Met +Met/Met | 1 (0.03) | 6 (0.06) | 0.09506 | 0.055-4.091 | 0.48 | 0.48865 | |
| Val | 66 (0.97) | 194 (0.97) | Ref | ||||
| Met | 2 (0.03) | 6 (0.03) | 0.980 | 0.193-4.973 | 0.00 | 1.03893 | |
| rs5743810 | Total | 35 | 101 | ||||
| Ser/Ser | 1 (0.03) | 3 (0.03) | Ref | ||||
| Ser/Pro | 8 (0.23) | 19 (0.19) | 1.263 | 0.114-14.05 | 0.04 | 0.84902 | |
| Pro/Pro | 26 (0.74) | 79 (0.78) | 0.987 | 0.098-9.909 | 0.00 | 0.99136 | |
| Ser/Pro+Pro/Pro | 34(0.97) | 98 (0.97) | 1.041 | 0.105-10.35 | 0.00 | 0.97276 | |
| Ser | 10 (0.14) | 25 (0.12) | Ref | ||||
| Pro | 60 (0.86) | 177 (0.88) | 0.847 | 0.385-1.867 | 0.17 | 0.68095 | |
* = p value < 0.05 is significant
OR= Odds Ratio, CI= confidence interval, χ2= chi-square value
The distribution of the heterozygous Val/Met mutation and double homozygous Met/Met mutation was 3% in the population having cancer in the colon area and 6% in the controls; this distribution was 0% in both sub-populations for the Met/Met genotype (Table 6A). The Val/Met and Met/Met genotype frequencies for TLR6 rs3796508 were, respectively, 0% and 3% in the CRC patients with rectal cancer and 6% and 0% in normal control patients with rectal cancer (Table 6B). The frequency for the mutant “Met” allele was similar in both sub-populations having either colon or rectal cancer when compared to healthy controls (p>0.05) (Tables 6A and 6B). The TLR6 rs5743810 SNP did not show any correlation with CRC susceptibility or cancer location for either the genotype or phenotype variations (Tables 6A and 6B).
Protein modeling and molecular dynamics (MD) simulations for the altered amino acids
We also examined whether amino acid changes might affect the protein function and stability by performing a prediction study for both the S249P and V327M mutations using SIFT and PolyPhen. Similar studies performed previously by our group [47] demonstrated that function prediction resulted in greater correlation of the gene expression results. In addition, the absorbed and compensated movements of the structural elements in the solvated TLR6 protein have not been studied in detail elsewhere. The current study is the first to report a solvated model for the mutated TLR6 protein and indicated a high association between the predicted structural and functional effects of TLR6.
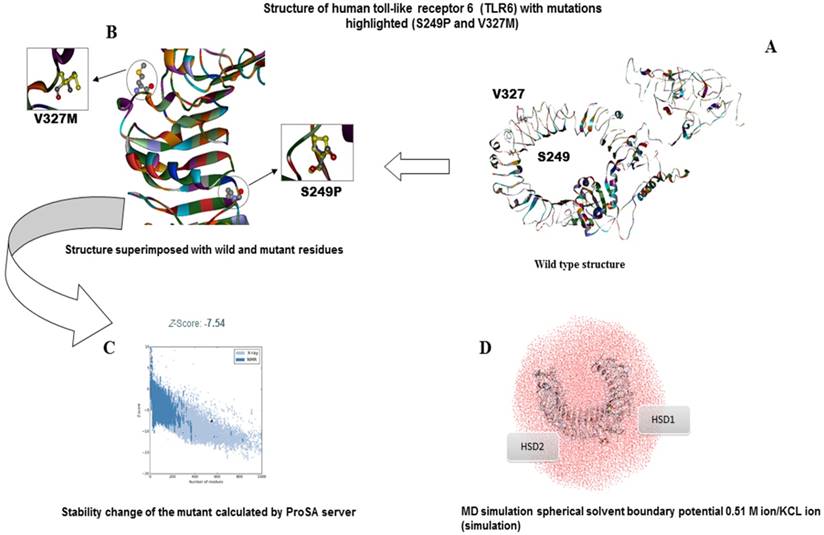
The SIFT and PolyPhen results indicated that both the S249P and V327M mutations were damaging, so they were further examined in silico for 3D structural functions. We used the X-Ray diffraction structure of Toll-like receptor 6 (3A79), available at the PDB database, as the reference model. A close comparison of the native protein (3A79) with the predicted mutant structure of all the amino acid variations and their solvent accessibility, including secondary structures, were modeled using the MD simulation. Prediction using I-TASSER returned the five best models according to C-score, which was below 0.3 after TM-alignment, and these were selected for homology modeling (threading template alignments). Once generated, the mutant protein stability was checked using the commercially available BioLuminate (Shrodinger) software. Residue scanning for predicting protein stability was performed using the mutant secondary structure. The generated 3D models were checked for quality using ProSA-web. The Z-score, which measures the total energy deviation of the wild type and the mutant protein, was calculated and was in an acceptable range (-7.54). The normal motion was simulated by performing energy minimization and molecular dynamics using CHARMM (Chemistry at HARvard Macromolecular Mechanics) [48,49]. The