Impact Factor ISSN: 1837-9664
J Cancer 2019; 10(15):3381-3388. doi:10.7150/jca.31553 This issue Cite
Research Paper
Asymmetric Division Gene Neurl2 Mediates Twist2 Regulation of Self-Renewal of Mouse Lewis Lung Cancer Stem Cells
1. Laboratory of Translational Cancer Stem Cell Research, Institute of Life Sciences, Chongqing Medical University, Chongqing, China.
2. State Key Laboratory of Ultrasound Engineering in Medicine Co-Founded by Chongqing and the Ministry of Science and Technology, Chongqing, China.
3. Chongqing Traditional Chinese Medicine Hospital, Chongqing, China.
*These authors contributed equally.
Received 2018-11-16; Accepted 2019-4-23; Published 2019-6-9
Abstract
Cancer stem cells (CSCs) play an important role in tumor development. While Epithelial-Mesenchymal Transition (EMT) has been shown to promote CSC self-renewal, underlying mechanisms are unclear. Here we identified and characterized the requirement of twist2, the EMT transcription factor, for the regulation of self-renewal thus stemness of mouse Lewis lung CSCs both in vitro and in vivo. Further, we elucidated the role of neurl2, an asymmetric division gene for normal stem cells, in mediating the self-renewal promoting activity of twist2. Moreover, analysis of TCGA showed a positive correlation between the expression of twist2 and the development of lung adenocarcinoma, and a negative correlation between neurl2 and lung adenocarcinoma development. In summary, our study provides a new mechanistic insight of regulation of CSC self-renewal by EMT.
Introduction
With the isolation and characteristication of cancer stem cells (CSCs) from different solid tumors, CSCs are regarded as a subset of cells that have the ability of self-renewal and generation of heterogeneous tumor cells. However, mechanisms underlying CSC self-renewal regulation remain elusive [1]. While pluripotent genes of normal stem cells (NSCs) have been shown to be capable of regulating the self-renewal of CSCs, including Oct4 and Nanog, differences exist between the NSCs and CSCs in self-renewal regulation [2-4]. A body of studies have showed the promoting effect of Epithelial-mesenchymal transition (EMT) on the stemness of CSCs [5-8]. EMT is an important step in embryonic development in which epithelial cells are polarized into stromal cells with stronger motility, which enables cells to move long distances into the inner layer of the embryo and participate in the formation of internal organs [9]. Similarly, EMT also plays a role in wound healing and in the development of some diseases, such as fibrosis. In cancers, EMT is required for the initiation of metastasis. Alberto Martin and Amparo Cano suggested that EMT is not only associated with tumor metastasis, but also with the acquisition of stem cells [10, 11].
Twist1, Twist2, Snai1 and Zeb1 can induce the stemness properties of CSCs by promoting EMT process [11-13]. Alain Puisieux et al. showed that Twist1 maintains self-renewal of CSCs by directly activating BMI1, which encodes a comb protein and inhibits INK4A/ARF [9]. Zeb1 could activate the transcription of stemness related genes Bmi1, Sox2, Klf4 and Sox9 by inhibiting their miRNA regulator - the miR-200 family [9, 14]. Twist2 could regulate the self-renewal of CSCs through the Twist2-CD24-STAT3-Nanog pathway in hepatocellular carcinoma which is a highly conserved transcription factor of helix-loop-helix structure and similar to Twist1 [6]. In this study, we identified the asymmetric division gene Neurl2, as a new gene target of Twist2 that mediates the regulation of CSC self-renewal.
Stem cell renewal can be achieved by symmetrical division, as well as asymmetric divisions. Through these two types of cell division, stem cells can produce differentiated cells needed for growth and development, and at the same time maintain a stable number of undifferentiated stem cells for the maintenance of stemness [15]. The balance between symmetrically dividing stem cells and asymmetrically dividing stem cells is critical to tissue homeostasis, which is broken in cancers [16]. Costa G et al. conducted in vivo experiments in zebrafish lead to the hypothesis that asymmetrical division plays a similar role in the development and progression of tumors [17].
In this study, we elucidate that Twis2 can regulate the self-renewal of lung CSCs both in vitro and in vivo, through transcriptional inactivation of Neurl2, a asymmetric division gene of normal stem cells. As importantly, the interaction between Twist2 and Neurl2 is clinically relevant. Neurl2 emerges as a potential marker for lung cancer prognosis. In summary, our research provides a new insight for mechanism under the self-renew regulation of CSCs by EMT.
