Impact Factor ISSN: 1837-9664
J Cancer 2017; 8(13):2487-2491. doi:10.7150/jca.18824 This issue Cite
Research Paper
Upregulation of Serum miR-10b Is Associated with Poor Prognosis in Patients with Melanoma
Department of Plastic and Reconstructive Surgery, Peking Union Medical College Hospital, Chinese Academy of Medical Science and Peking Union Medical College, Beijing, China
Received 2016-12-19; Accepted 2017-2-19; Published 2017-8-2
Abstract
Aberrant expression of microRNAs (miRNAs) are believed to play a central role in the initiation and development of cancer. The aim of our study was to determine the clinical significance of serum miR-10b in melanoma. A total of 85 and 30 serum samples were obtained from patients with melanoma and healthy volunteers respectively. qRT-PCR was performed to evaluate the expression level of miR-10b in the melanoma cell lines and the serum samples from the participants. Then the clinical significance of serum miR-10b was further determined. Our results showed that the expression level of miR-10b was significantly increased in metastasis melanoma cells and melanoma patients compared to their respective controls. In addition, serum miR-10b expression level was able to discriminate melanoma patients from healthy volunteers as well differentiate melanoma patients at different clinical stage with high accuracy. Moreover, upregulation of serum miR-10b was positively associated with enhanced lymph node metastasis, advanced clinical stage and a shortened survival rate. Finally serum miR-10b was an independent prognostic factor for melanoma. Collectively, our study suggests that serum miR-10b level is upregulated in melanoma and associated with poor prognosis. It may be used as a potential prognostic biomarker for melanoma.
Keywords: Serum, miR-10b, melanoma, Prognosis.
Introduction
Melanoma, derived from the pigment-producing melanocytes, is one of the most invasive tumors (1). It causes a large majority of skin cancer related deaths but accounts for only 1% of all skin cancers. Unfortunately, despite the great advance in the treatment of melanoma, the five year overall survival of melanoma patients in advanced stage remains stagnant (2). In addition, the incidence of melanoma has been rising in the past few decades (3). Therefore, it is important to uncover the molecular mechanisms that regulate the initiation and development of melanoma.
MicroRNAs (miRNAs) are small non-coding RNAs of 20-22 nucleotides which regulate gene expression at the post-transcriptional level (4). These molecules play a central role in regulating virtually all the biological processes. Aberrant expression of miRNAs has been reported in a number of human diseases including cancer (5). miRNAs might behave as oncogenes and tumor suppressors in the carcinogenesis process (6). RAB22A acted as an oncogene in melanoma and its activity was regulated by miR-203. In addition, the expression level of miR-203 was significantly reduced in melanoma tissues and cell lines compared to the controls, indicating miR-203 was a tumor suppressor in melanoma (7). The expression level of miR-769 was remarkably increased in melanoma samples. Ectopic expression of miR-769 enhanced the proliferative capacity of melanoma cells, and vice versa. In addition, glycogen synthase kinase-3 beta was demonstrated to be a direct target of miR-769, suggesting miR-769 was a tumor promoter in the progression of melanoma (8).
The mammalian miR-10 family, including miR-10a and miR-10b, is highly conserved. Although miR-10a and miR-10b have identical seed sequences, they regulate various different pathways (9). Deregulation of miR-10b has been found in many types of cancer such as breast cancer, colerectal cancer, hepatocellular carcinoma and melanoma (10-13). In addition, upregulated tissue miR-10b levels were associated with poor prognosis of melanoma. However, the clinical significance of serum miR-10b in patients with melanoma remains unknown. Thus the aim of current study was to investigate the potential prognostic value of serum miR-10b in melanoma.
Materials and Methods
Cell culture
All five melanoma cell lines (A375M, K1735, BLM, WM115 and WM793) were cultured in Dulbecco's Modified Eagle's Medium containing 10% fetal bovine serum and streptomycin/penicillin at 37 °C in a humidified atmosphere containing 5% CO2.