MD simulations were conducted under conditions of 5-fs time, 400 K temperature, 1 atm constant pressure, and below periodic solvent boundary conditions using GROMACS 4.0.5 [50].The KoBaMIN program [51,52] was used to refine the predicted TLR6 structure. The Particle Mesh Ewald methodology [53] was utilized for electrostatic interaction using a 12 Å cut-off for vdW interactions. KCl (51 M) ions were added to the simulation box to neutralize the overall negative charge of the structures. The native and mutated 3D structures were analyzed by CCP4 (QtMG). NOMAD-Ref was used for energy minimization [54]. The whole protein structure including water molecules was generated by immersing it in solvent-based simulation parameters and energy minimization to analyze potential energies, structural fluctuations, coordinate stability, and geometrical features (Figure 2).
Structure of human toll-like receptor 6 (TLR6) with mutations highlighted (S249P and V327M). A) Ribbon diagram of human TLR6 protein showing all the secondary structures. B) Location of mutations (S249P and V327M) identified in the TLR6 protein of colon cancer patients, Structure of TLR6 superimposed with wild and mutant residues with mutation S249P and V327M enlarged. C). Stability change of the mutant calculated by ProSA server. D) Molecular dynamics (MD) simulation showing truncated octahedron boundary explicit water solvated and hydrogen atoms. The visual inspection also allow to identify the side chain of histidine residues (HSD1 - HSD2) involved in hydrogen bonding with the surrounding molecules
Discussion
CRC is one of the most frequently diagnosed malignancies around the world [55] and lifestyle is identified as one of the most important risk factors for CRC. The first line of human defense against pathogens such as fungi, viruses, and bacteria is generally an innate immune system, which is responsible for initiation of immune responses to repair damaged tissues and eliminate infections caused by microorganisms [13,56]. Detecting and eliminating microorganisms is a key function of our immune system, which can discriminate between normal native molecules and abnormal foreign molecules undesirable to our bodies. In humans, the immune system consists of two main classes: innate immunity and adaptive immunity [6]. Innate immunity is mainly activated and regulated by TLRs present in the majority of immune cells, such as neutrophils, dendritic cells, and macrophages [57]. Current evidence now supports a correlation between malignancy and loss of immune function, resulting in a direct association with several types of malignancies [7-9,44,58]. Therefore, the TLRs appear to could play an essential role in innate immunity [17] and could therefore be associated with the risk of malignancy.
TLRs, which are transmembrane proteins characterized by an extracellular leucine-rich domain and a cytoplasmic tail, are known as innate immunity receptors that are essential for detecting invading pathogens and are associated with altered susceptibility to many diseases, including cancers [9,58-61]. Multiple SNPs have been found in TLRs that have the potential to induce functional changes with respect to cancer or infectious disease [62]. In Saudi patients CRC is typically diagnosed at an advance stage, so a new method is needed to diagnose CRC at early stage. Therefore, SNPs represent a major common genetic variation that could be used as biomarkers to detect the risk of development of CRC [25]. Furthermore, SNPs are associated with susceptibility to inflammatory diseases and other certain infections in different human populations [26].
