Impact Factor ISSN: 1837-9664
J Cancer 2019; 10(4):821-828. doi:10.7150/jca.27899 This issue Cite
Research Paper
KRAS G12V Mutation is an Adverse Prognostic Factor of Chinese Gastric Cancer Patients
1. Guangdong Institute of Gastroenterology, The Sixth Affiliated Hospital, Sun Yat-sen University, Guangdong, China
2. Guangdong Provincial Key Laboratory of Colorectal and Pelvic Floor Diseases, The Sixth Affiliated Hospital, Sun Yat-sen University, Guangdong, China
3. Department of Pathology, The Sixth Affiliated Hospital, Sun Yat-sen University, Guangdong, China
4. Department of Information and Technology, The Sixth Affiliated Hospital, Sun Yat-sen University, Guangdong, China
5. Department of GI Surgery, The Sixth Affiliated Hospital, Sun Yat-sen University, Guangdong, China
* These authors have contributed equally to this work.
Received 2018-6-15; Accepted 2018-12-14; Published 2019-1-29
Abstract
This study aims to investigate the molecular characteristics of Chinese gastric cancer patients. In our study, the KRAS, BRAF, and PIK3CA mutation status of 485 GC patients were analyzed by Sanger sequencing. Kaplan-Meier analysis was used to plot survival curves according to different genotypes. The results show that the frequency of KRAS, BRAF and PIK3CA mutations were 4.1%, 1.2% and 3.5%, respectively. BRAF mutations were significantly concentrated in stage III and IV gastric cancer (P=0.009). KRAS G12V mutation carriers have much shorter OS than other mutation carriers and wild-type group patients (P=0.013). In conclusion, only the KRAS G12V mutation has an adverse effect on patient survival.
Keywords: KRAS, BRAF, PIK3CA, Mutation, Gastric cancer
Introduction
Gastric cancer (GC) is the fifth most common malignancy and the third leading cause of cancer-related death in the world (http://globocan.iarc.fr/Default.aspx) [1]. In China, the incidence of GC is much higher than in any other countries and it is the third most common cancer and the leading cause of death [2]. The geographical differences may partly reflect differences in population-specific genetic risk factors and the prevalence of Helicobacter pylori infection, which plays critical roles in GC pathogenesis [3, 4]. Surgery is the primary treatment for patients with early-stage GC. However, GC is often diagnosed at an advanced stage [5]. In most of the world, GC continues to pose a significant challenge for health care professionals.
In the present, management and prognosis of patients with GC are based entirely on the TNM staging system. However, TNM staging information is not enough for individual treatment and potential targeted therapy. Increased understanding of oncogenic mutations and cell signaling pathways led to the successful application of targeted therapies in various cancers.
The RAS/RAF/MEK and phosphoinositide 3-kinase (PI3K)/Akt/mTOR signaling pathway are key signals that are activated in the different tumors. These pathways are involved in the diverse cellular process, including cell growth, survival and motility [6-8]. Mutations in KRAS, BRAF and PIK3CA in the above pathways have been detected in various malignancies. Identification of these mutations in tumor has predictive value or prognostic value for clinical application. In colorectal cancer, anti-EGFR drugs should only be provided to patients with wild-type KRAS [9]. BRAF mutation has an adverse prognosis in patients with metastatic colorectal cancer [10, 11]. In melanoma with BRAF mutation, vemurafenib therapy improves rates of overall and progression-free survival of melanoma patients [12]. Mutated PIK3CA in colorectal cancer patients indicated low dose aspirin use improves patient's survival [13, 14]. Several studies have partially analyzed KRAS, BRAF and PIK3CA mutation in GC patients [15-18], but the clinical implications of these mutations in GC patients are not addressed. Further investigation for these genetic alterations in GC is required.
In the present study, we analyzed the molecular characteristics of GC in Chinese patients. We accessed the status of KRAS, BRAF and PIK3CA mutations by using Sanger sequencing, and investigated the clinicopathological characteristics and prognostic role of gene mutations in GC patients.
Materials and Methods
Patients
The study retrospectively analyzed 485 GC patients who underwent surgical resection at the Sixth Affiliated Hospital of Sun Yat-sen University from December 2009 to May 2016. All patients underwent informed a consent process approved by the Institutional Review Board of the hospital. The criteria for patient inclusion were: (1) Aged 18-80 years; (2) Primary lesion was pathologically diagnosed as gastric carcinoma; (3) Clinical information, including follow-up data, was completed. The criteria for exclusion were: (1) With a history of other tumors or hematological malignancy; (2) Accompanied with severe infection, severe kidney dysfunction, or severe hepatic dysfunction; (3) Accepted preoperational chemotherapy, radiotherapy, targeted therapy, and immunotherapy. Formalin-fixed, paraffin-embedded tumor tissues were obtained. Clinical data was collected. The study protocol was approved by the Institute Research Medical Ethics Committee of Sun Yat-sen University. Overall survival (OS) was defined as the time from the beginning of surgical resection to death or last follow-up.
