Impact Factor ISSN: 1837-9664
J Cancer 2018; 9(21):3950-3961. doi:10.7150/jca.25967 This issue Cite
Research Paper
CDK5 Functions as a Tumor Promoter in Human Lung Cancer
1. Department of Respiratory Medicine, Shanghai Tenth People's Hospital, Tongji University, Shanghai 200072, PR China;
2. Department of Biological Sciences, Arkansas State University, State University, AR 72467, USA;
3. Molecular Biosciences Program, Arkansas State University, State University, AR 72467, USA.
# These authors have contributed equally to this work.
Received 2018-3-9; Accepted 2018-8-19; Published 2018-10-10
Abstract
Cyclin-dependent kinase 5 (CDK5), an atypical member of the cyclin-dependent kinase family, plays an important role in the nervous system. Recent studies have shown that CDK5 is also associated with tumors. However, few studies have been done to investigate the mechanism underlying the connection between CDK5 and cancers. To explore the role of CDK5 in cancers by using an extensive bioinformatics data mining process. We mined the transcriptional, survival, functions and structure of CDK5 gene through databases and in vitro experiments. We found that higher CDK5 expression levels in most cancer cell lines while lower expression in liver and brain cancer cell lines. High expression of CDK5 was associated with shorter overall survival (OS) in lung cancer. In addition, high expression level of CDK5 promoted lung cancer cells proliferation and metastasis. Inhibited CDK5 decreases CAP1 phosphorylation. CDK5 may prove to be a valid target of anticancer therapies.
Keywords: CDK5, data mining, overall survival, tumor promoter, lung cancer
Introduction
Cancer is a critical cause of death globally [1]. Although standard chemotherapy and better supportive care have improved the length of survival and quality of life, the prognosis of advanced cancer patients remains poor. Thus, it is important to investigate underlying mechanisms of cancer tumorigenesis and tumor progression, and identify potential prognostic biomarkers that could even be used as drug targets. Deregulation of cell cycle is a fundamental process that underlies cancer proliferation [2]. Progression through cell cycle is regulated by coordinated actions of CDKs, which including CDK5 as the major one [3].
CDK5 is considered as a neuron-specific kinase in the past decade due to the abundant existence of its activator p35 in post-mitotic neurons [4]. Increasing evidence has demonstrated that the karyoplasm localization of CDK5 is important for its multiple pathological and physiological functions, including induction of cell motility, apoptosis, cell cycle progression, neuronal migration, neuronal cell survival, lymphatic system, vascularization, and insulin secretion [5-13]. Recent studies have shown that CDK5 also participates in a series of biological and pathological processes in non-neuronal cells, and is generally dysregulated in various cancer cells [14].
In this study, we first explored the role of CDK5 in cancers by using an extensive bioinformatics data mining process to find out the expression of CDK5 miRNA in different cancers, and then predict the connection between the expression level and OS. The protein network of predicted associations for CDK5 and alterations in cancer genomics were analyzed. Some experiments were performed to validate the reliance of database and the connection between CDK5 and lung cancer.
Materials and Methods
Oncomine database analysis
The transcription level of CDK5 gene in different cancers was identified by Oncomine database (https://www.oncomine.org/resource/main.html) [15]. The mRNA expression level in cancer tissue compared to the normal was obtained as the parameters of p-value<1E-4, fold change>2 and top gene rank 10%, the analyses were summarized in Table 1.
CDK5 expression in lung cancer
| Cancer subtype | P-value | Fold change | Rank (10%) | Sample | Reference |
|---|---|---|---|---|---|
| Lung Carcinoid Tumor | 2.72E-7 | 7.946 | 10 | 37 | [16] |
| Large Cell Lung Carcinoma | 1.00E-8 | 2.122 | 10 | 84 | [17] |
| Lung Adenocarcinoma | 2.30E-7 | 2.168 | 10 | 39 | [18] |
| Lung Adenocarcinoma | 9.26E-6 | 2.203 | 10 | 57 | [19] |
PrognoScan database analysis
The relationship between CDK5 expression and survival in various types of cancers was analyzed by PrognoScan database (http://www.prognoscan.org/) [20]. The threshold was set as cox p-value<0.05 and the analyses were summarized in Fig. 2, Table 2 and Table S2.
CDK5 gene in lung, skin and blood cancer (Prognoscan Database). (A, B) Lung cancer, (C) skin cancer, (D) blood cancer was plotted from PrognoScan database as the threshold of cox p-value < 0.05. The survival curve comparing the patient with high (red) and low (blue) expression was plotted from PrognoScan database.
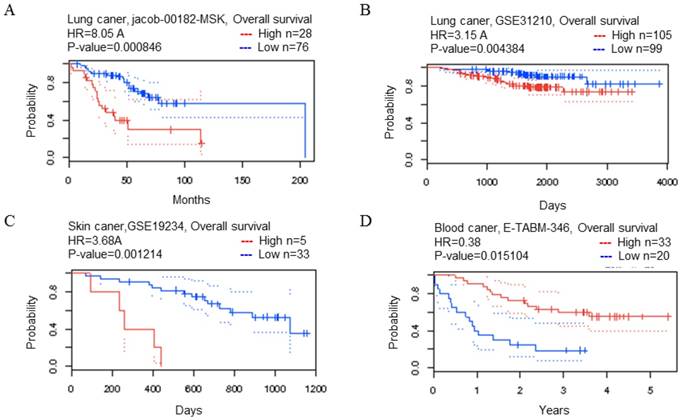
The relationship between transcription level of CDK5 and OS in cancer patients
| Cancer type | N | Cox p-value | HR | Endpoint | Dataset | Probe ID |
|---|---|---|---|---|---|---|
| Lung | 104 | 0.000846 | 2.09 | OS | HG_U133A | 204247_S_at |
| 204 | 0.004384 | 1.15 | OS | HG_U133_Plus_2 | 204247_S_at | |
| Skin | 38 | 0.001214 | 1.30 | OS | HG_U133_Plus_2 | 204247_S_at |
| Blood | 53 | 0.015104 | -0.97 | OS | HG_U133A | 204247_S_at |
Kaplan-Meier plotter database analysis
The Kaplan Meier plotter is capable to assess the effect of 54,675 genes on survival using 10,461 cancer samples. These include 5,143 breast, 1,816 ovarian, 2,437 lung and 1,065 gastric cancer patients. The correlation of CDK5 expression and survival in breast, gastric, ovarian and lung cancer was analyzed by Kaplan-Meier plotter database (http://kmplot.com/analysis/) [21] and the result was showed in Fig. 3. The hazard ratio with 95% confidence intervals and log-rank p-value was also computed.
CDK5 gene in ovarian, breast, lung, gastric cancer (Kaplan-Meier Plotter). (A) Ovarian, (B) Breast, (C) Lung, (D) Gastric cancer. The survival curve comparing the patient with high (red) and low (black) expression was plotted from Kaplan-Meier Plotter database. The p-value was 0.73, 0.43, 0.0013 and 0.32 respectively.