The human TLR6 gene is generally expressed by epithelial cells, which are considered the first cell types that are exposed to foreign agents and can join with TLR2 to make heterodimers that can lead to activation of NFκB and MAPK intracellular signaling pathways [63]. The hypothesis of the current study was that CRC may be initiated either by changing a nucleotide within the TLR6 gene or by an effect on TLR6 gene expression. Few studies have focused on the possible association of CRC development risk and TLR6 SNPs. For this reason, the focus of the present study was to validate both the TLR6 expression and TLR6 genetic polymorphisms (such as rs3796508 and rs5743810) in the Saudi population. Our results indicated that the TLR6 protein is found at higher levels in normal colon epithelial cells than in colon cancer tissues, suggesting a role for this protein in the development of normal colon tissues. The TLR6 mRNA expression was also significantly increased in normal colon tissues compared to matching colon malignancy tissues, in agreement with previous work that showed decreased levels of TLR6 expression in different malignancies, including hepatocellular carcinoma [64], esophageal carcinoma [65], and lymphoma [66]. Other studies have shown significant increases in expression of this gene in human renal carcinoma cell lines [67] and in head and neck squamous cell carcinoma [68] when compared to healthy cells. The present study findings suggest that TLR6 may have an anticancer function in terms of the risk of colon cancer development, since TLR6 expression was significantly reduced in cancer tissues compared to normal tissues. This difference may reflect an epigenetic mechanism within TLR6 in colon cancer tissue.
Many genetic and epigenetic mechanisms can be involved in the decrease in TLR6 expression and lead to alteration of TLR6 gene expression. These modifications include cytosine-phosphate-guanine (CpG) hypermethylation, alterations of non-coding RNAs, acetylation of the lysine in the histones [69,70], and microRNAs [71]. Another mechanism that could play a role in gene expression reduction is somatic mutation [70]. These mutations or single nucleotide changes, particularly in the TLR6 exons, can contribute to colon cancer formation. In the current study, we also examined a possible relationship of two common TLR6 polymorphisms with CRC in the KSA population. Our global statistical analysis of two TLR6 SNPs showed no clear evidence for an association between either TLR6 rs3796508 or rs5743810 polymorphisms and susceptibility to CRC development in the Saudi Arabian population, indicating that these SNPs are located in an exon region of the TLR6 gene that may have an impact on TLR6 expression and function. Similar findings have been reported previously regarding the influence of genotype of diverse TLRs and the close linkage between deregulation of TLR gene expression and inflammatory cytokine secretion. Veltkamp et al. (2007) showed that, upon TLR2 stimulation, the mutant homozygous genotype of the TLR2 rs4696480 SNP was correlated with a higher pro-inflammatory cytokine secretion, as well as TNF- alpha, IL-6 and IL-12 secretion, when compared to the double mutant heterozygote or wild genotype, indicating an increased risk of development of chronic diseases [72]. Similarly, Liu et al. (2011) reported that the AA genotype of the TLR2 promoter variant rs4696480 SNP was associated with a higher FOXP3 expression in regulatory T cells of maternal atopy [73].
Evidence from the latest studies shows that polymorphisms within TLR6 gene are correlated with mild malaria [74], pancreatic cancer [41], and long-term smokers [75]. In addition, TLR6 SNP rs5743810 is linked with endometritis in the American population [76]. Yang H et al. (2014) also showed that polymorphism of TLR6 rs3796508 increased the Klebsiella pneumoniae infection risk [77] and polymorphism of TLR6 rs5743810 was correlated with an increased risk of asthma increase [78]. However, our previous work on the same study population showed that SNP rs3796508 was not associated with cigarette smokers, but a correlation was evident between Saudi cigarette smokers compared to non-smokers for the rs5743810 SNP of TLR6 [75]. By contrast, our study demonstrated a significant association between genetic variations of TLR6 polymorphism and male CRC cases and CRC cases over 57 years of age at the time of diagnosis. We found that the Val/Met genotype of SNP rs3796508 had a significantly higher frequency in the control group than in the CRC cases. Therefore, this result suggests that this genotype is a protective against CRC in this sub-group of this ethnic population. This can be explained by the considerable loss of sex hormone that protects against the development of colon cancer at this age, especially for women in menopause, where a drop occurs in sex hormones like estrogen. The presence of harmful habits of life hygiene, as well as the problem of consanguinity among the Saudi population, was predominant factors that explain these genetic variations and their implications in increasing the CRC risk in the Saudi Arabian population. In this population, the rate of CRC has continued to climb considerably during the past 10 years and far exceeds the international average to now represent the first ranked cancer in males in the KSA.