KRAS, BRAF and PIK3CA mutation analysis
Assessment of KRAS, BRAF and PIK3CA mutation was performed in the Molecular Diagnostic Laboratory of the Sixth Affiliated Hospital of Sun Yat-sen University, using an adequate quality-control procedure. All tissue samples were formalin-fixed paraffin-embedded and histologically confirmed. Genomic DNA from analyzed samples was extracted with Hipure FFPE DNA Kit (Cat No: D3126-02, Magen, China) according to the manufacturer's protocol. Exon 2 (codon 12 and 13) of KRAS, exon 9 (codon 542 and 545) and exon 20 (codon 1047) of PIK3CA, and exon 15 (codon 600) of BRAF were assessed.
The prior PCR amplification was performed on an ABI 9700 PCR system. Amplification was done in 20μL reaction contain 50-100ng of DNA template and 500nM primers, with the following program: 5min at 98°C for initial denaturation followed by 45cyclers of 25sec at 95°C, 25 sec at 58°C and 25 sec at 72°C, and a final extension at 72°C for 10 min. The primers were listed in Table 1. PCR products were purified, sequenced by using BigDye Terminator v3.1 Sequencing Standard Kit (Thermo Fisher Scientific, USA) with an ABI Prism 3500Dx genetic Analyzer (Applied Biosystems, Foster City, CA).
Primers and conditions of Sanger sequencing.
| Gene | Primers | Length (bp) | Conditions |
|---|---|---|---|
| KRAS | 98°C 5min, (95°C 25sec, 58°C 25 sec, 72°C 25 sec ) 45 cycles, 72°C 10min | ||
| Exon 2 | F: ATGTTCTAATATAGTCACATTTTC | 202 | |
| R: GTCCTGCACCAGTAATATGC | |||
| BRAF | |||
| Exon 15 | F: TCATAATGCTTGCTCTGATAGGA | 224 | |
| R: GGCCAAAAATTTAATCAGTGGA | |||
| PIK3CA | |||
| Exon 9 | F: ATCCAGAGGGGAAAAATATG | 194 | |
| R: TTAGCACTTACCTGTGACTC | |||
| Exon 20 | F: CGAAAGACCCTAGCCTTAGAT | 215 | |
| R: GTCTTTGCCTGCTGAGAGTTATT | |||
Statistical analysis
Associations of KRAS/BRAF/PIK3CA mutation status with demographic and clinical characteristics were evaluated using continuous variables, categorical data analysis. Statistical analyses were performed with SPSS software (SPSS, Chicago, IL, USA). Statistical analysis for Kaplan-Meier survival curves for OS was performed using GraphPad Prism 5 (Graph Pad Software Inc., San Diego, CA, USA). A two-sided probability value of less than 0.05 was considered to be statistically significant.
Results
Patient characteristics
Table 2 summarizes the clinicopathological characteristics of study subjects. Of these 485 patients, males were over twice females (68.0% vs. 32.0%). A majority of patients (79.1%) were older than 50 at diagnosis. Most patients (65.2%) had stage III or stage IV tumor. Nearly half of the tumors (41.4%) were located in the lower gastric.
Clinicpathological characteristics of 485 GC patients.
| Characteristics | Frequency %(n) |
|---|---|
| Gender | |
| Male | 68.0 (330) |
| Female | 32.0 (155) |
| Age, years | |
| <45 | 12.2 (59) |
| 45-49 | 8.7 (42) |
| 50-70 | 59.5 (289) |
| ≥70 | 19.6 (95) |
| TNM stage | |
| I | 14.0 (68) |
| II | 20.8 (101) |
| III | 40.0 (194) |
| IV | 25.2 (122) |
| Tumor location | |
| Upper | 30.3 (147) |
| Middle | 18.8 (91) |
| Lower | 41.4 (201) |
| Residual gastric or total gastric | 9.5 (46) |
KRAS, BRAF and PIK3CA mutations, and their correlations with patient characteristics
The mutation rate of KRAS was 4.1% (20 out of 485). Five different substitutions were detected, including G13D (n=6), G12S (n=3), G12D (n=5), G12V (n=5) and G12A (n=1). Six BRAF V600E was detected, which was KRAS wild-type. The mutation rate of PIK3CA was 3.5% (17 out of 485). Among 17 patients, 10 carried mutations within exon 9 and 7 carried mutations within exon 20. Mutation types identified in exon 9 included E542K (n=5), E545K (n=4), Q546R (n=1), whereas H1047R accounted for all the mutations in exon 20. One patient was identified with concomitant PIK3CA mutation (E545K) and KRAS mutation (G13D).