Results
The establishment and characterization of mouse Lewis lung cancer stem cell lines
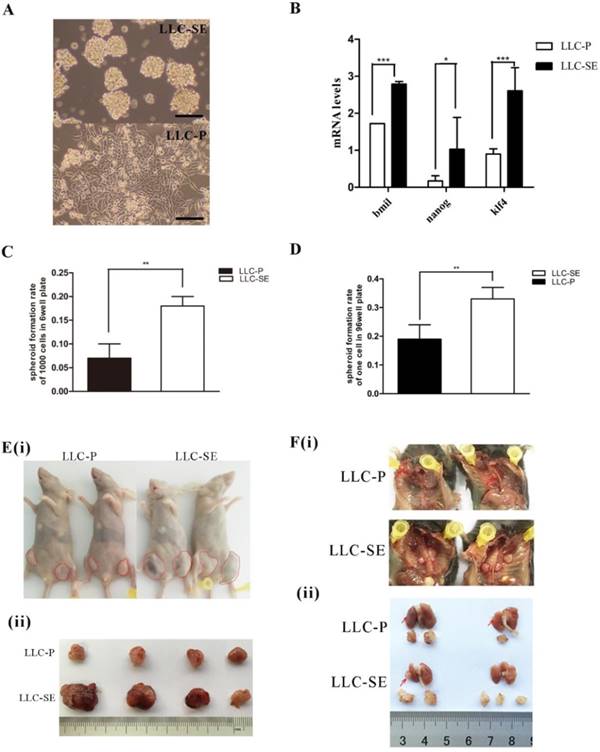
The mouse Lewis lung cancer stem cell line LLC-SE was obtained by selecting floating spheroidal cells from the culture of Lewis lung cancer parental cells (LLC-P) (Figure 1A). The enhancement in the stemness of LLP-SE cells was confirmed by elevated expression of stemness genes (Bmil, Klf4 and Nanog) (Figure 1B) and the increased spheroid formation rate and signle cell cloning rate (Figures 1C and D). In vivo, subcutaneous transplantation of 103 LLC-SE cells to the hinder legs of nude mice resulted in faster tumor growth and larger tumor burden compared to tumors derived from 103 LLC-P cells (Figure 1E(i) and (ii)). Further, orthotopic implantation of LLC-SE cells on the left lung in the syngeneic C57 mice showed more robust orthotopic tumor growth and more aggressive course of tumor progression than the LLC-P cells (Figure 1F(i) and (ii)). Collectively, these in vitro and in vivo observations indicate that LLC-SE cells exhibit enhanced stem cell properties and augmented tumorigeneicity.
The establishment and identification of mouse Lewis lung cancer stem cell lines. A.The morphology of LLC-P cells and LLC-SE cells, bar=120um. B. the analysis of mRNA expression of Bmi, Nanog and Klf4 in LLC-SE and LLC-P cells, the tata-Box binding protein gene (Tbp) was used as a reference control. C. the analysis of spheroid formation rate of 1000 cells in 6 well plate in LLC-SE cells and LLC-P cells. D. the analysis of spheroid formation rate of single cell in 96 well plate in LLC-SE cells and LLC-P cells. E. (i) and (ii) tumor formation in nude mice after injection. F. (i) and (ii) the procedure of orthotopic tumor transplantation in C57BL/6 mice.
Twist2 regualtes the stemness of LLC-SE
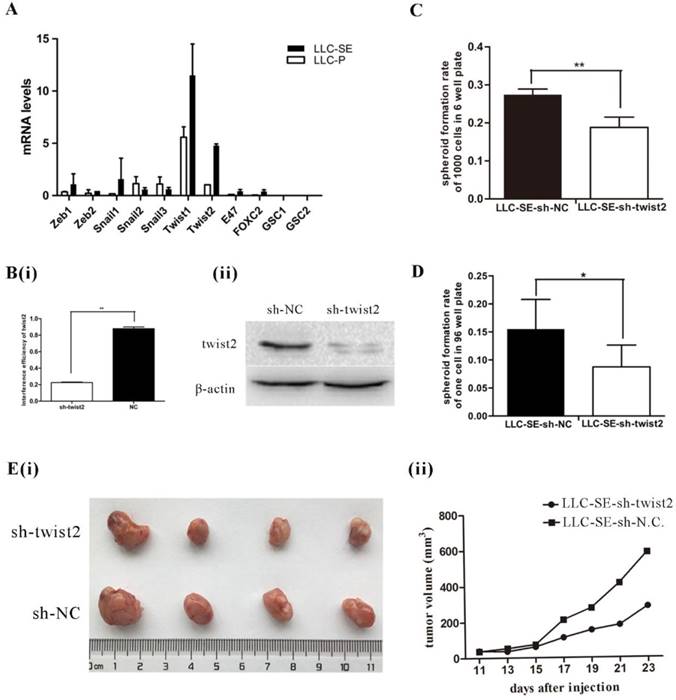
EMT transcript factors (TFs) are shown to promote the stemness of cancer stem cells (CSCs). Thus, the expression of 11 EMT TFs (Zeb1, Zeb2, Snail1, Snail2, Snail3, Twist1, Twist2, E47, FOXC2, GSC1 and GSC2) were measured by qPCR in LLC-SE and LLC-P, among which Zeb1, Zeb2, Snail1, Twist1, Twist2 and E47, FOXC2 were higher in LLC-SE than in LLC-P (Figure 2A). The expression of Twist2 showed the most significant difference between LLC-SE and LLC-P in all of the EMT TFs we assayed. Therefore, we