The TLR6 Valine 327 and Serine 249 substitutions altered protein stability due to the size differences between wild-type and mutant residues, which caused the new residue to be in an incorrect position to make the required hydrogen bond (Figure 2; B) as the resulting difference in hydrophobicity affects hydrogen bond formation. The wild-type residue is predicted to be located in an α-helix (Figure 2; C).
Proline disrupts the α-helix when it is not located at one of the first 3 positions of that helix. In case of the mutation at hand, the helix will be disturbed and this can have severe effects on the structure of the protein as the wild-type residue is not conserved at this position. The observed mutant residue was not among the residue types observed at this position in other proteins and is located near a highly conserved position. Homologous sequences might indicate that the mutation is possibly damaging to the protein. The mutated residue is located in a domain that is important for the activity of the protein and in contact with another domain that is known to be involved in binding (Figure 2; D). The interaction between these domains could be disturbed by the mutation, which might affect signal transduction between the domains. One possibility is that this interaction is important for the correct function of the protein. The mutation can affect this interaction, thereby affecting protein function.
In summary: Due to the large number of studies being carried out on the role ofTLR6 on the development of diverse molecular therapies for different types of cancers, further research on the role of TLR6 with more samples and with other ethnic populations with potentially malignant disorders should be considered. Our study revealed the possible relationship between the TLR6- rs3796508 polymorphism and the development of malignant CRC. Our data suggest that TLR6 rs3796508 has a crucial role as a protective factor against CRC in the older male Saudi population.
Abbreviations
CRC: Colorectal cancer; KSA: Kingdom of Saudi Arabia; PAMPs: Specific pathogen-associated molecular patterns; PRRs: Pattern recognition receptors; TLR: Toll-like receptor; TIR: Toll/interleukin-1 receptor; MAL: MyD88-adaptor-like; TLR6: Toll-like receptor 6; SNP: Single nucleotide polymorphism; KKUH: King Khalid University Hospital; Bps: Base pairs; MOE: Molecular Operating Environment; DNA: Deoxyribonucleic acid; cDNA: Complementary DNA; PCR: Polymerase chain reaction; SNAP: Screening for Non-Acceptable Polymorphisms; MD: molecular dynamics; CHARMM: Chemistry at HARvard Macromolecular Mechanics; qRT-PCR: Real-Time Quantitative reverse transcription PCR; GAPDH: Glyceraldehyde 3-phosphate dehydrogenase; IHC: Immunohistochemistry; HWE: Hardy—Weinberg equilibrium test; SPSS: Statistical Package of the Social Sciences; OR: Odds ratio; CIs: Confidence intervals; SD: Standard deviation; CpG: Cytosine-phosphate-guanine.
Supplementary Material
Supplementary table.