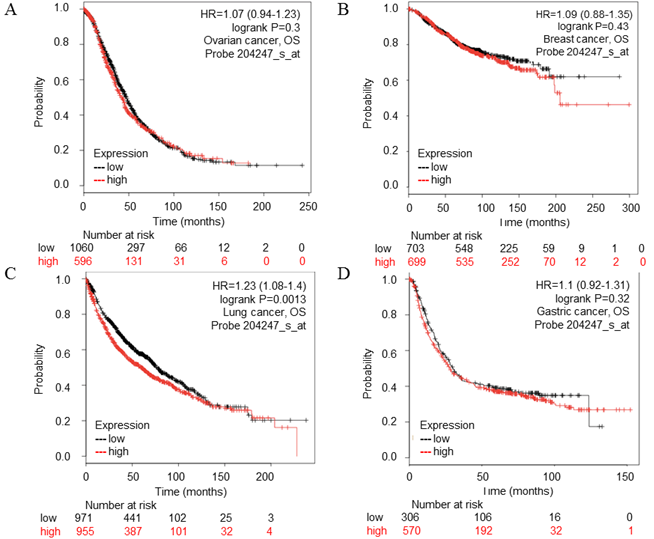
STRING database analysis
We analyzed the functional protein association network by STRING database (https://string-db.org/) [22]. Protein name: CDK5 and organism; homo sapiens were analyzed. The results showed in Fig. 4.
Identification of known and predicted structural proteins essential for CDK5 function (STRING). Interacting nodes are displayed in colored circles using String. Predicted functional partners of CDK5 are shown based upon peer reviewed published data and curated database entries.
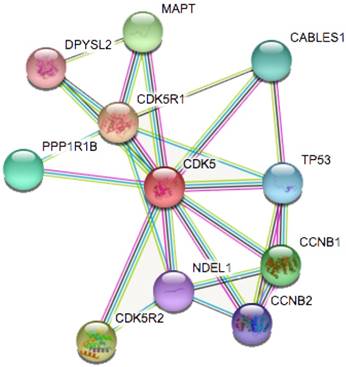
cBioPortal database analysis
The mutations and copy number changes were analyzed by cBioPortal database (http://www.cbioportal.org/) [23]. We selected 13 small cell lung cancer and non-small cell lung cancer datasets. Data Type Priority: Mutation and CAN, Gene Set: CDK5, CDK5R1, CDK5R2, MAPT were analyzed. The results showed in Table 3 and Fig. 5-6.
The alteration frequency of summary for CDK5, CDK5R1, CDK5R2 and MAPT genes in lung cancer subtypes
| Cancer | Data source | N | Frequency (%) | Amplification (%) | Deletion (%) | Mutation (%) | Multiple alteration (%) |
|---|---|---|---|---|---|---|---|
| Lung squ | TCGA, Provisional | 178 | 7.9 | 2.2 | 2.2 | 2.8 | 0.6 |
| Lung squ | TCGE,Nature 2012 | 178 | 6.2 | 0.6 | 1.7 | 3.4 | 0.6 |
| NSCLC | TCGA,Nat Genet 2016 | 1144 | 5.9 | 2.2 | 0.4 | 3.1 | 0.2 |
| Lung adeno | TCGA,Nat Genet 2014 | 230 | 5.7 | 3.9 | 1.3 | 0.4 | |
| Lung adeno | TCGA, Provisional | 230 | 5.7 | 3.9 | 1.3 | 0.4 |
Copy number alteration of CDK5 gene and lung cancer (cBioPortal). (A) The alteration frequency of CDK5, CDK5R1, CDK5R2 and MAPT genes in lung cancer were analyzed by cBioPortal database. (B) The percentage of alteration in CDK5, CDK5R1, CDK5R2 and MAPT in lung squamous cell carcinoma. (C) CDK5 mutations in lung squamous cell carcinoma. Minimal percentage of alteration was set as 1%. The alteration frequency included amplification (red), deep deletions (blue), mutation (green) or multiple alterations (grey).
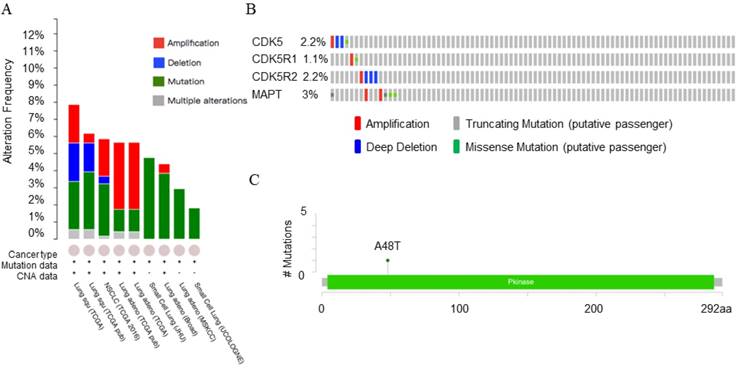
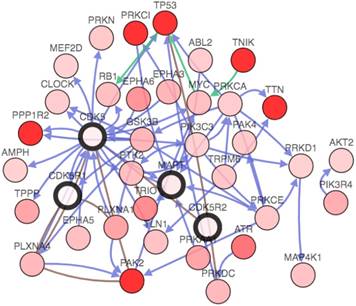
Genetic analysis of CDK5, p35, p39 and MAPT in lung cancer subtypes (cBioPortal). Network view of the CDK5 neighborhood in lung cancer. The network contains 54 nodes, including 4 query genes and the 50 most frequently altered neighbor genes (out of a total of 360). The depth of color represents the degree of alteration.
Cell lines and cell culture reagents
Human lung carcinoma cell lines A549 and PC9 were obtained from FDCC (Shanghai, China) and maintained in RPMI 1640 medium or DMEM (High Glucose) (Thermo, Suzhou, China) supplemented with 10% FBS (Gemini, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA). Human brain cancer cell line U87 were obtained from ATCC (USA) and maintained in DMEM (High Glucose) (Thermo, Suzhou, China) supplemented with 10% FBS (Gemini, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA). Human liver cancer cell line HepG2, breast cancer cell line MCF-7, bladder cancer cell line EJ were obtained from SIBS (Shanghai, China) and maintained in RPMI 1640 or DMEM (High Glucose) (Thermo, Suzhou, China) medium supplemented with 10% FBS (Gemini, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA). Human retinal pigment epithelium (ARPE-19) cells were bought from iCell Bioscience (Shanghai, China) and maintained in DMEM/F12 cell medium (Hyclone, USA) supplemented with 10% FBS (Gemini, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA). MCF-10A were obtained from SIBS (Shanghai, China) and maintained in DMEM (Invitrogen, USA) supplemented with 20% Horse Serum (Invitrogen, USA), 0.02% EGF (Peprotech, USA) , 0.05% Hydrocortisone (Sigma, USA), 0.1%Insulin (Sigma, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA). Roscovitine was from Selleck (Shanghai, China).