The associations of patients' clinicopathological characteristics and KRAS, BRAF, PIK3CA gene mutations were summarized in Table 3. BRAF mutations were significantly concentrated in stage III and IV gastric cancer (P=0.009). The associations of patients' clinicopathological characteristics and different KRAS mutations were summarized in Table 4. KRAS G12V mutation was associated with female (P=0.038). KRAS G12D mutation was significantly correlated with tumor location (P=0.020) and lymph node status (P=0.045).
Associations between gene mutations and clinicpathological characteristics of patients.
| KRAS | P -value | PIK3CA | P -value | BRAF | P -value | ||||
|---|---|---|---|---|---|---|---|---|---|
| Mutation | Wild type | Mutation | Wild type | Mutation | Wild type | ||||
| n (%) | n (%) | n (%) | n (%) | n (%) | n (%) | ||||
| Gender | |||||||||
| Male | 12 (60.0) | 318 (68.4) | 0.432 | 12 (66.7) | 318 (68.1) | 1.000 | 3 (50.0) | 327 (68.3) | 0.341 |
| Female | 8 (40.0) | 147 (31.6) | 6 (33.3) | 149 (31.9) | 3 (50.0) | 152 (31.7) | |||
| Age, years | |||||||||
| <45 | 2 (10.0) | 57 (12.3) | 0.933 | 4 (22.2) | 55 (11.8) | 0.332 | 1 (16.7) | 58 (12.1) | 0.572 |
| 45-49 | 2 (10.0) | 40 (8.6) | 1 (5.6) | 41 (8.8) | 0 | 42 (8.8) | |||
| 50-70 | 13 (65.0) | 276 (59.4) | 10 (55.6) | 279 (59.7) | 5 (83.3) | 284 (59.3) | |||
| ≥70 | 3 (15.0) | 92 (19.7) | 3 (16.6) | 92 (19.7) | 0 | 95 (19.8) | |||
| TNM stage | |||||||||
| I | 1(5.0) | 67 (14.4) | 0.525 | 3 (16.7) | 65 (13.9) | 0.179 | 0 | 68 (14.2) | 0.009 |
| II | 7 (35.0) | 94 (20.2) | 6 (33.3) | 95 (20.3) | 0 | 101 (21.1) | |||
| III | 5 (25.0) | 189 (40.6) | 7 (38.9) | 187 (40.0) | 1 (16.7) | 193 (40.3) | |||
| IV | 7 (35.0) | 115 (24.8 ) | 2 (11.1) | 120 (25.8) | 5 (83.3) | 117 (24.4) | |||
| Tumor location | |||||||||
| Upper | 7 ( 35.0) | 140 (30.1) | 0.646 | 5 (27.8) | 142 (30.4) | 0.634 | 3 (50.0) | 144 (30.1) | 0.937 |
| Middle | 6 (30.0) | 85 (18.3) | 6 (33.3) | 85 (18.2) | 0 | 91 (19.0) | |||
| Lower | 3 (15.0) | 198 (42.6) | 6 (33.3) | 195 (41.8) | 1 (16.7) | 200 (41.8) | |||
| Residual gastric or total gastric | 4 (20.0) | 42 (9.0) | 1 (5.6) | 45 (9.6) | 2 (33.3) | 44 (9.1) | |||
| Lymph node status | |||||||||
| pN0 | 10 (50.0) | 138 ( 29.7) | 0.139 | 22.2(4) | 144 (30.8) | 0.556 | 0 | 148 (30.9) | 0.320 |
| pN1 | 5 (25.0) | 132 ( 28.4) | 38.9(7) | 130 (27.8) | 3 (50.0) | 134 (28.0) | |||
| pN2 | 2 (10.0) | 139 (29.9) | 16.7(3) | 138 (29.6) | 2 (33.3) | 139 (29.0) | |||
| pN3a/b | 3 (15.0) | 56 (12.0) | 22.2(4) | 55 (11.8) | 1 (16.7) | 58 (12.1) | |||
Associations between different mutations of KRAS and clinicpathological characteristics of patients.