Patients and sample collection
A total of 115 participants were included in this study. The investigation was approved by the Research Ethics Committee of Peking Union Medical College Hospital and written consent was obtained from all subjects and their relatives. The peripheral blood samples were obtained from 85 patients who were diagnosed with melanoma before any treatment (surgery, chemo- or radiotherapeutic treatment etc.) and from 30 healthy volunteers at Department of Plastic and Reconstructive Surgery, Peking Union Medical College Hospital. The melanoma patients were staged according to the American Joint Committee on Cancer (AJCC) system. Briefly, the samples were centrifuged at 1200 g for 10 min at 4 °C within 30 min after collection. Then the supernatant was aliquoted to a fresh tube and stored at -80 °C until used.
Realtime PCR
Total RNA was isolated from cells and the serum samples using Trizol and miRNeasy Serum/Plasma kit (Qiagen, Hilden, Germany) according to manufacturer's instructions respectively. Reverse transcription was performed using the TaqMan MicroRNA Reverse Transcription kit (Applied Biosystems, Foster City, CA, USA). cDNA was amplified with SYBR Green PCR Master Mix (TOYOBO, Osaka, Japan) by Applied Biosystems 7500 real-time PCR System (Applied Biosystems). The relative expression of serum miR-10b was calculated using the 2-ΔΔCt method and U6 snRNA was used as the internal control. Each sample was analyzed in triplicate.
Statistical analysis
All the data was expressed as mean ± SEM. One-way ANOVA was used to compare the mean differences of serum miR-10b among different groups.
The receiver operator characteristic curve (ROC) was performed, and the area under the ROC curve (AUC) was calculated to evaluate the sensitivity and specificity of the serum miR-10b in the diagnosis of melanoma. Correlations between serum miR-10b and clinicopathological parameters were analyzed using the Chi-square test. Survival analysis was performed following the Kaplan-Meier method, and comparisons were made using the log rank test. Multivariable Cox proportional hazards model was used to identify the independent prognostic factor for melanoma. P<0.05 was considered to indicate a statistically significant difference.
Results
miR-10b was upregulated in invasive melanoma cell lines
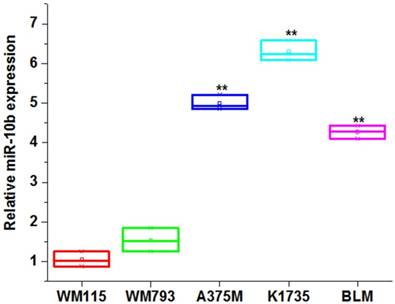
We first compared the expression level of miR-10b among the melanoma cell lines. Our results showed the miR-10b was significantly increased in high metastasis melanoma cell lines (A375M, K1735 and BLM) compared to non-metastasis melanoma cell lines (WM115 and WM793) (**P<0.01) (Figure 1), indicating miR-10b expression level was associated with the invasive capability of melanoma cells.
The expression level of miR-10b in melanoma cell lines
Serum miR-10b level was upregulated in patients with melanoma and had potential diagnostic value
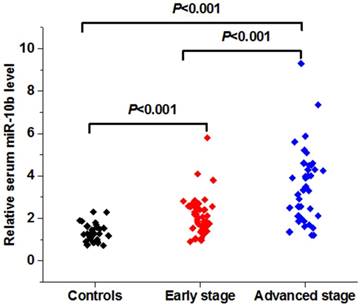
The expression level of serum miR-10b was remarkably increased in patients with melanoma when compared to the healthy volunteers. In addition, serum miR-10b level was overexpressed in melanoma patients in the advanced stage (Ⅲ-Ⅳ) than those in the early stage (Ⅰ-Ⅱ) (***P<0.001) (Figure 2A).