Acknowledgements
The authors extend their appreciation to the Deanship of Scientific Research at King Saud University for funding this work through research group No (RG-1440-044).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Zhang Y, Zarrabi K, Hou W, Madajewicz S, Choi M, Zucker S. et al. Assessing Clinical Outcomes in Colorectal Cancer with Assays for Invasive Circulating Tumor Cells. Biomedicines. 2018:6
2. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359-86
3. Siegel RL, Miller KD, Fedewa SA, Ahnen DJ, Meester RGS, Barzi A. et al. Colorectal cancer statistics, 2017. CA Cancer J Clin. 2017;67:177-93
4. Marques I, Araujo A, de Mello RA. Anti-angiogenic therapies for metastatic colorectal cancer: current and future perspectives. World J Gastroenterol. 2013;19:7955-71
5. Ibrahim EM, Zeeneldin AA, El-Khodary TR, Al-Gahmi AM, Bin Sadiq BM. Past, present and future of colorectal cancer in the Kingdom of Saudi Arabia. Saudi journal of gastroenterology: official journal of the Saudi Gastroenterology Association. 2008;14:178-82
6. Takeda K, Kaisho T, Akira S. Toll-like receptors. Annual review of immunology. 2003;21:335-76
7. Semlali A, Jalouli M, Parine NR, Al Amri A, Arafah M, Al Naeem A. et al. Toll-like receptor 4 as a predictor of clinical outcomes of estrogen receptor-negative breast cancer in Saudi women. Onco Targets Ther. 2017;10:1207-16
8. Semlali A, Parine NR, Al Amri A, Azzi A, Arafah M, Kohailan M. et al. Association between TLR-9 polymorphisms and colon cancer susceptibility in Saudi Arabian female patients. Onco Targets Ther. 2017;10:1-11
9. Semlali A, Reddy Parine N, Arafah M, Mansour L, Azzi A, Al Shahrani O. et al. Expression and Polymorphism of Toll-Like Receptor 4 and Effect on NF-kappaB Mediated Inflammation in Colon Cancer Patients. PLoS One. 2016;11:e0146333
10. Kawai T, Akira S. TLR signaling. Cell death and differentiation. 2006;13:816-25
11. O'Neill LA. When signaling pathways collide: positive and negative regulation of toll-like receptor signal transduction. Immunity. 2008;29:12-20
12. Takeda K, Akira S. Toll-like receptors. Current protocols in immunology / edited by John E Coligan [et al]. 2015;109:14.2.1-2.0
13. Akira S, Takeda K, Kaisho T. Toll-like receptors: critical proteins linking innate and acquired immunity. Nature immunology. 2001;2:675-80
14. Kumar H, Kawai T, Akira S. Pathogen recognition by the innate immune system. Int Rev Immunol. 2011;30:16-34
15. Meylan E, Tschopp J, Karin M. Intracellular pattern recognition receptors in the host response. Nature. 2006;442:39-44
16. Aderem A, Ulevitch RJ. Toll-like receptors in the induction of the innate immune response. Nature. 2000;406:782-7
17. Rich AM, Hussaini HM, Parachuru VP, Seymour GJ. Toll-like receptors and cancer, particularly oral squamous cell carcinoma. Front Immunol. 2014;5:464
18. Messaritakis I, Stogiannitsi M, Koulouridi A, Sfakianaki M, Voutsina A, Sotiriou A. et al. Evaluation of the detection of Toll-like receptors (TLRs) in cancer development and progression in patients with colorectal cancer. PLoS One. 2018;13:e0197327
19. Liew FY, Xu D, Brint EK, O'Neill LA. Negative regulation of toll-like receptor-mediated immune responses. Nature reviews Immunology. 2005;5:446-58
20. Cook DN, Pisetsky DS, Schwartz DA. Toll-like receptors in the pathogenesis of human disease. Nature immunology. 2004;5:975-9
21. Himmel ME, Hardenberg G, Piccirillo CA, Steiner TS, Levings MK. The role of T-regulatory cells and Toll-like receptors in the pathogenesis of human inflammatory bowel disease. Immunology. 2008;125:145-53
22. Manavalan B, Basith S, Choi S. Similar Structures but Different Roles - An Updated Perspective on TLR Structures. Frontiers in physiology. 2011;2:41