| KRAS G12V mutation status | P - value | KRAS G12S mutation status | P- value | KRAS G12D mutation status | P- value | KRAS G12A mutation status | P- value | KRAS G13D mutation status | P- value | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mutationn (%) | Wild typen (%) | Mutationn (%) | Wild typen (%) | Mutationn (%) | Wild typen (%) | Mutationn (%) | Wild typen (%) | Mutationn (%) | Wild typen (%) | ||||||
| Gender | |||||||||||||||
| Male | 1 (20.0) | 329 (68.5) | 0.038 | 3 (100 ) | 327 (67.8) | 0.555 | 3 (60.0) | 327 (68.1) | 1.000 | 0 | 330 (68.2) | 0.320 | 5 (83.3) | 325 (67.8) | 0.670 |
| Female | 4 (80.0) | 151 (31.5) | 0 | 155 (32.2) | 2 (40.0) | 153 (31.9) | 1 (100.0) | 154 (31.8) | 1 (16.7) | 154 (32.2) | |||||
| Age, years | |||||||||||||||
| <45 | 1 (20.0) | 58 (12.1) | 1.000 | 0 | 59 (12.2) | 0.272 | 0 | 59 (12.3) | 0.300 | 0 | 59 (12.2) | 1.000 | 1 (16.7) | 58 (12.1) | 1.000 |
| 45-49 | 0 | 42 (8.8) | 1 (33.3) | 41 ( 8.5) | 0 | 42 (8.8) | 0 | 42 (8.7) | 1 (16.7) | 41 (8.6) | |||||
| 50-70 | 3 (60.0) | 286 (59.6) | 1 (33.3) | 288 (59.8) | 5(100) | 284 (59.2) | 1 (100.0) | 288 (59.5) | 3 (50.0) | 286 (59.7) | |||||
| >70 | 1 (20.0) | 94 (19.5) | 1 (33.3) | 94 (19.5) | 0 | 95 (19.7) | 0 | 95 (19.6) | 1 (16.7) | 94 (19.6) | |||||
| TNM stage | |||||||||||||||
| I | 1 (20.0) | 67 (14.0) | 0.928 | 0 | 68 (14.1) | 0.385 | 0 | 68 (14.2) | 0.065 | 0 | 68 (14.0) | 1.000 | 0 | 68 (14.2) | 0.303 |
| II | 1 (20.0) | 100 (20.8) | 0 | 101 (21.0) | 3 (60.0) | 98 (20.4) | 0 | 101 (20.9) | 3 (50.0) | 98 (20.5) | |||||
| III | 1 (20.0) | 193 (40.2) | 1 (33.3) | 193 (40.0) | 0 | 194 (40.4) | 1 (100.0) | 193 (39.9) | 2 (33.3) | 192 (40.1) | |||||
| IV | 2 (40.0) | 120 (25.0) | 2 (66.7) | 120 (24.9) | 2 (40.0) | 120 (25.0) | 0 | 122 (25.2) | 1 (16.7) | 121 (25.3) | |||||
| Tumor location | |||||||||||||||
| Upper | 2 (40.0) | 145 (30.2) | 1.000 | 1 (33.3) | 146 (30.3) | 0.322 | 1 (20.0) | 146 (30.4) | 0.020 | 1 (100.0) | 146 (30.2) | 0.586 | 2 (33.3) | 145 (30.3) | 0.557 |
| Middle | 1 (20.0) | 90 (18.8) | 1 (33.3) | 90 (18.7) | 2 (40.0) | 89 (18.5) | 0 | 91 (18.8) | 2 (33.3) | 89 (18.6) | |||||
| Lower | 2 (40.0) | 199 (41.5) | 0 | 201 (41.7) | 0 | 201 (41.9) | 0 | 201 (41.5) | 1 (16.7) | 200 (41.8) | |||||
| Residual gastric or total gastric | 0 | 46 (9.5) | 1 (33.3) | 45 (9.3) | 2 (40.0) | 44 (9.2) | 0 | 46 (9.5) | 1 (16.7) | 42 (9.4) | |||||
| Lymph node status | |||||||||||||||
| pN0 | 3 (60.0) | 145 (30.2) | 0.562 | 0 | 148 (30.7) | 0.144 | 4 (80.0) | 144 (30.0) | 0.045 | 0 | 148 (30.6) | .122 | 3 (50.0) | 145 (30.3) | 0.664 |