Transfection
CDK5 siRNAs were purchased from Sangon Biotech (Shanghai, China). The sequences of the S1, S2 and NC sense strand were 5'- CAGAACCUUCUGAAGUGUAAC-3', 5'- GGGCUGGGAUUCUGUCAUATT-3', and 5'-UUCUCCGAACGUGUCACGUTT-3' respectively. Cells were seeded into six-well plates (Corning, USA) at 1x105/ well and cultured overnight to 40-50% confluence. The cells were transfected with CDK5 siRNA by Lipofectamine 2000 (Invitrogen, USA) according to the manufacturer's instructions. Silencing of CDK5 was examined by Western blots after transfected for 48h.
Antibodies and Western blotting
The A549 cells were homogenized in RIPA lysis buffer containing protease inhibitor mixture (meilunbio, Shanghai, China) on ice for 30 min, then removed from six-well plates (Corning, USA) using a cell scraper and centrifuged for 10 min at 12,000 rpm in a 4℃ Eppendorf microfuge (Germany). Loading buffer was added to the protein lysate and boiled for 10 minutes. Equal amounts of protein were loaded, separated by 10% SDS-PAGE, and transferred to NC membranes (Merck Millipore, Germany). The membranes were incubated first in a 5% powdered milk solution (BBI, Shanghai, China) for 1 hour, then incubated with primary antibodies overnight 4℃, followed by appropriate secondary antibodies. The following primary antibodies were used by CAP1 (1:2000; Santa Cruz, USA), pCAP1 (1:500, donated by Professor Field [24]), CDK5 (1:1000, Cell Signaling Technology, USA), β-Actin (1:1000, Sigma, USA) and GAPDH (1:2000, Bioworld, USA).
MTT cell proliferation assay
Approximately 1000 cells were seeded into each well of a 96-well plate (Corning, USA). At each time point, the cells were incubated with 10μl Dye solution (Solarbio, Beijing, China) for 4 hours at 37 °C followed by cytolysis. The released formazan product was detected with an Epoch Microplate Spectrometer (BioTek, USA) by absorbance reading at 490 nM.
Colony formation assay
For colony formation assay, 1000 transfected cells were seeded into 6-well plates and cultured for 10 days at RPMI 1640 medium or DMEM (High Glucose) (Thermo, Suzhou, China) supplemented with 10% FBS (Gemini, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA). The colonies were imaged and counted after fixed with 75% ethanol and staining with 0.1% crystal violet solution.
Wound healing and transwell assays
Cell migration was detected by wound healing and transwell assays. Cells were seeded into 6-well plates and cultured until the confluence reached 95%. A sterile 10 μl pipette tip was used to generate a scratch through each well. The wound closure was observed after 0h and 24h and photographed under a microscope (Olympus, Tokyo, Japan).
For migration assays, 3x104 transfected cells in serum-free medium were seeded in the upper well of the chambers equipped with 8 μm membranes were used (Corning Incorporated, Corning, NY, USA), and 500 μl of RPMI 1640 medium or DMEM (High Glucose) (Thermo, Suzhou, China) supplemented with 10% FBS (Gemini, USA) and 1% antibiotics (penicillin-streptomycin, Gibco, USA) were added into the lower chambers. After incubated at 37°C for 24 hours, cells in the upper chamber were removed with a cotton tip, the chambers were washed by 1xPBS for 3 times, the cells migrating to the lower surface of the chamber were fixed with 95% ethanol for 10 min, stained with 0.1% crystal violet solution for 10 min, washed for three times, air dried, photographed and counted in five randomly selected fields for each well by a light microscope (Olympus, Tokyo, Japan), cells were counted by ImageJ software (National Institutes of Health, USA).
Statistical analysis
All experiments were performed at least three times. Data are presented as mean ± standard deviation (SD). Statistical analysis was performed with GraphPad Prism (version 6.01, GraphPad Software, USA). Statistical significance was assumed if p≤0.05.
Results
The transcription level of CDK5 in different cancers
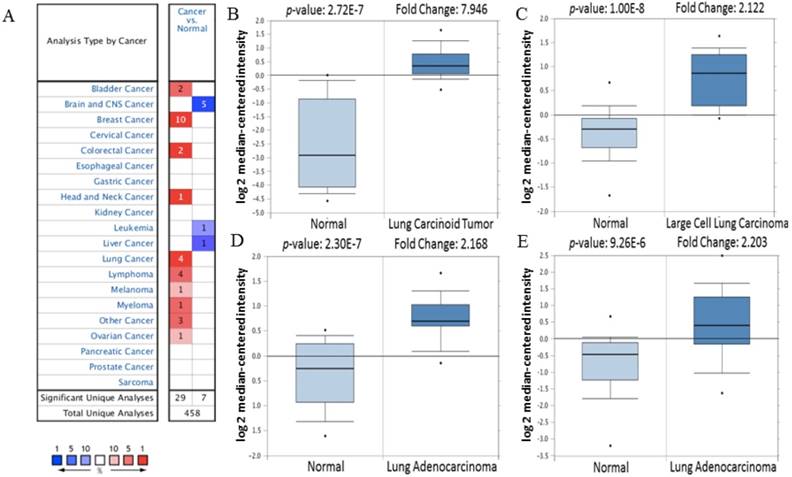
Firstly, the mRNA expression of CDK5 in cancer was analyzed. The main functions of Oncomine database are gene expression differential analysis, gene expression and clinical relevance analysis and multi-gene co-expression analysis. Oncomine database was used to detect the expression of CDK5 mRNA in cancer and normal clinical specimens, and the expression of CDK5 mRNA in lung cancer are shown in Table 1, other cancers results are shown in Table S1. The threshold was set as: p-value<1E-4, fold change>2 and top gene rank 10%. Eventually, 162 datasets, including 18,295 samples, were selected. Compared with normal tissues, the mRNA expression of CDK5 was significantly up-regulated in some cancers, and down-regulated in others (Fig. 1). CDK5 was hyper-expressed in bladder, breast, colorectal, head and neck, lung, lymphoma, melanoma, myeloma, ovarian, uterine corpus leiomyoma, yolk sac tumor and seminoma cancers, and hypo-expressed in brain and central nervous system (CNS), leukemia and liver cancers, suggesting that the transcription level of CDK5 mainly depended on the cancer type.
The transcription level of CDK5 in various cancer types (Oncomine Database). (A) The left column in red showed the number of datasets with CDK5 overexpression and the right column in blue show the under expression datasets number after compared clinical specimen of cancer vs. normal tissue. The box plot comparing specific CDK5 expression in normal (left plot) and cancer tissue (right plot) was derived from Oncomine database. The analysis was shown in (B) lung carcinoid tumor, (C) large cell lung carcinoma, (D, E) lung adenocarcinoma relative to normal lung. The threshold set as follow: p-value: 1E-4, fold change: 2, gene rank: 10%.
The relationship between transcription level of CDK5 and prognosis of cancer
The results of the previous database showed that the expression level of CDK5 mRNA in most cancers including bladder, breast, colorectal, head and neck, lung, lymphoma, melanoma, myeloma, ovarian, uterine corpus leiomyoma, yolk sac tumor and seminoma cancers were higher than that in the normal group and lower in brain and CNS, leukemia and liver cancers. Thus, it is necessary to explore the relationship between the transcription level of CDK5 and prognosis of cancer, and clarify whether CDK5 is a cancer promoter or a tumor suppressor. PrognoScan searches the relation between gene expression and patient prognosis across a large collection of publicly available cancer microarray datasets, provides a platform for evaluating potential tumor markers and therapeutic targets.