The expression level and diagnostic value of serum miR-10b in melanoma
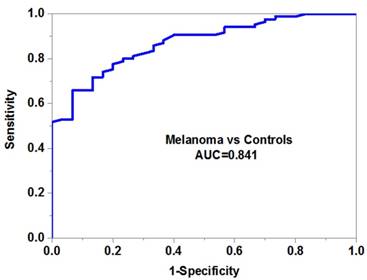
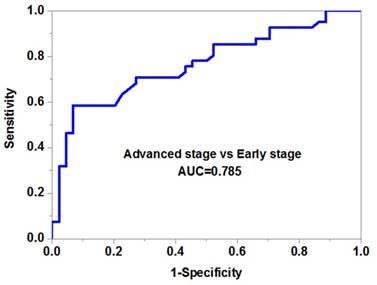
We then evaluated the potential diagnostic value of serum miR-10b in melanoma. Our data demonstrated that serum miR-10b was able to discriminate melanoma patients from healthy volunteers with high accuracy (AUC=0.841, specificity=0.88 and sensitivity=0.76) (Figure 2B). More importantly, serum miR-10b level could also differentiate the patients in the advanced stage from those in the early stage (AUC=0.785, specificity=0.82 and sensitivity=0.74), indicating serum miR-10b could be a useful biomarker for melanoma staging (Figure 2C).
The association between serum miR-10b and clinicopathological parameters of melanoma
Our Chi-square analysis showed that high serum miR-10b was positively associated with advanced TNM stage (P=0.0008) and positive lymph node metastasis (P=0.0013). However, it was not correlated with other clinical features including age, gender, site, tumor thickness, ulceration and differentiation (Table 1).
The correlation between serum miR-10b and the clinical features of melanoma
| Variables | Cases | Serum miR-10b | P |
|---|---|---|---|
| Low High | |||
| Gender | 0.5913 | ||
| Male | 45 | 21 24 | |
| Female | 40 | 21 19 | |
| Age | 0.0630 | ||
| <60 | 46 | 27 19 | |
| ≥60 | 39 | 15 24 | |
| Site | 0.5884 | ||
| Sun exposed | 42 | 22 20 | |
| Sun protected | 43 | 20 23 | |
| Ulceration | 0.7231 | ||
| No | 51 | 26 25 | |
| Yes | 34 | 16 18 | |
| Tumor thickness(mm) | 0.4483 | ||
| <1 | 31 | 17 14 | |
| ≥1 | 54 | 25 29 | |
| Lymph node metastasis | 0.0013 | ||
| No | 50 | 32 18 | |
| Yes | 35 | 10 25 | |
| TNM stage | 0.0008 | ||
| Ι-Ⅱ | 41 | 28 13 | |
| Ⅲ-Ⅳ | 44 | 14 30 | |
| Differentiation | 0.1509 | ||
| Well/moderate | 48 | 27 21 | |
| Poor | 37 | 15 22 |
Correlation between serum miR-10b and survival of melanoma
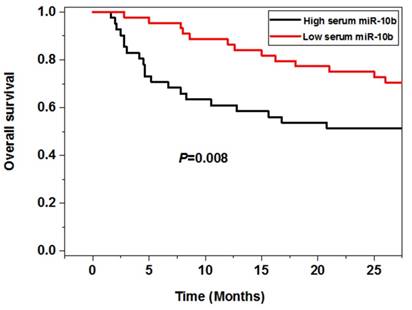
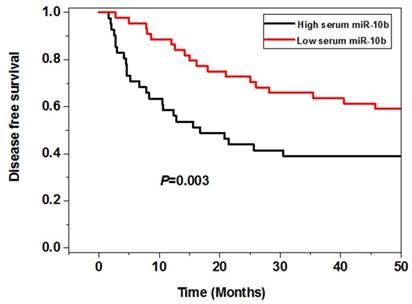
The survival analysis showed that the melanoma patients in the high serum miR-10b level group had worse 5 year overall (P=0.008) and disease free survival (P=0.003) than those in the low serum miR-10b level group, indicating high serum miR-10b was correlated with poor prognosis of melanoma (Figure 3).
The association between serum miR-10b and overall survival as well as disease free survival
Serum miR-10b was an independent prognostic factor for melanoma
For the overall survival, the multivariate analysis showed that lymph node metastasis (HR=2.82, 95%CI=1.14-4.21, P=0.038), TNM stage (HR=4.81, 95%CI=1.84-7.29, P=0.005) and serum miR-10b expression level (HR=3.54, 95%CI=1.47-5.80, P=0.012) were independent prognostic predictors of melanoma. Similar findings were observed on the disease free survival. Lymph node metastasis (HR=3.45, 95%CI=1.28-5.45, P=0.021), TNM stage (HR=5.62, 95%CI=2.04-8.93, P=0.002) and serum miR-10b expression level (HR=3.64, 95%CI=1.35-5.60, P=0.014) were also independent risk factors (Table 2).