| pN1 | 1 (20.0) | 136 (28.3) | 2 (66.7) | 135 (28.0) | 0 | 137 (28.5) | 0 | 137 (28.3) | 2 (33.3) | 135 (28.2) | |||||
| pN2 | 1 (20.0) | 140 (29.2) | 0 | 141 (29.3) | 0 | 141 (29.4) | 1 (100.0) | 141 (29.1) | 1 (16.7) | 140 (29.2) | |||||
| pN3a/b | 0 | 59 (12.3) | 1 (33.3) | 58 (2.0) | 1 (20.0) | 58 (12.1) | 0 | 58 (12.0) | 0 | 59 (12.3) | |||||
Survival analysis
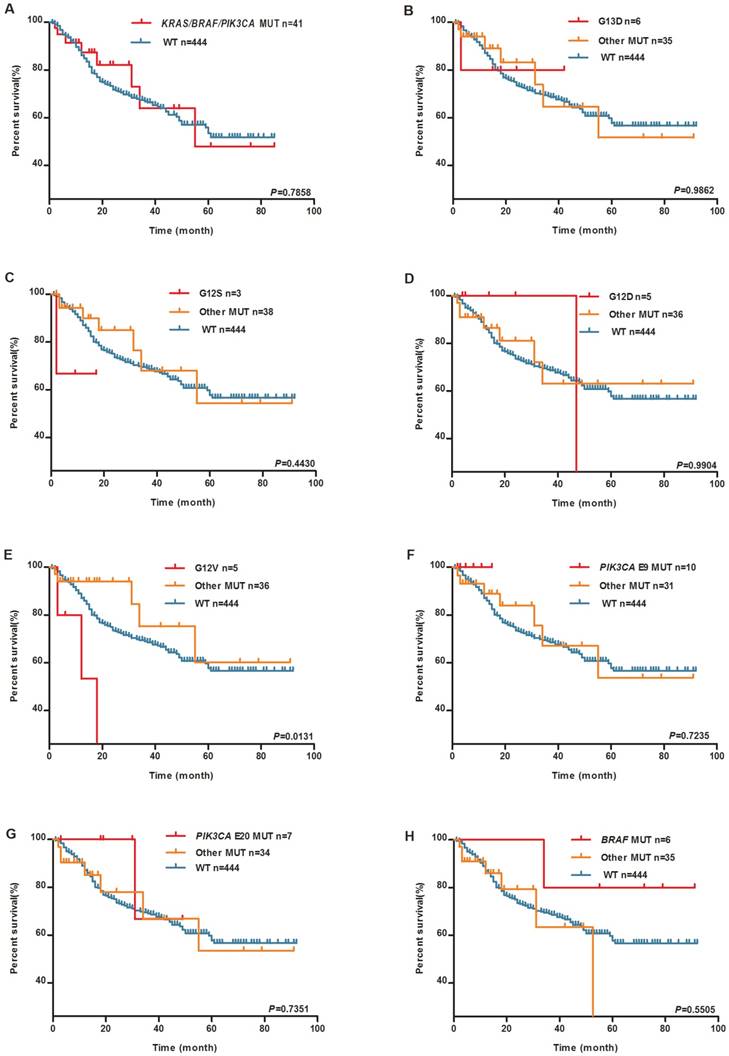
Kaplan-Meier survival analysis was performed to clarify the prognostic effect of these mutations on the GC patients. During the follow-up, 136 patients died. No significant differences were reported between patients with and without any mutation of KRAS, BRAF, and PIK3CA in the survival analysis (The median survival time in mutation group is 55 months, the median survival time of wild-type group does not reach, log-rank P=0.7858, Figure 1A). Individual KRAS mutation types were further examined. As shown in Figure 1E, KRAS G12V mutation carriers experienced much shorter OS than wild-type group patients (The median survival time in KRAS G12V mutation group is 18 months, the median survival time of other mutation group and wild-type group has not reached, log-rank P=0.0131). As shown in Figure 2, there was no significant difference in OS among the patients with different location of the tumor.
KRAS G12V is associated with worse patient survival. Kaplan-Meier plots of overall survival (OS) for GC patients by tumor KRAS, BRAF and PIK3CA mutations. (A) KRAS/BRAF/PIK3CA mutation and WT groups. (B) KRAS G13D mutation, (C) KRAS G12S mutation, (D) KRAS G12D mutation, (E) KRAS G12V mutation, (F) PIK3CA exon9 mutation, (G) PIK3CA exon20 mutation, (H) BRAF V600E mutation, other mutation and WT groups.
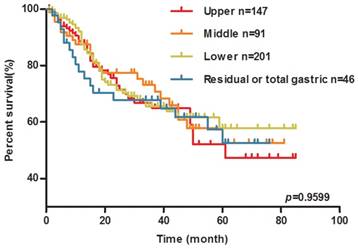
Tumor location has no effect on the OS of GC patients. Kaplan-Meier plots of overall survival (OS) for GC patients by tumor location (upper group/middle group / lower group and residual or total gastric group).
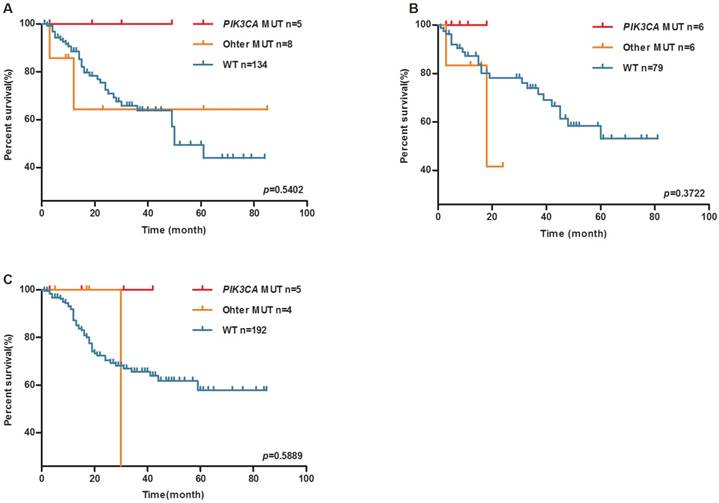
Whereas, PIK3CA mutation was associated with a trend towards longer OS in upper (The median survival time in wild-type group is 50 months, the median survival time of PIK3CA mutation group and other mutation group have not reached, log-rank P=0.5402, Figure 3A), middle (The median survival time in other mutation group is 18 months, median survival time of PIK3CA mutation group and wild-type group have not reached, log-rank P=0.3722, Figure 3B) and lower (The median survival time in other mutation group is 34 months, median survival time of PIK3CA mutation group and wild-type group does not reach, log-rank P=0.5889, Figure 3C) gastric tumors.