focused the rest of the study on characterizing the role of Twist2 in regulating the stemness of lung cancer using the LLC-SE model. Interference of Twist2 expression by lentivirus effectively inhibited its expression, confirmed by qPCR and WB (Figure 2B(i) and 2B(ii)). Upon Twist2 interference, spheroid formation and single cloning rate were significantly reduced (Figure 2C and figure 2D), indicating hindered stemness. In vivo, subcutaneous transplantation of 104 LLC-SE-sh-twist2 in which Twist2 was targeted lead to inhibition of tumor growth compared to LLC-SE-sh-NC control tumors [Figure 2E(i) and 2E(ii)]. These observations indicate that Twist2 regulate the stemness of LLC-SE.
Twist2 enhances the stemness properties of LLC-SE cells. A. mRNA expression analysis of 11 Epithelial-Mesenchymal Transition (EMT) factors in the LLC-SE and LLC-P cells, the tata-Box binding protein gene (Tbp) was used as a reference control. B.(i) and (ii) verification of the efficiency of sh-mediated knockdown of Twist2 at mRNA and protein levels. C. the analysis of spheroid formation rate of 1000 cells in 6-well plate in LLC-SE-sh-NC cells and LLC-SE-sh-twist2 cells. D. the analysis of spheroid formation rate of single cell in 96 well plate in LLC-SE-sh-NC cells and LLC-SE-sh-twist2 cells. E.(i) and (ii) tumor formation in nude mice after injection derived from LLC-SE-sh-NC and LLC-SE-sh-twist2.
Neulr2 - the asymmetric division gene mediates Twist2 regualtion of stemness in LLC-SE
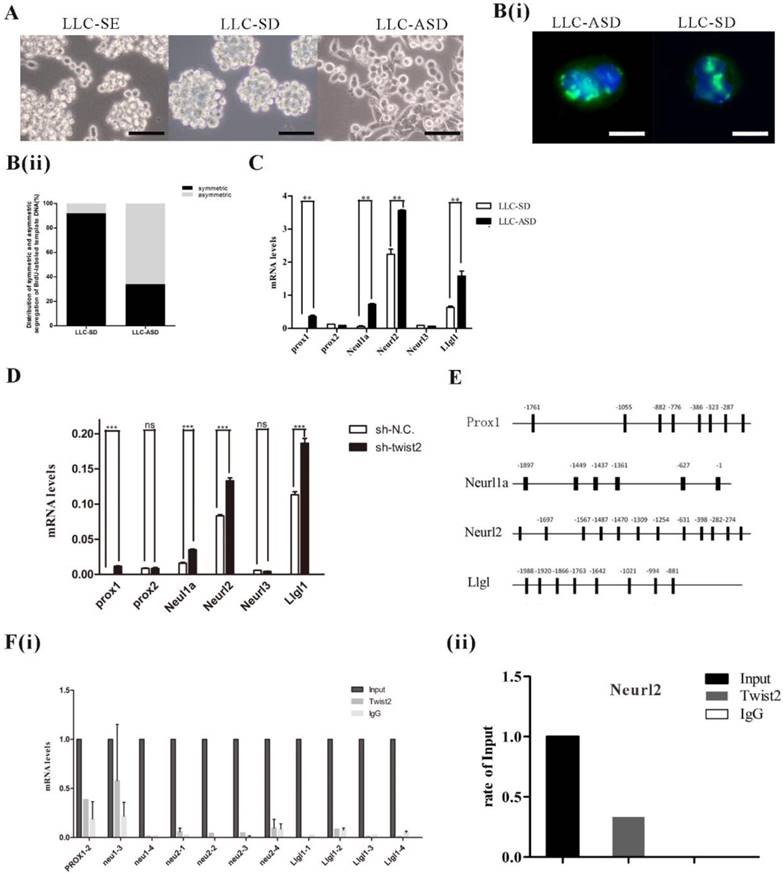
The proportion and balance between symmetrically dividing stem cells (SD) and asymmetrically dividing (ASD) stem cells that are tightly controlled and regulated in normal stem cells are broken in cancers. However, mechanisms regulating SD and ASD in CSCs remain largely unknown. In our previous study, the LLC-SD and LLC-ASD were derived from the LLC-SE and the expression of EMT TFs was higher in LLC-SD cells than that in LLC-ASD cells [14, 18]. SD and ASD ratio were detected by BrdU labeling (Figure 3A, 3B(i) and 3B(ii)). We assayed the expression of a set of known genes that regulate the asymmetric division of normal stem cells including Prox1, Neurl1a, Neurl2, and LIgl1. All displayed higher expression in LLC-ASD cells than in LLC-SD cells (Figure 3C). In order to explore the relationship between Twist2 and asymmetric division