Serum miR-10b was an independent prognostic factor for melanoma
| Variable | Overall survival | Disease free survival | ||||
|---|---|---|---|---|---|---|
| HR | 95% CI | P | HR | 95% CI | P | |
| Lymph node metastasis (Yes versus. No) | 2.82 | 1.14-4.21 | 0.038 | 3.45 | 1.28-5.45 | 0.021 |
| TNM stage (Ⅲ-Ⅳversus.Ⅰ-Ⅱ) | 4.81 | 1.84-7.29 | 0.005 | 5.62 | 2.04-8.93 | 0.002 |
| Serum miR-10b (High versus. low) | 3.54 | 1.47-5.80 | 0.012 | 3.64 | 1.35-5.60 | 0.014 |
Discussion
Serum miRNA profile is a promising research tool currently being applied to a number of human cancers. Although tissue biopsy is still the gold standard for the diagnosis of many diseases, it cannot monitor the disease status in real time. Deregulation of miRNAs expression has been reported in the serum or plasma samples of many cancer patients as cancer cells can secret miRNAs into the circulation system. As the levels of serum miRNAs are highly stable, reproducible and consistent (14), detecting their changes is an efficient tool for disease diagnosis and prognosis prediction. For instance, the expression of serum miR-206 was significantly decreased in patients with melanoma. In addition, reduced serum miR-206 was associated with unfavorable clinical outcome of melanoma, indicating miR-206 suppression might drive the progression of this malignancy (15).
Our data demonstrated that miR-10b was increased in highly invasive melanoma cell lines and the serum samples derived from patients with melanoma. In addition, serum levels of miR-10b were able to differentiate melanoma patients at different clinical stages and associated with various clinicopathological parameters. Moreover, melanoma patients with higher serum miR-10b had poorer overall and disease free survival. Finally serum miR-10b was an independent prognostic factor for melanoma. We speculated that melanoma cells could secret miR-10 into the blood, leading to its upregulation in the serum samples. In addition, miR-10 might play an oncogenic role in melanoma. Thus increased expression of serum miR-10 was correlated with worse clinical outcome of melanoma. Consistent with our results, tissue miR-10b was significantly increased in the melanoma with metastasis compared to those without metastasis. In addition, upregulation of tissue miR-10b was a poor prognostic predictor of melanoma (13). The expression level of a panel of miRNAs (miR-10b, 21, 200c, 373 and 520c) were upregulated in melanospheres, indicating these molecules might be important regulators of both metastasis and stemness potential (16). Jukic et al. (17) compared the differentially expressed miRNAs between the melanoma of young adults and older adults. MiR-10b expression was increased in older patients with melanoma, indicating miR-10b might be a useful biomarkers accounting for differences of melanoma at different ages. These data further confirmed our findings that increased serum miR-10b was a risk factor for prognosis of melanoma.
In addition to melanoma, serum miR-10b level has been shown to be upregulated in many types of cancer. The expression levels of serum miR-10b were significantly increased in breast patients with bone metastases in comparison with those without bone metastases or healthy subjects, indicating that serum miR-10b was a sensitive biomarker for predicting the bone metastasis of breast cancer (18). Jiang et al. (19) reported that circulating miR-10b was upregulated in patients with HCC. Its level was able to discriminate the HCC from chronic liver disease as well as healthy controls with high accuracy. Similarly, Huang et al. (20) recently identified a set of serum miRNAs (miR-10b-5p, miR-132-3p, miR-185-5p, miR-195-5p, miR-20a-3p and miR-296-5p) were increased in patients with gastric cancer compared with the healthy volunteers and had a good performance in diagnosis of gastric cancer. These studies suggest upregulation of serum miR-10b is not unique to melanoma.