PIK3CA mutation may be a favorable prognosis marker of the OS of upper, middle and lower GC patients. Kaplan-Meier plots of overall survival (OS) for GC patients by PIK3CA mutation (A) upper GC patients, (B) middle GC patients, (C) lower GC patients.
Discussion
Of these 485 GC patients in the study, males were over twice females. The mutations in KRAS exon2 (codons 12, 13), BRAF codon 600 and PIK3CA exon 9/20 (codons 542, 545 and 1047) were not frequent in Chinese GC patients.
KRAS is a downstream effector of EGFR. Activating mutation of KRAS is thought to stimulate the RAS/RAF/MEK/signaling pathway independent of EGFR activation. We found the overall KRAS mutation rate was 4.1%, which were close to most of the previous studies in China and Japan (4% - 4.9% in Japan and 4.5% in China) [15, 16, 18]. In patients and animal models, the malign function of KRAS G12V mutation has been identified in other tumors, such as colorectal cancer, lung cancer, and pancreatic cancer by multiple techniques [16, 19-25]. As to gastric cancer, we found KRAS G12V is a poor prognostic marker, which has not been reported before. Moreover, our cohort is the biggest one in the related reports [15, 26], which can more effectively reflect the southern China GC patients' molecular profile. Also, we found tumor locations were not associated with the overall survival of GC patients or with mutations of KRAS, BRAF, and PIK3CA. KRAS is a key biomarker for predicting response to anti-EGFR therapy in colorectal cancer[11, 27] .Some phase III trials on addition of cetuximab or panitumumab to system chemotherapy reported that in advanced GC or oesophagogastric cancer, the anti-EGFR antibodies provided no additional benefit[28, 29]. Thus, more evidence is needed to elucidate KRAS mutations' predictive value to GC.
BRAF is a downstream effector of KRAS, its prognostic value for colorectal cancer is wildly accepted [10, 11]. In GC, the frequency of BRAF mutation is very low. Large-scale trials are needed to test its clinical value in GC.
PIK3CA, encodes the p110 catalytic submit of PI3K, frequently mutated in some human tumors [30]. Exon 9 and exon 20 are two PIK3CA mutational hotspots that affected the helical and catalytic protein domains, respectively. The PI3K/AKT/mTOR pathway has a close association with the RAS/RAF/MEK/signaling pathway, that active RAS can interactive catalytic submit of PI3K and lead its activation in the regulation of cellular functions [31].
Previous studies indicated that the mutation rate of PIK3CA in GC was 3.8% to 12% [15, 16, 32-35]. In our analysis, 3.5% of Chinese GC patients had PIK3CA mutations. It has been reported that PIK3CA mutation was frequently concomitant with KRAS or BRAF mutations in a wide variety of tumors especially in colorectal cancer [30, 36-38]. In GC, simultaneous mutations in PIK3CA and KRAS were observed in rare cases [16, 33]. In our study, only one patient had concomitant PIK3CA mutation (E545K) and KRAS mutation (G13D). Among the 17 PIK3CA mutation carries, only one patient died whose tumor located in the residual gastric. The other PIK3CA mutation carriers may have longer OS than wild-type group patients.
To validate our main conclusion, we analyzed clinical and somatic mutations data of the TCGA database on LinkedOmics website (http://www.linkedomics.org). However, the result showed that there is no significant association between KRAS G12V mutation and patients' survival (P=0.87, Supplementary Figure 1) [39]. The deviation between our cohort and TCGA cohort may be due to the small sample of KRAS G12V mutated patients in the database and the genetic background difference between Asian people and western people. Further validation will be carried out in our center.
As to the tumor stage of diagnosed GC patients, we found only 14% of patients were diagnosed at stage I. The overwhelming majority of patients were diagnosed at advanced stage. Considering 8.7% of the GC patient were aged between 45 years to 49 years, we recommend that individuals of 45 years age and older should be screened for gastric cancer.
There are several limitations to the studies. First, the small sample size was the major limitation. We could not adequately evaluate the prognostic impact of each gene mutation on GC patients with this small cohort. Second, the study population was collected from a single center. The genetic variations we observed may mainly reflect the signatures of southern China. Insufficient event outcome was the third limitation. More significant finding needs longer follow-up time in the future.