The PrognoScan database included 13 types of cancers including bladder, blood, brain, breast, colorectal, esophageal, eye, head and neck, lung, ovarian, prostate, skin and soft tissue cancers. The result of analysis revealed that cancers with statistical significance regarding OS were lung, skin and blood cancers, the p-value was 0.000846, 0.004384, 0.001214 and 0.015104 respectively (Table 2). The high transcription level of CDK5 showed poor prognosis in lung and skin cancer patients, the HR were 8.05, 3.15 and 3.68 respectively (Fig. 2 A-C), whereas good prognosis in blood cancer the HR was 0.38 (Fig. 2D). The Kaplan Meier plotter is capable to assess the effect of 54,675 genes on survival using 10,461 cancer samples, which included data from breast, ovarian, lung and gastric cancers. In order to verify the search results of Prognoscan, we searched the Kaplan-Meier database to predict the relationship between the transcription level of CDK5 and prognosis of cancer. Only in lung cancer, the relationship between CDK5 and prognosis was statistically significant (p=0.0013; HR: 1.23), overexpression of CDK5 related to poor OS (Fig. 3C). There was no statistically significant difference in ovarian, breast and gastric cancer, the p-value was 0.73, 0.43 and 0.32 respectively (Fig. 3A.3B.3D). Both of the two databases indicated that CDK5 was associated with poor prognosis in lung cancer.
CDK5 predicted protein-protein interaction analysis
STRING is a database known to be used to predict protein-protein interactions. The co-expression analysis revealed that CDK5 was co-expressed with cyclin-dependent kinase 5 regulatory subunit 1(CDK5R1/p35), cyclin-dependent kinase 5 regulatory subunit 2(CDK5R2/p39), microtubule-associated protein tau (MAPT), cyclin B1(CCNB1), protein phosphatase 1(PPP1R1B), CDK5 and Abl enzyme substrate 1(CABLES1), tumor protein p53 (TP53), cyclin B2(CCNB2), nudE nuclear distribution E homolog(A. nidulans)-like 1(NDEL1) and dihydropyrimidinase-like 2(DPYSL2), whose correlation score was 0.999, 0.994, 0.991, 0.988, 0.986, 0.986, 0.983, 0.982, 0.978 and 0.976 respectively (Fig. 4). p35 and p39 are widely recognized as activators of CDK5, and the p35/CDK5 complex is essential for the development of the nervous system. MAPT is thought to be associated with the formation and maintenance of the nervous system polarity. We found that the above three proteins (CDK5R1, CDK5R2 and MAPT) had the highest correlation with CDK5, and therefore require more in-depth analysis.
CDK5, p35, p39 and MAPT genomic mutation and CNA in lung cancer subtypes
The cBioPortal for Cancer Genomics provides visualization and analysis of large-scale cancer genomics datasets. Genetic alterations in cancer mainly include mutation, amplification, deletion and multiple alterations. Both PrognoScan and Kaplan-Meier plotter database showed that there was a connection between CDK5 and lung cancer. In order to provide a theoretical basis for further research experiments, CDK5, p35, p39 and MAPT genomic changes in lung cancer subtypes (4104 samples) were studied using 13 datasets of cBioPortal database. Minimal percentage altered samples were set as 1%, and the results are shown in Fig. 5A. Five datasets including lung squamous cell carcinoma (lung squ), non-small cell lung cancer (NSCLC) and lung adenocarcinoma (lung adeno) samples showed gene alteration rates were varied from 5.7% to 7.9%. The sample sizes for these datasets were 178, 178, 1144, 230 and 230 respectively. The detailed information is shown in Table 3.
The OncoPrint database was applied to explore the specific alteration in each gene (Fig. 5A). For example, alteration's percentage in CDK5, CDK5R1, CDK5R2 and MAPT gene among lung squamous cell carcinoma varied from 1.1% to 3.0% in individual genes (CDK5, 2.2%; CDK5R1, 1.1%; CDK5R2, 2.2%; MAPT, 3.0%). The alterations included amplification, deep deletion, truncating mutation and missense mutation (Fig. 5B). Mutations page showed that the 48th amino acid in CDK5 was susceptible to mutations, and the mutation type belonged to missense mutations (Fig. 5C).
Genetic analysis of CDK5, p35, p39 and MAPT in lung cancer subtype
The gene network, which could further analyze genes that interacted with CDK5, p35, p39 and MAPT in lung cancer. The network contains 54 nodes, including 4 query genes and the 50 most frequently altered neighbor genes (out of a total of 360). The depth of color represents the degree of alteration. Darker red nodes including TP53, protein phosphatase 1 regulatory inhibitor subunit 2 (PPP1R2), protein kinase C iota(PRKCI), p21 (RAC1) activated kinase 2(PAK2), titin(TTN) and TRAF2 and NCK interacting kinase(TNIK) gene showed an increased frequency of alteration in lung squamous cell carcinoma (Fig. 6).
Expression of CDK5 in human cancer cell lines
The expression of CDK5 was detected in 8 cell lines, which include 6 human cancer cell lines and 2 normal cell lines (Fig.7A). Analysis showed that expression of CDK5 was up-regulated in most cancer cell lines MCF-7, EJ, A549, PC9, while obviously down-regulated in HepG2 and miner down-regulated in U87 cell line, and nearly no expression of CDK5 was detected in normal cell lines. This was in line with the research outcome of Oncomine database. According to Oncomine CDK5 was hyper-expressed in bladder, breast, colorectal, head and neck, lung, lymphoma, melanoma, myeloma, ovarian, uterine corpus leiomyoma, yolk sac tumor and seminoma cancers, and hypo-expressed in brain and central nervous system (CNS), leukemia and liver cancers. This set of data points to the reliability of database and CDK5 was high expression in human lung cancer cell lines.
Expression of CDK5 in human cancer cell lines (A) and CDK5 promoted the proliferation ability of lung cancer cells in vitro (B-D). (A) The expression of CDK5 was detected in six cancer cell lines and two normal cell lines by western blot. β-actin was used as internal control. (B) Proliferation assay of MTT experiments was performed in A549 cells transfected with NC siRNA or CDK5 siRNA, NC: Normal control. Error bars represent the mean ± s.d. of three independent experiments. (C) The growth inhibitory rate of A549 after treated with roscovitine at the concentration for 10um, 20um, 40um. Error bars represent the mean ± s.d. of three independent experiments. (D) Colony-formation assays was used for detecting the proliferation ability in A549 cell line after treated with roscovitine. (# p<0.05; * p<0.01; ** p<0.001).