In conclusion, we provide strong evidence supporting that serum miR-10b is overexpressed in melanoma and associated with poor prognosis of melanoma. This metastasis related miRNA may be clinically applicable for the management of patients with melanoma.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Miller AJ, Mihm MC Jr. Melanoma. N Engl J Med. 2006;355:51-65
2. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10-29
3. Bandarchi B, Ma L, Navab R, Seth A, Rasty G. From melanocytes to metastatic malignant melanoma. Dermatol Res Pract 2010. 2010
4. Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancers: biomarkers, functions and therapy. Trends Mol Med. 2014;20:460-469
5. Ha TY. MicroRNAs in human diseases: from cancer to cardiovascular disease. Immune Netw. 2011;11:135-154
6. Shenouda SK, Alahari SK. MicroRNA function in cancer: oncogene or a tumor suppressor? Cancer Metastasis Rev. 2009;28:369-378
7. Su F, Chen Y, Zhu S. et al. RAB22A overexpression promotes the tumor growth of melanoma. Oncotarget. 2016;7:71744-71753
8. Qiu HJ, Lu XH, Yang SS, Weng CY, Zhang EK, Chen FC. MiR-769 promoted cell proliferation in human melanoma by suppressing GSK3B expression. Biomed Pharmacother. 2016;82:117-123
9. Tehler D, Høyland-Kroghsbo NM, Lund AH. The miR-10 microRNA precursor family. RNA Biol. 2011;8:728-734
10. Parrella P, Barbano R, Pasculli B. et al. Evaluation of microRNA-10b prognostic significance in a prospective cohort of breast cancer patients. Mol Cancer. 2014;13:142
11. Jiang H, Liu J, Chen Y, Ma C, Li B, Hao T. Up-regulation of mir-10b predicate advanced clinicopathological features and liver metastasis in colorectal cancer. Cancer Med. 2016;5:2932-2941
12. Zhu Q, Gong L, Wang J. et al. miR-10b exerts oncogenic activity in human hepatocellular carcinoma cells by targeting expression of CUB and sushi multiple domains 1 (CSMD1). BMC Cancer. 2016;16:806
13. Saldanha G, Elshaw S, Sachs P. et al. microRNA-10b is a prognostic biomarker for melanoma. Mod Pathol. 2016;29:112-121
14. Chen X, Ba Y, Ma L. et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997-1006
15. Tian R, Liu T, Qiao L, Gao M, Li J. Decreased serum microRNA-206 level predicts unfavorable prognosis in patients with melanoma. Int J Clin Exp Pathol. 2015;8:3097-3103
16. Fomeshi MR, Ebrahimi M, Mowla SJ, Khosravani P, Firouzi J, Khayatzadeh H. Evaluation of the expressions pattern of miR-10b, 21, 200c, 373 and 520c to find the correlation between epithelial-to-mesenchymal transition and melanoma stem cell potential in isolated cancer stem cells. Cell Mol Biol Lett. 2015;20:448-465
17. Jukic DM, Rao UN, Kelly L. et al. Microrna profiling analysis of differences between the melanoma of young adults and older adults. J Transl Med. 2010;8:27
18. Zhao FL, Hu GD, Wang XF, Zhang XH, Zhang YK, Yu ZS. Serum overexpression of microRNA-10b in patients with bone metastatic primary breast cancer. J Int Med Res. 2012;40:859-866
19. Jiang L, Cheng Q, Zhang BH, Zhang MZ. Circulating microRNAs as biomarkers in hepatocellular carcinoma screening: a validation set from China. Medicine (Baltimore). 2015;94:e603
20. Huang Z, Zhu D, Wu L. et al. Six serum-based miRNAs as potential diagnostic biomarkers for gastric cancer. Cancer Epidemiol Biomarkers Prev. 2016 [Epub ahead of print]
Author contact
Corresponding author: Dr. Ru Zhao, Department of Plastic and Reconstructive Surgery, Peking Union Medical College Hospital, Chinese Academy of Medical Science and Peking Union Medical College, No.1 Shuaifuyuan Dongcheng District, Beijing, 100730, China. Tel: +86-10-69156114; E-mail: ruzhaopumccom

 Global reach, higher impact
Global reach, higher impact