Overall, we found the frequency of KRAS, BRAF and PIK3CA mutations in GC patients were 4.1%, 1.2%, and 3.5%, respectively. Meanwhile, we found KRAS G12V is an adverse prognostic factor for gastric cancer patients.
Supplementary Material
Supplementary figure.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (grant number 2017YFC1308800), National Natural Science Foundation of China (grant number 81201545, 81201581), Young Teacher Training Program of Sun Yat-sen University (grant number 14YKPY31),Science and Technology Planning Project of Guangdong Province (grant number 2012B031800355), “985” Project of Sun Yat-sen University (grant number 4202037), National Basic Research Program of China (grant number 2015CB554001), China Scholarship Council (grant number 201706385049), the frontier and key technology innovation project of Guangdong Province (grant number 2014B010118003), the Science and Technology Planning Project of Guangdong Province (grant number 2015B010129008), and National Key Clinical Discipline.
Compliance with ethical standards
Ethical approval: All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent: Informed consent was obtained from all individual participants included in the study.
Competing Interests
The authors have declared that no competing interest exists.
References
1. World Health Organization. GLOBOCAN2012: Estimated Cancer Incidence, Mortality and Prevalence Worldwide in 2012. http://globocan.iarc.fr/Default.aspx
2. Chen W, Zheng R, Zeng H. et al. Annual report on status of cancer in China, 2011. Chin J Cancer Res. 2015;27:2-12
3. Arnold M, Karim-Kos HE, Coebergh JW. et al. Recent trends in incidence of five common cancers in 26 European countries since 1988: Analysis of the European Cancer Observatory. Eur. J. Cancer. 2015;51:1164-1187
4. Uemura N, Okamoto S, Yamamoto S. et al. Helicobacter pylori infection and the development of gastric cancer. N Engl J Med. 2001;345:784-789
5. Kim HJ, Karpeh MS. Surgical approaches and outcomes in the treatment of gastric cancer. Semin. Radiat. Oncol. 2002;12:162-169
6. Qu JL, Qu XJ, Zhao MF. et al. Gastric cancer exosomes promote tumour cell proliferation through PI3K/Akt and MAPK/ERK activation. Dig Liver Dis. 2009;41:875-880
7. Friday BB, Adjei AA. Advances in targeting the Ras/Raf/MEK/Erk mitogen-activated protein kinase cascade with MEK inhibitors for cancer therapy. Clin. Cancer Res. 2008;14:342-346
8. Yu HG, Ai YW, Yu LL. et al. Phosphoinositide 3-kinase/Akt pathway plays an important role in chemoresistance of gastric cancer cells against etoposide and doxorubicin induced cell death. Int. J. Cancer. 2008;122:433-443
9. Deng Y, Wang L, Tan S. et al. KRAS as a predictor of poor prognosis and benefit from postoperative FOLFOX chemotherapy in patients with stage II and III colorectal cancer. Mol. Oncol. 2015;9:1341-1347
10. Bokemeyer C, Van Cutsem E, Rougier P. et al. Addition of cetuximab to chemotherapy as first-line treatment for KRAS wild-type metastatic colorectal cancer: pooled analysis of the CRYSTAL and OPUS randomised clinical trials. Eur. J. Cancer. 2012;48:1466-1475
11. Van Cutsem E, Kohne CH, Lang I. et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J. Clin. Oncol. 2011;29:2011-2019
12. Chapman PB, Hauschild A, Robert C. et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med. 2011;364:2507-2516
13. Liao X, Lochhead P, Nishihara R. et al. Aspirin use, tumor PIK3CA mutation, and colorectal-cancer survival. N Engl J Med. 2012;367:1596-1606
14. Bibbins-Domingo K. Aspirin Use for the Primary Prevention of Cardiovascular Disease and Colorectal Cancer: U.S. Preventive Services Task Force Recommendation Statement. Ann. Intern. Med. 2016;164:836-845
15. Lu W, Wei H, Li M. et al. Identification of KRAS and PIK3CA but not BRAF mutations in patients with gastric cancer. Mol. Med. Rep. 2015;12:1219-1224
16. Takahashi N, Yamada Y, Taniguchi H. et al. Clinicopathological features and prognostic roles of KRAS, BRAF, PIK3CA and NRAS mutations in advanced gastric cancer. BMC Res Notes. 2014;7:271
17. Lee SH, Lee JW, Soung YH. et al. BRAF and KRAS mutations in stomach cancer. Oncogene. 2003;22:6942-6945
