Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(8):2359-2370. doi:10.7150/jca.50462 This issue Cite
Research Paper
Classification of Clear Cell Renal Cell Carcinoma based on Tumor Suppressor Genomic Profiling
1. The Fifth Hospital of Xiamen, Xiamen 361101, Fujian Province, China.
2. Key Laboratory of Optoelectronic Science and Technology for Medicine of Ministry of Education, College of Life Sciences, Fujian Normal University, Fuzhou 350117, Fujian Province, China.
Abstract
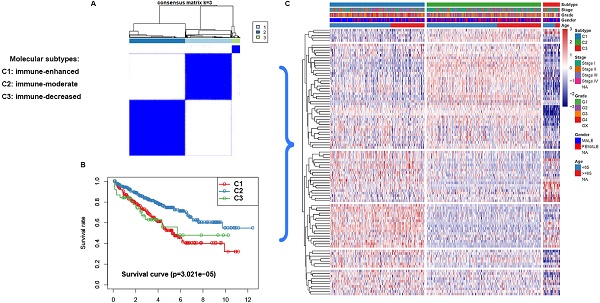
Clear cell renal cell carcinoma (ccRCC) is the most prevalent type of malignancy in adults. However, the clinical significance of tumor suppressor genes (TSG) is largely elusive. Herein, the expression profile TSGs and its clinical response in ccRCC were investigated. A total of 603 ccRCC samples from two cohorts (TCGA and ICGC) were retrieved in this study. Three molecular subtypes (C1, C2, and C3) were identified based on the TSGs expression profile in the TCGA dataset. Through Weighted Gene Correlation Network Analysis (WGCNA), six modules associated with three subtypes were identified. Pathway enrichment for the modules revealed that crucial pathways including p53 signaling and immune-related pathways were significantly enriched. We further focused on the relationship between immune infiltration level and subtypes, and found that subtype C1 was associated with higher immune infiltration level, subtype C2 was corresponding with medium immune infiltration level, whereas subtype C3 was correlated with lower immune infiltration level. Interestingly, C2 have a better survival outcome, while C1 and C3 showed a poor prognosis. Considering their survival difference, we then performed a differentially expression analysis between C2 and C1&3, and a total of 99 differentially expressed tumor suppressor genes (DETSGs) were identified. According to these DETSGs, 59 potential compounds with 28 mechanisms of action (MOA) were predicted using the Connectivity Map (CMap) database. Among these compounds, leflunomide, naftopidil, and ribavirin were the most prospective compounds for the treatment of ccRCC. In addition, we found that subtype C2 is more sensitive to sorafenib and sunitinib drugs, and C2 have more likelihood to be responded to immunotherapy. In summary, the three subtypes hinged on the tumor suppressor gene expression for ccRCC might contribute to understanding the underlying molecular mechanisms of ccRCC. Also, its potential compounds might offer guidelines for developing a novel treatment strategy of ccRCC.
Keywords: clear cell renal cell carcinoma, tumor suppressor genes, molecular subtype, tumor microenvironment, compounds, drug sensitivity, immunotherapy

 Global reach, higher impact
Global reach, higher impact