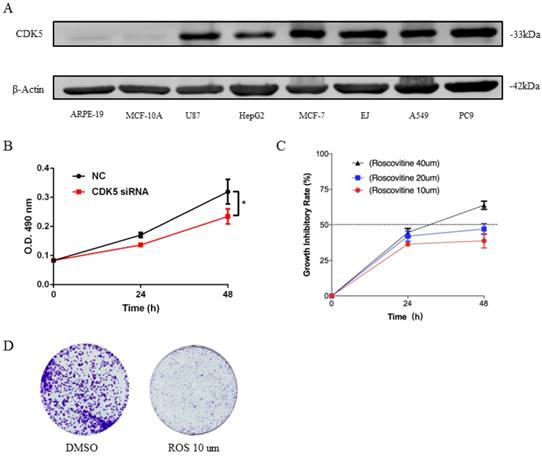
CDK5 promoted the proliferation ability of lung cancer cells in vitro
To investigate the biological function of CDK5 in lung cancer cells, we transfected A549 and PC9 cells with CDK5 siRNA and pharmacological inhibited CDK5 by small molecule roscovitine. MTT and colony formation assays were performed to validate the impact of CDK5 in proliferation. Experiments showed that S2 interference effect was better than S1, without specific notification CDK5 siRNA means S2.MTT assay showed that after down-regulated the expression of CDK5 by CDK5 siRNA significantly inhibited the proliferation (Fig.7B, Fig.S1A). Roscovitine inhibited the proliferation of A549 and PC9, and showed the IC50 was between 20um and 40um (Fig.7C, Fig.S1B). Colony formation assays showed the similar results, roscovitine significantly inhibited the colony formation. (Fig.7D, Fig.S1C)
CDK5 promoted lung cancer cell migration in vitro
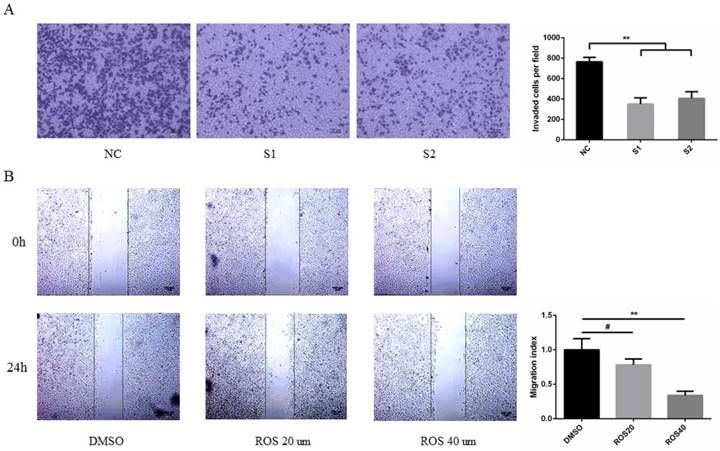
To determine the effect of CDK5 on metastasis of lung cancer cells. A549 and PC9 cells were transfected with CDK5 siRNA or treated with CDK5 specific inhibitor roscovitine then performed transwell invasion and wound healing assay respectively in indicated time. The results showed that transfected lung cancer cells by CDK5 siRNA will inhibit cells migration (Fig.8A, Fig.S2A). A concentration-dependent decrease of migration by Cdk5 inhibitor roscovitine was observed (Fig.8B, Fig.S2B).
CDK5 promoted lung cancer cell migration in vitro. (A) Transwell assay was performed in A549 cells transfected with NC siRNA and CDK5 siRNA. NC: Normal control, S1: si-RNA1, S2: si-RNA2. (B) Wound healing assay was performed in A549 cells after treated with roscovitine at the concentration of 20um or 40um. Error bars represent the mean ± s.d. of three independent experiments. (# p<0.05; * p<0.01; ** p<0.001).
Inhibition of CDK5 decreases CAP1 phosphorylation
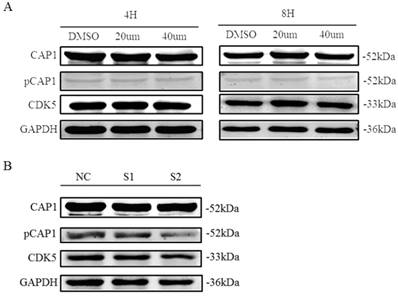
Recent findings suggested CDK5 as a kinase that phosphorylates the S307/S309 regulatory site on cyclase-associated protein 1 (CAP1; H. Zhang & G.L. Zhou, unpublished results). We tested the impact of CDK5 inhibition on the phosphorylation of CAP1 in lung cancer cell line A549 by performing different assays using the CDK5 inhibitor roscovitine and CDK5 siRNA. We found that phosphorylated CAP1 (pCAP1) was decreased after treatment with CDK5 inhibitor roscovitine and knockdown CDK5 by siRNA in lung cancer cell (Fig. 9). These results support that CDK5 regulates CAP1 phosphorylation in lung cancer cells as well.
CDK5 regulates CAP1 phosphorylation. Western blots analysis showed the protein of pCAP1 was decreased after treatment with CDK5 inhibitor roscovitine and knockdown CDK5 by siRNA in lung cancer cell A549. NC: Normal control, S1: si-RNA1, S2: si-RNA2.
Discussion
The present study aimed at exploring the role of CDK5 in cancers by using an extensive bioinformatics data mining process to find out the expression of CDK5 mRNA in different cancers, and then predict the connection between the expression level and OS. The protein network of predicted associations for CDK5 and alterations in cancer genomics were analyzed. Western blots, cell proliferation and migration experiments were performed to validate the connection between CDK5 and lung cancer.
CDKs, a group of proline-directed serine/ threonine protein kinases, are conserved throughout evolution and can be found in species from saccharomyces cerevisia to humans. In humans, there are 13 different CDKs (CDK1-CDK13) highly expressed in mitotic cells [25]. But CDK5 is unusual because it is not typically activated upon binding with a cyclin and does not require T-loop phosphorylation for activation; instead, it is activated through specific binding with p35 or p39 protein, or their respective cleaved counterparts p25 and p29 [26]. CDK5 is well characterized for its role in the CNS. While increased levels of CDK5 target proteins are being considered as possible biomarkers of specific cancers. Zhou et al. found that CDK5 activated FAK/AKT signaling pathway to generated vasculogenic mimicry in NSCLC, and identification of CDK5 as a key factor regulating migration and metastasis by regulated actin [27]. Zhuang et al. revealed that CDK5 promotes human colorectal cancer via ERK5-AP-1 axis [28]. Several researchers established that CDK5 enhanced cell-cycle progression and facilitated medullary thyroid carcinoma (MTC) proliferation by phosphorylated retinoblastoma protein (Rb) at Ser807/811 [29]. R. Dixon Dorand found that after injecting tumor cells that block the activity of CDK5 into mice, more than half of the mice survived, while almost all the mice in the control group died, which verified the CDK5 associated with tumor development [30]. The regulation of CDK5 activity is now emerging as a candidate therapeutic target. The inhibition or knockdown of CDK5 has been proven to play an anti-cancer role through various mechanisms, and can synergize the killing effect of chemotherapeutics. The results are similar to our study. Our analysis predicted CDK5 were higher expressed in most cancer patients compared to normal by Oncomine database. In addition, to further explore the relationship between CDK5 and OS in various cancer types by PrognoScan and Kaplan-Meier Plotter databases. However, the results between OS and other cancers were not statistically significant except lung cancer. Those results suggest that a high transcription level of CDK5 was correlated with poor prognosis in lung cancer patients.