23. Kang JY, Nan X, Jin MS, Youn SJ, Ryu YH, Mah S. et al. Recognition of lipopeptide patterns by Toll-like receptor 2-Toll-like receptor 6 heterodimer. Immunity. 2009;31:873-84
24. Charames GS, Bapat B. Genomic instability and cancer. Curr Mol Med. 2003;3:589-96
25. Ulrich CM, Robien K, McLeod HL. Cancer pharmacogenetics: polymorphisms, pathways and beyond. Nat Rev Cancer. 2003;3:912-20
26. Vijay K. Toll-like receptors in immunity and inflammatory diseases: Past, present, and future. Int Immunopharmacol. 2018;59:391-412
27. Stevens VL, Hsing AW, Talbot JT, Zheng SL, Sun J, Chen J. et al. Genetic variation in the toll-like receptor gene cluster (TLR10-TLR1-TLR6) and prostate cancer risk. International journal of cancer Journal international du cancer. 2008;123:2644-50
28. Kupcinskas J, Wex T, Bornschein J, Selgrad M, Leja M, Jonaitis L. et al. Lack of Association between Gene Polymorphisms of Angiotensin Converting Enzyme, Nod-Like Receptor 1, Toll-Like Receptor 4 and Fas/Fasl with the Presence of Helicobacter Pylori-Induced Premalignant Gastric Lesions and Gastric Cancer in Caucasians. Helicobacter. 2011;16:79 -
29. Hold GL, Rabkin CS, Gammon MD, Berry SH, Smith MG, Lissowska J. et al. CD14-159C/T and TLR9-1237T/C polymorphisms are not associated with gastric cancer risk in Caucasian populations. Eur J Cancer Prev. 2009;18:117-9
30. Singh V, Srivastava N, Kapoor R, Mittal RD. Single-nucleotide polymorphisms in genes encoding toll-like receptor -2, -3, -4, and -9 in a case-control study with bladder cancer susceptibility in a North Indian population. Arch Med Res. 2013;44:54-61
31. Roszak A, Lianeri M, Sowinska A, Jagodzinski PP. Involvement of toll-like receptor 9 polymorphism in cervical cancer development (vol 39, pg 8425, 2012). Molecular biology reports. 2013;40:731 -
32. Roszak A, Lianeri M, Sowinska A, Jagodzinski PP. Involvement of toll-like receptor 9 polymorphism in cervical cancer development. Molecular biology reports. 2012;39:8425-30
33. Etokebe GE, Knezevic J, Petricevic B, Pavelic J, Vrbanec D, Dembic Z. Single-Nucleotide Polymorphisms in Genes Encoding Toll-Like Receptor-2,-3,-4, and-9 in Case-Control Study with Breast Cancer. Genet Test Mol Bioma. 2009;13:729-34
34. Ashton KA, Proietto A, Otton G, Symonds I, McEvoy M, Attia J. et al. Toll-Like Receptor (TLR) and Nucleosome-binding Oligomerization Domain (NOD) gene polymorphisms and endometrial cancer risk. BMC cancer. 2010:10
35. Miedema KG, Tissing WJ, Te Poele EM, Kamps WA, Alizadeh BZ, Kerkhof M. et al. Polymorphisms in the TLR6 gene associated with the inverse association between childhood acute lymphoblastic leukemia and atopic disease. Leukemia. 2012;26:1203-10
36. Xie JJ, Jiang SY, Shi MM, Song YY, Shen BY, Deng XX. et al. The association between Toll-like receptor 2 single-nucleotide polymorphisms and hepatocellular carcinoma susceptibility. BMC cancer. 2012:12
37. Mollaki V, Georgiadis T, Tassidou A, Ioannou M, Daniil Z, Koutsokera A. et al. Polymorphisms and haplotypes in TLR9 and MYD88 are associated with the development of Hodgkin's lymphoma: a candidate-gene association study. J Hum Genet. 2009;54:655-9
38. Noack J, Jordi M, Zauner L, Alessi D, Burch A, Tinguely M. et al. TLR9 agonists induced cell death in Burkitt's lymphoma cells is variable and influenced by TLR9 polymorphism. Cell death & disease. 2012:3
39. Etokebe GE, Knezevic J, Petricevic B, Pavelic J, Vrbanec D, Dembic Z. Single-nucleotide polymorphisms in genes encoding toll-like receptor -2, -3, -4, and -9 in case-control study with breast cancer. Genet Test Mol Biomarkers. 2009;13:729-34
40. Lu CC, Kuo HC, Wang FS, Jou MH, Lee KC, Chuang JH. Upregulation of TLRs and IL-6 as a marker in human colorectal cancer. International journal of molecular sciences. 2015;16:159-77