18. van Grieken NC, Aoyama T, Chambers PA. et al. KRAS and BRAF mutations are rare and related to DNA mismatch repair deficiency in gastric cancer from the East and the West: results from a large international multicentre study. Br J Cancer. 2013;108:1495-1501
19. Alamo P, Gallardo A, Di Nicolantonio F. et al. Higher metastatic efficiency of KRas G12V than KRas G13D in a colorectal cancer model. Faseb J. 2015;29:464-476
20. Fiala O, Buchler T, Mohelnikova-Duchonova B. et al. G12V and G12A KRAS mutations are associated with poor outcome in patients with metastatic colorectal cancer treated with bevacizumab. Tumour Biol. 2016;37:6823-6830
21. Olmedillas LS, Garcia-Olmo DC, Garcia-Arranz M. et al. KRAS G12V Mutation Detection by Droplet Digital PCR in Circulating Cell-Free DNA of Colorectal Cancer Patients. Int. J. Mol. Sci. 2016;17:484
22. Raub CB, Lee CC, Shibata D. et al. HistoMosaic Detecting KRAS G12V Mutation Across Colorectal Cancer Tissue Slices through in Situ PCR. Anal. Chem. 2016;88:2792-2798
23. Renaud S, Falcoz PE, Schaeffer M. et al. Prognostic value of the KRAS G12V mutation in 841 surgically resected Caucasian lung adenocarcinoma cases. Br J Cancer. 2015;113:1206-1215
24. Chaft JE, Litvak A, Arcila ME. et al. Phase II study of the GI-4000 KRAS vaccine after curative therapy in patients with stage I-III lung adenocarcinoma harboring a KRAS G12C, G12D, or G12V mutation. Clin. Lung Cancer. 2014;15:405-410
25. Provost E, Bailey JM, Aldrugh S. et al. The tumor suppressor rpl36 restrains KRAS(G12V)-induced pancreatic cancer. Zebrafish. 2014;11:551-559
26. Amado RG, Wolf M, Peeters M. et al. Wild-type KRAS is required for panitumumab efficacy in patients with metastatic colorectal cancer. J. Clin. Oncol. 2008;26:1626-1634
27. Lordick F, Kang YK, Chung HC. et al. Capecitabine and cisplatin with or without cetuximab for patients with previously untreated advanced gastric cancer (EXPAND): a randomised, open-label phase 3 trial. Lancet Oncol. 2013;14:490-499
28. Waddell T, Chau I, Cunningham D. et al. Epirubicin, oxaliplatin, and capecitabine with or without panitumumab for patients with previously untreated advanced oesophagogastric cancer (REAL3): a randomised, open-label phase 3 trial. Lancet Oncol. 2013;14:481-489
29. Ihle NT, Byers LA, Kim ES. et al. Effect of KRAS oncogene substitutions on protein behavior: implications for signaling and clinical outcome. J Natl Cancer Inst. 2012;104:228-239
30. Samuels Y, Wang Z, Bardelli A. et al. High frequency of mutations of the PIK3CA gene in human cancers. Science. 2004;304:554
31. Katso R, Okkenhaug K, Ahmadi K. et al. Cellular function of phosphoinositide 3-kinases: implications for development, homeostasis, and cancer. Annu Rev Cell Dev Biol. 2001;17:615-675
32. Shi J, Yao D, Liu W. et al. Highly frequent PIK3CA amplification is associated with poor prognosis in gastric cancer. BMC Cancer. 2012;12:50
33. Li VS, Wong CW, Chan TL. et al. Mutations of PIK3CA in gastric adenocarcinoma. BMC Cancer. 2005;5:29
34. Janku F, Hong DS, Fu S. et al. Assessing PIK3CA and PTEN in early-phase trials with PI3K/AKT/mTOR inhibitors. Cell Rep. 2014;6:377-387
35. Harada K, Baba Y, Shigaki H. et al. Prognostic and clinical impact of PIK3CA mutation in gastric cancer: pyrosequencing technology and literature review. BMC Cancer. 2016;16:400
36. Hsieh LL, Er TK, Chen CC. et al. Characteristics and prevalence of KRAS, BRAF, and PIK3CA mutations in colorectal cancer by high-resolution melting analysis in Taiwanese population. Clin. Chim. Acta. 2012;413:1605-1611
37. Guedes JG, Veiga I, Rocha P. et al. High resolution melting analysis of KRAS, BRAF and PIK3CA in KRAS exon 2 wild-type metastatic colorectal cancer. BMC Cancer. 2013;13:169
38. Janku F, Lee JJ, Tsimberidou AM. et al. PIK3CA mutations frequently coexist with RAS and BRAF mutations in patients with advanced cancers. PLoS One. 2011;6:e22769
39. Vasaikar SV, Straub P, Wang J. et al. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2018;46:D956-D963
Author contact
Corresponding authors: Jian-Ping Wang, Email: wangjpgzcom; Phone: 0086-020-38255495; Address: 26 Er-Heng Road, Yuan-Cun,Tian-He District, Guangzhou 510655, Guangdong, China. Lei Wang, Email: wangl9sysu.edu.cn; Phone: 0086-020-38767131;Address: 26 Er-Heng Road, Yuan-Cun, Tian-He District, Guangzhou 510655, Guangdong, China

 Global reach, higher impact
Global reach, higher impact