The cause of human cancer is mainly due to irreparable structural mutations in cells. These mutations at very specific genomic locations can alter the function of the gene and DNA copy number [31, 32]. Pollack et al. found that copy number alterations (CNAs) uncover all gene expression, which could be a critical element in the tumor development [33]. Here in identifying CNAs is a new method for linking CNA with the disease phenotype. The present study aimed to determine if the CNAs of the CDK5 correlate with aggressive cancer sub-types, based on the cBioPortal [34, 35]. The mutations alteration (2.8-3.4%) was the most common gene alteration in lung squamous cell carcinoma, while in lung adenocarcinoma patients amplifications (3.9%) were the most common gene alteration. Meanwhile, the 48th amino acid in CDK5 was susceptible to missense mutations (Fig. 5C).
Furthermore, the cBioPortal can be used for interactive analysis and visualization of altered networks. According to STRING analysis, CDK5 is mainly activated by p35/p25, p39/p29 (Fig. 4). In addition, CDK5 is also co-expressed with MAPT, CCNB1, PPP1R1B, CABLES1, TP53, CCNB2, NDEL1 and DPYSL2 in lung cancer (Fig. 4). MAPT and NDEL1 related to microtubule, DPYSL2 related to metastasis which subsequent remodeling of the cytoskeleton [36-38]. Fig. 6 showed that the network view of the CDK5 in lung squamous cell carcinoma, those results will help researchers to better understand the molecular mechanisms of CDK5 in lung cancer. Serine/threonine protein kinases encoded by PRKCI gene is a member of the protein kinase C family. It plays a role in microtubule dynamics in the early secretory pathway, which promotes metastasis of esophageal cancer via PKCι-SKP2-AKT pathway [39]. The p21 activated kinases (PAK) are critical effectors that link Rho GTPases to cytoskeleton reorganization and nuclear signaling [40]. TTN gene encodes a large number of muscle related proteins [41]. Protein encoded by TNIK gene is a serine/threonine kinase that functions as an activator of the Wnt signaling pathway which plays important roles in carcinogenesis [42]. These data analysis results show that CDK5 is related to cell migration and skeleton, providing a direction for further research.
In order to verify the reliance of databases we performed some experiments. Western blots showed that the expression level of CDK5 were up-regulated in most cancer cell lines MCF-7, EJ, A549, PC9, while obviously down-regulated in HepG2 and miner down-regulated in U87 cell line, and nearly no expression of CDK5 was detected in normal cell lines. This was in line with the research outcome of Oncomine database and some previous reports [43-45]. In addition, we also found that knockdown of CDK5 inhibited lung cancer cells proliferation and migration (Fig. 7.8). These results indicated that CDK5 may function as a tumor activator in lung cancer. CDK5 has been reported up-regulated and to act as a tumor activator in many human cancers, including colorectal cancer [43], breast cancer [44], pancreatic cancer[45]. These results showed that CDK5 can be a promising anticancer target.
Caldesmon phosphorylation and actin polymerization can be observed in CDK5 activity [46]. CAP1, a protein encoded by the CAP gene, plays an important role in cell movement and morphological changes by acting synergistically with cofilin to regulate cytoskeleton movement [47]. As is known to all tumor cell growth, differentiation, apoptosis and motility are accompanied by abnormal phosphorylation of the protein in cell signal transduction [48, 49]. Our group has done a long study of CAP1 and found that CAP1 is abnormal in lung cancer cells and lung cancer tissues, and may be related to poor prognosis and the signal transduction of cytoskeleton and movement [50]. Compare with normal tissue, CAP1 is overexpressed in lung cancer tissue, particularly in the metastasis state. Clinical and experimental results are consistent with the results of the database analysis [51, 52].
Zhou et al. found that there were 9 CAP1 phosphorylation sites; one of the phosphorylation sites S309 of CAP1 can recognize the sequence of CDKs (S/T-P-X-K/R/H) [53]. CDKs are a large family of serine/threonine kinases. Research showed that this sequence is a specific sequence of phosphorylated substrate of CDK5 [54]. Therefore CDK5 may be potential kinase of CAP1. After treated with the inhibitor of CDK5, the level of phosphorylated CAP1 was decreased (Fig. 9). Therefore, the phosphorylation of CAP1 is regulated by CDK5.
Database is a powerful tool can help us to screen some target molecules that are worth studying and provide more basis for research, which can predict the direction of the experiment and save research funding. Our study aimed to better understand the molecular mechanisms by extensive oncogenic databases. This study provides a theoretical basis for clinicians and researchers, and finds out a new signaling pathway or biomarker in cancer, which may help develop novel therapeutic approaches for early intervention in cancer prevention. However, this study still has some limitations, the role of CDK5 needed to be further clarified. We intend to further verify the role and function of CDK5 in lung cancer by regulating the phosphorylation site S309 of CAP1.
Conclusion
In summary, this study explored the role of CDK5 in cancers by using an extensive bioinformatics data mining process and seeks new strategies for targeting CDK5 and its downstream mechanisms. CDK5 may prove to be a valid target for anticancer therapies.