41. Cotterchio M, Lowcock E, Bider-Canfield Z, Lemire M, Greenwood C, Gallinger S. et al. Association between Variants in Atopy-Related Immunologic Candidate Genes and Pancreatic Cancer Risk. PLoS One. 2015;10:e0125273
42. Alanazi M, Pathan AA, Abduljaleel Z, Shaik JP, Alabdulkarim HA, Semlali A. et al. Association between PARP-1 V762A polymorphism and breast cancer susceptibility in Saudi population. PloS one. 2013;8:e85541
43. Semlali A, Jacques E, Plante S, Biardel S, Milot J, Laviolette M. et al. TGF-beta suppresses EGF-induced MAPK signaling and proliferation in asthmatic epithelial cells. American journal of respiratory cell and molecular biology. 2008;38:202-8
44. Semlali A, Leung KP, Curt S, Rouabhia M. Antimicrobial decapeptide KSL-W attenuates Candida albicans virulence by modulating its effects on Toll-like receptor, human beta-defensin, and cytokine expression by engineered human oral mucosa. Peptides. 2011;32:859-67
45. Parker RL, Huntsman DG, Lesack DW, Cupples JB, Grant DR, Akbari M. et al. Assessment of interlaboratory variation in the immunohistochemical determination of estrogen receptor status using a breast cancer tissue microarray. American journal of clinical pathology. 2002;117:723-8
46. Ng PC, Henikoff S. Predicting the effects of amino acid substitutions on protein function. Annu Rev Genom Hum G. 2006;7:61-80
47. Alanazi M, Al-Arfaj AS, Abduljaleel Z, Al-Arfaj HF, Parine NR, Shaik JP. et al. Novel Hypoxanthine Guanine Phosphoribosyltransferase Gene Mutations in Saudi Arabian Hyperuricemia Patients. BioMed research international. 2014
48. Brooks BR, Brooks CL, Mackerell AD, Nilsson L, Petrella RJ, Roux B. et al. CHARMM: The Biomolecular Simulation Program. J Comput Chem. 2009;30:1545-614
49. Jo S, Jiang W, Lee HS, Roux B, Im W. CHARMM-GUI Ligand Binder for Absolute Binding Free Energy Calculations and Its Application. J Chem Inf Model. 2013;53:267-77
50. van der Spoel D, van Maaren PJ, Caleman C. GROMACS molecule & liquid database. Bioinformatics. 2012;28:752-3
51. Delarue M, Dumas P. On the use of low-frequency normal modes to enforce collective movements in refining macromolecular structural models. Proceedings of the National Academy of Sciences of the United States of America. 2004;101:6957-62
52. Rodrigues JPGLM, Levitt M, Chopra G. KoBaMIN: a knowledge-based minimization web server for protein structure refinement. Nucleic Acids Res. 2012;40:W323-W8
53. Darden T, Perera L, Li LP, Pedersen L. New tricks for modelers from the crystallography toolkit: the particle mesh Ewald algorithm and its use in nucleic acid simulations. Struct Fold Des. 1999;7:R55-R60
54. Suhre K, Sanejouand YH. ElNemo: a normal mode web server for protein movement analysis and the generation of templates for molecular replacement. Nucleic Acids Res. 2004;32:W610-W4
55. Jahani-Sherafat S, Alebouyeh M, Moghim S, Ahmadi Amoli H, Ghasemian-Safaei H. Role of gut microbiota in the pathogenesis of colorectal cancer; a review article. Gastroenterol Hepatol Bed Bench. 2018;11:101-9
56. Woo SR, Corrales L, Gajewski TF. Innate immune recognition of cancer. Annu Rev Immunol. 2015;33:445-74
57. Basith S, Manavalan B, Yoo TH, Kim SG, Choi S. Roles of toll-like receptors in cancer: a double-edged sword for defense and offense. Archives of pharmacal research. 2012;35:1297-316
58. Gu J, Liu Y, Xie B, Ye P, Huang J, Lu Z. Roles of toll-like receptors: From inflammation to lung cancer progression. Biomed Rep. 2018;8:126-32
59. Bhatelia K, Singh K, Singh R. TLRs: linking inflammation and breast cancer. Cell Signal. 2014;26:2350-7
60. Liu YD, Ji CB, Li SB, Yan F, Gu QS, Balic JJ. et al. Toll-like receptor 2 stimulation promotes colorectal cancer cell growth via PI3K/Akt and NF-kappaB signaling pathways. Int Immunopharmacol. 2018;59:375-83