Abbreviations
CDK5: Cyclin-Dependent Kinase 5; OS: overall survival; Lung Squ: lung squamous cell carcinoma; NSCLC: non-small cell lung cancer; Lung Adeno: lung adenocarcinoma; CNS: central nervous system; CAP1: cyclase-associated protein 1; pCAP1: phosphorylated CAP1; MTC: medullary thyroid carcinoma; Rb: retinoblastoma protein; CAN: copy number alteration; CDK5R1(p35): Cyclin-dependent kinase 5 regulatory subunit 1; CDK5R2(p39): Cyclin-dependent kinase 5 regulatory subunit 2; MAPT: Microtubule-associated protein tau; CCNB1: Cyclin B1; PPP1R1B: Protein phosphatase 1; CABLES1: Cdk5 and Abl enzyme substrate 1; TP53: Tumor protein p53; CCNB2: Cyclin B2; NDEL1: nudE nuclear distribution E homolog (A. nidulans)-like 1; DPYSL2: Dihydropyrimidinase-like 2; PPP1R2: Protein phosphatase 1 regulatory inhibitor subunit 2; PRKCI: protein kinase C iota; PAK2: p21(RAC1) activated kinase 2; TTN: titin, TNIK, TRAF2 and NCK interacting kinase.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81472180, 81802262) and Shanghai Tenth Hospital's improvement plan for NSFC (04.03.17.032). GLZ is supported by an Institutional Development Award (IDeA) from the NIGMS of the National Institutes of Health [grant no. P20GM103429].The funder has no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author Contributions
Conception and design: Shuangshuan Xie, Changhui Wang and Guo-Lei Zhou; Administrative support: Xiaolian Song, Changxing Shen; Provision of study materials: Jie Zeng, Yang Liu; Collection and assembly of data: Jie Zeng, Shuanshuan Xie; Data analysis and interpretation: All authors; Manuscript writing: All authors; Final approval of manuscript: All authors.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67:7-30
2. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. CELL. 2011;144:646-674
3. Asghar U, Witkiewicz AK, Turner NC. et al. The history and future of targeting cyclin-dependent kinases in cancer therapy. NAT REV DRUG DISCOV. 2015;14:130-146
4. Dhavan R, Tsai LH. A decade of CDK5. Nat Rev Mol Cell Biol. 2001;2:749-759
5. Lopes JP, Agostinho P. Cdk5: multitasking between physiological and pathological conditions. PROG NEUROBIOL. 2011;94:49-63
6. Watanabe G, Pena P, Shambaugh GR. et al. Regulation of cyclin dependent kinase inhibitor proteins during neonatal cerebella development. Brain Res Dev Brain Res. 1998;108:77-87
7. Modi PK, Komaravelli N, Singh N. et al. Interplay between MEK-ERK signaling, cyclin D1, and cyclin-dependent kinase 5 regulates cell cycle reentry and apoptosis of neurons. MOL BIOL CELL. 2012;23:3722-3730
8. Odajima J, Wills ZP, Ndassa YM. et al. Cyclin E constrains Cdk5 activity to regulate synaptic plasticity and memory formation. DEV CELL. 2011;21:655-668
9. Patrick GN, Zukerberg L, Nikolic M. et al. Conversion of p35 to p25 deregulates Cdk5 activity and promotes neurodegeneration. NATURE. 1999;402:615-622
10. Pozo K, Castro-Rivera E, Tan C. et al. The role of Cdk5 in neuroendocrine thyroid cancer. CANCER CELL. 2013;24:499-511
11. Bhandari D, Lopez-Sanchez I, To A. et al. Cyclin-dependent kinase 5 activates guanine nucleotide exchange factor GIV/Girdin to orchestrate migration-proliferation dichotomy. Proc Natl Acad Sci U S A. 2015;112:E4874-E4883
12. Grant NJ, Coates PJ, Woods YL. et al. Phosphorylation of a splice variant of collapsin response mediator protein 2 in the nucleus of tumour cells links cyclin dependent kinase-5 to oncogenesis. BMC CANCER. 2015;15:885
13. Fang WQ, Ip JP, Li R. et al. Cdk5-mediated phosphorylation of Axin directs axon formation during cerebral cortex development. J NEUROSCI. 2011;31:13613-13624
14. Liebl J, Furst R, Vollmar AM. et al. Twice switched at birth: cell cycle-independent roles of the "neuron-specific" cyclin-dependent kinase 5 (Cdk5) in non-neuronal cells. CELL SIGNAL. 2011;23:1698-1707
15. Xie ZC, Dang YW, Wei DM. et al. Clinical significance and prospective molecular mechanism of MALAT1 in pancreatic cancer exploration: a comprehensive study based on the GeneChip, GEO, Oncomine, and TCGA databases. Onco Targets Ther. 2017;10:3991-4005
16. Bhattacharjee A, Richards WG, Staunton J. et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci U S A. 2001;98:13790-13795
17. Hou J, Aerts J, den Hamer B. et al. Gene expression-based classification of non-small cell lung carcinomas and survival prediction. PLOS ONE. 2010;5:e10312
18. Stearman RS, Dwyer-Nield L, Zerbe L. et al. Analysis of orthologous gene expression between human pulmonary adenocarcinoma and a carcinogen-induced murine model. AM J PATHOL. 2005;167:1763-1775
19. Su LJ, Chang CW, Wu YC. et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC GENOMICS. 2007;8:140
20. Mizuno H, Kitada K, Nakai K. et al. PrognoScan: a new database for meta-analysis of the prognostic value of genes. BMC MED GENOMICS. 2009;2:18
21. Hou GX, Liu P, Yang J. et al. Mining expression and prognosis of topoisomerase isoforms in non-small-cell lung cancer by using Oncomine and Kaplan-Meier plotter. PLOS ONE. 2017;12:e174515
22. Crosara K, Moffa EB, Xiao Y. et al. Merging in-silico and in vitro salivary protein complex partners using the STRING database: A tutorial. J PROTEOMICS. 2018;171:87-94
23. Gao J, Aksoy BA, Dogrusoz U. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. SCI SIGNAL. 2013;6:l1
24. Freeman NL, Field J. Mammalian homolog of the yeast cyclase associated protein, CAP/Srv2p, regulates actin filament assembly. Cell Motil Cytoskeleton. 2000;45:106-120
25. Shah K, Lahiri DK. Cdk5 activity in the brain - multiple paths of regulation. J CELL SCI. 2014;127:2391-2400
26. Malumbres M. Cyclin-dependent kinases. GENOME BIOL. 2014;15:122
27. Zhou X, Gu R, Han X. et al. Cyclin-dependent kinase 5 controls vasculogenic mimicry formation in non-small cell lung cancer via the FAK-AKT signaling pathway. Biochem Biophys Res Commun. 2017;492:447-452
28. Zhuang K, Zhang J, Xiong M. et al. CDK5 functions as a tumor promoter in human colorectal cancer via modulating the ERK5-AP-1 axis. CELL DEATH DIS. 2016;7:e2415
29. Pozo K, Hillmann A, Augustyn A. et al. Differential expression of cell cycle regulators in CDK5-dependent medullary thyroid carcinoma tumorigenesis. ONCOTARGET. 2015;6:12080-12093
30. Dorand RD, Nthale J, Myers JT. et al. Cdk5 disruption attenuates tumor PD-L1 expression and promotes antitumor immunity. SCIENCE. 2016;353:399-403
31. Fridlyand J, Snijders AM, Ylstra B. et al. Breast tumor copy number aberration phenotypes and genomic instability. BMC CANCER. 2006;6:96
32. Pinkel D, Segraves R, Sudar D. et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. NAT GENET. 1998;20:207-211
33. Pollack JR, Sorlie T, Perou CM. et al. Microarray analysis reveals a major direct role of DNA copy number alteration in the transcriptional program of human breast tumors. Proc Natl Acad Sci U S A. 2002;99:12963-12968