61. Xu C, Li H, Yin M, Yang T, An L, Yang G. Osteopontin is involved in TLR4 pathway contributing to ovarian cancer cell proliferation and metastasis. Oncotarget. 2017;8:98394-404
62. El-Omar EM, Ng MT, Hold GL. Polymorphisms in Toll-like receptor genes and risk of cancer. Oncogene. 2008;27:244-52
63. Turner ML, Cronin JG, Healey GD, Sheldon IM. Epithelial and stromal cells of bovine endometrium have roles in innate immunity and initiate inflammatory responses to bacterial lipopeptides in vitro via Toll-like receptors TLR2, TLR1, and TLR6. Endocrinology. 2014;155:1453-65
64. Xu C, Qiao Y, Zhang D. [Expression of Toll-like receptors in human hepatocellular carcinoma and its correlation with inflammasome-associated cytokines]. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi. 2017;33:1182-6
65. Huhta H, Helminen O, Lehenkari PP, Saarnio J, Karttunen TJ, Kauppila JH. Toll-like receptors 1, 2, 4 and 6 in esophageal epithelium, Barrett's esophagus, dysplasia and adenocarcinoma. Oncotarget. 2016;7:23658-67
66. Smith TJ, Yamamoto K, Kurata M, Yukimori A, Suzuki S, Umeda S. et al. Differential expression of Toll-like receptors in follicular lymphoma, diffuse large B-cell lymphoma and peripheral T-cell lymphoma. Exp Mol Pathol. 2010;89:284-90
67. Wild CA, Brandau S, Lindemann M, Lotfi R, Hoffmann TK, Lang S. et al. Toll-like Receptors in Regulatory T Cells of Patients With Head and Neck Cancer. Arch Otolaryngol Head Neck Surg. 2010;136:1253-9
68. Yu H, Xu SS, Cheng QQ, He LM, Li Z. [Expression and clinical significance of Toll-like receptors in human renal carcinoma cell 786-0 and normal renal cell HK-2]. Zhonghua yi xue za zhi. 2011;91:129-31
69. Baretti M, Azad NS. The role of epigenetic therapies in colorectal cancer. Curr Probl Cancer. 2018
70. Lao VV, Grady WM. Epigenetics and colorectal cancer. Nat Rev Gastroenterol Hepatol. 2011;8:686-700
71. El Bairi K, Tariq K, Himri I, Jaafari A, Smaili W, Kandhro AH. et al. Decoding colorectal cancer epigenomics. Cancer Genet. 2018;220:49-76
72. Veltkamp M, Wijnen PA, van Moorsel CH, Rijkers GT, Ruven HJ, Heron M. et al. Linkage between Toll-like receptor (TLR) 2 promotor and intron polymorphisms: functional effects and relevance to sarcoidosis. Clinical and experimental immunology. 2007;149:453-62
73. Liu J, Radler D, Illi S, Klucker E, Turan E, von Mutius E. et al. TLR2 polymorphisms influence neonatal regulatory T cells depending on maternal atopy. Allergy. 2011;66:1020-9
74. Leoratti FM, Farias L, Alves FP, Suarez-Mutis MC, Coura JR, Kalil J. et al. Variants in the toll-like receptor signaling pathway and clinical outcomes of malaria. J Infect Dis. 2008;198:772-80
75. Kohailan M, Alanazi M, Rouabhia M, Alamri A, Parine NR, Alhadheq A. et al. Effect of smoking on the genetic makeup of toll-like receptors 2 and 6. Onco Targets Ther. 2016;9:7187-98
76. Taylor BD, Darville T, Ferrell RE, Ness RB, Haggerty CL. Racial variation in toll-like receptor variants among women with pelvic inflammatory disease. J Infect Dis. 2013;207:940-6
77. Yang H, Zhang X, Geng J, Zheng Z, Fu Q. Toll-like receptor 6 V327M polymorphism is associated with an increased risk of Klebsiella pneumoniae infection. Pediatr Infect Dis J. 2014;33:e310-5
78. Koponen P, Vuononvirta J, Nuolivirta K, Helminen M, He Q, Korppi M. The association of genetic variants in toll-like receptor 2 subfamily with allergy and asthma after hospitalization for bronchiolitis in infancy. Pediatr Infect Dis J. 2014;33:463-6
Author contact
Corresponding author: Dr. Abdelhabib Semlali: Groupe de Recherche en Écologie Buccale, Faculté de Médecine Dentaire, Université Laval, Québec, Québec, Canada. Office +418 6562131#400062. E. mail: abdelhabib.semlaliulaval.ca

 Global reach, higher impact
Global reach, higher impact