34. Cerami E, Gao J, Dogrusoz U. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. CANCER DISCOV. 2012;2:401-404
35. Beroukhim R, Mermel CH, Porter D. et al. The landscape of somatic copy-number alteration across human cancers. NATURE. 2010;463:899-905
36. Sampedro F, Marin-Lahoz J, Martinez-Horta S. et al. Early Gray Matter Volume Loss in MAPT H1H1 de Novo PD Patients: A Possible Association With Cognitive Decline. FRONT NEUROL. 2018;9:394
37. Chansard M, Hong JH, Park YU. et al. Ndel1, Nudel (Noodle): flexible in the cell? Cytoskeleton (Hoboken). 2011;68:540-554
38. Pham X, Song G, Lao S. et al. The DPYSL2 gene connects mTOR and schizophrenia. Transl Psychiatry. 2016;6:e933
39. Liu SG, Wang BS, Jiang YY. et al. Atypical protein kinase Ciota (PKCiota) promotes metastasis of esophageal squamous cell carcinoma by enhancing resistance to Anoikis via PKCiota-SKP2-AKT pathway. MOL CANCER RES. 2011;9:390-402
40. Rane CK, Minden A. P21 activated kinase signaling in cancer. SEMIN CANCER BIOL. 2018
41. Bobylev AG, Galzitskaya OV, Fadeev RS. et al. Smooth muscle titin forms in vitro amyloid aggregates. Biosci Rep. 2016:36
42. Masuda M, Sawa M, Yamada T. Therapeutic targets in the Wnt signaling pathway: Feasibility of targeting TNIK in colorectal cancer. Pharmacol Ther. 2015;156:1-9
43. Zhuang K, Zhang J, Xiong M. et al. CDK5 functions as a tumor promoter in human colorectal cancer via modulating the ERK5-AP-1 axis. CELL DEATH DIS. 2016;7:e2415
44. Mandl MM, Zhang S, Ulrich M. et al. Inhibition of Cdk5 induces cell death of tumor-initiating cells. Br J Cancer. 2017;116:912-922
45. Jin X, Yang C, Fan P. et al. CDK5/FBW7-dependent ubiquitination and degradation of EZH2 inhibits pancreatic cancer cell migration and invasion. J BIOL CHEM. 2017;292:6269-6280
46. Bisht S, Nolting J, Schutte U. et al. Cyclin-Dependent Kinase 5 (CDK5) Controls Melanoma Cell Motility, Invasiveness, and Metastatic Spread-Identification of a Promising Novel therapeutic target. TRANSL ONCOL. 2015;8:295-307
47. Wang C, Zhou GL, Vedantam S. et al. Mitochondrial shuttling of CAP1 promotes actin- and cofilin-dependent apoptosis. J CELL SCI. 2008;121:2913-2920
48. Ntantie E, Fletcher J, Amissah F. et al. Polyisoprenylated cysteinyl amide inhibitors disrupt actin cytoskeleton organization, induce cell rounding and block migration of non-small cell lung cancer. ONCOTARGET. 2017;8:31726-31744
49. Jacquemet G, Hamidi H, Ivaska J. Filopodia in cell adhesion, 3D migration and cancer cell invasion. CURR OPIN CELL BIOL. 2015;36:23-31
50. Tan M, Song X, Zhang G. et al. Overexpression of adenylate cyclase-associated protein 1 is associated with metastasis of lung cancer. ONCOL REP. 2013;30:1639-1644
51. Zhang H, Ghai P, Wu H. et al. Mammalian adenylyl cyclase-associated protein 1 (CAP1) regulates cofilin function, the actin cytoskeleton, and cell adhesion. J BIOL CHEM. 2013;288:20966-20977
52. Xie S, Shen C, Tan M. et al. Systematic analysis of gene expression alterations and clinical outcomes of adenylate cyclase-associated protein in cancer. ONCOTARGET. 2017;8:27216-27239
53. Zhou GL, Zhang H, Wu H. et al. Phosphorylation of the cytoskeletal protein CAP1 controls its association with cofilin and actin. J CELL SCI. 2014;127:5052-5065
54. Cheung ZH, Ip NY. Cdk5: mediator of neuronal death and survival. NEUROSCI LETT. 2004;361:47-51
55. Sanchez-Carbayo M, Socci ND, Lozano J. et al. Defining molecular profiles of poor outcome in patients with invasive bladder cancer using oligonucleotide microarrays. J CLIN ONCOL. 2006;24:778-789
56. Bredel M, Bredel C, Juric D. et al. Functional network analysis reveals extended gliomagenesis pathway maps and three novel MYC-interacting genes in human gliomas. CANCER RES. 2005;65:8679-8689
57. Rickman DS, Bobek MP, Misek DE. et al. Distinctive molecular profiles of high-grade and low-grade gliomas based on oligonucleotide microarray analysis. CANCER RES. 2001;61:6885-6891
58. Sun L, Hui AM, Su Q. et al. Neuronal and glioma-derived stem cell factor induces angiogenesis within the brain. CANCER CELL. 2006;9:287-300
59. Curtis C, Shah SP, Chin SF. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. NATURE. 2012;486:346-352
60. Zhao H, Langerod A, Ji Y. et al. Different gene expression patterns in invasive lobular and ductal carcinomas of the breast. MOL BIOL CELL. 2004;15:2523-2536
61. Gluck S, Ross JS, Royce M. et al. TP53 genomics predict higher clinical and pathologic tumor response in operable early-stage breast cancer treated with docetaxel-capecitabine +/- trastuzumab. Breast Cancer Res Treat. 2012;132:781-791
62. TCGA. http://tcga-data.nci.nih.gov/tcga/
63. Gaedcke J, Grade M, Jung K. et al. Mutated KRAS results in overexpression of DUSP4, a MAP-kinase phosphatase, and SMYD3, a histone methyltransferase, in rectal carcinomas. Genes Chromosomes Cancer. 2010;49:1024-1034
64. Sabates-Bellver J, Van der Flier LG, de Palo M. et al. Transcriptome profile of human colorectal adenomas. MOL CANCER RES. 2007;5:1263-1275
65. Estilo CL, O-charoenrat P, Talbot S. et al. Oral tongue cancer gene expression profiling: Identification of novel potential prognosticators by oligonucleotide microarray analysis. BMC CANCER. 2009;9:11
66. Basso K, Margolin AA, Stolovitzky G. et al. Reverse engineering of regulatory networks in human B cells. NAT GENET. 2005;37:382-390
67. Chen X, Cheung ST, So S. et al. Gene expression patterns in human liver cancers. MOL BIOL CELL. 2002;13:1929-1939
68. Piccaluga PP, Agostinelli C, Califano A. et al. Gene expression analysis of peripheral T cell lymphoma, unspecified, reveals distinct profiles and new potential therapeutic targets. J CLIN INVEST. 2007;117:823-834
69. Talantov D, Mazumder A, Yu JX. et al. Novel genes associated with malignant melanoma but not benign melanocytic lesions. CLIN CANCER RES. 2005;11:7234-7242
70. Yoshihara K, Tajima A, Komata D. et al. Gene expression profiling of advanced-stage serous ovarian cancers distinguishes novel subclasses and implicates ZEB2 in tumor progression and prognosis. CANCER SCI. 2009;100:1421-1428
71. Korkola JE, Houldsworth J, Chadalavada RS. et al. Down-regulation of stem cell genes, including those in a 200-kb gene cluster at 12p13.31, is associated with in vivo differentiation of human male germ cell tumors. CANCER RES. 2006;66:820-827
72. Crabtree JS, Jelinsky SA, Harris HA. et al. Comparison of human and rat uterine leiomyomata: identification of a dysregulated mammalian target of rapamycin pathway. CANCER RES. 2009;69:6171-6178
Author contact
Corresponding author: Changhui Wang, No.301, Mid Yanchang Rd, Department of Respiratory Medicine, Shanghai Tenth People's Hospital, Tongji University, Shanghai, China, 200072. Email: wang-chang-huicom, Fax number: 86-021-66301685, Telephone: 86-021-66301685

 Global reach, higher impact
Global reach, higher impact