related genes, the expression of Prox1, Prox2, Neul1a, Neurl2 and LIgl1 were detected after Twist2 knockdown in LLC-SE cells. Inhibition of Twist2 in LLC-SE cells lead to the upregulation of this panel of asymmetric division genes (Figure 3D), exhibiting an inverse relationship. Twist2 functions often as a transcriptional inhibiting factor. We thus hypothesized that Twist2 could regulate the stemess of LLC-SE by inhibiting at least one of these asymmetric division genes. We analyzed probable Twist2 binding sites in Prox1, Neul1a, Neurl2 and LIgl1 promoter region predicted by JASPAR database (Figure 3E). ChIP assay was conducted to identify direct transcriptional targeting of Twist2. We found that Twist2 could bind to the Neurl2 promoter region (Figure 3F(i) and 3F(ii)), indicative of direct transcriptional inactivation of Neural2 by Twist2 in LLC-SE.
Twist2 regulates the stemness of LLC-SE via regulating the expression of asymmetric division related genes. A. The morphology of LLC-SE, LLC-SD and LLC-ASD cells, bar=120um. B.(i) Analysis of symmetric and asymmetric segregation of BrdU-labeled DNA during mitosis in anaphase LLC-SD and LLC-ASD, bar=120um. (ii) Quantification of SD and ASD cells in 100 dividing anaphase LLC-SD and LLC-ASD, respectively. C. the analysis of mRNA expression of asymmetric division genes in the LLC-SD and LLC-ASD, the tata-Box binding protein gene (Tbp) was used as a reference control. D. the analysis of mRNA expression of asymmetric division genes in LLC-SE-sh-NC and LLC-SE-sh-twist2, Tbp was used as a reference control. E. potential twist2 binding sites on the promoter regions of asymmetric division genes predicted through JASPAR database. G. (i) ChIP assay for determining the nature of interaction between Twist2 and asymmetric division genes. (ii) ChIP assay for determining the nature of interaction between Twist2 and Neurl2.
The analysis of the relationship between Twist2 and Neurl2 in clinical TCGA data
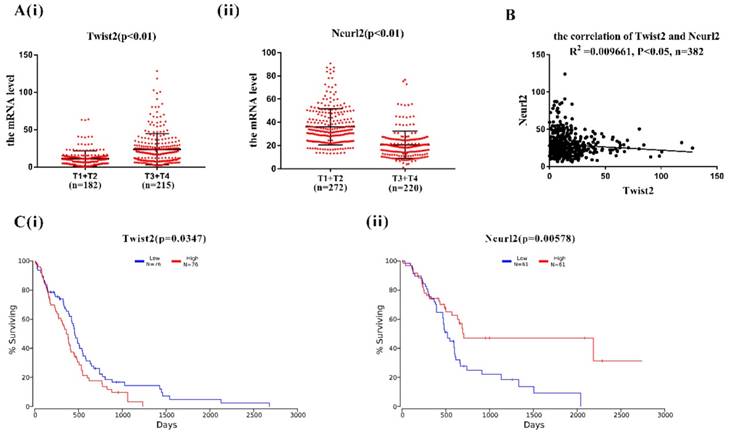
In order to explore the clinical relevance of Twist2-Neurl2, the expression of Twsit2 and Neurl2 were analysed in T1+T2 group and T3+T4 group according to human lung adenocarcinoma TMN classification. The expression of Twsit2 was higher in T1+T2 than T3+T4 and it was opposite for the Neurl2 (Figure 4A(i) and 4(ii)). The analysis of the correlation of Twsit2 and Neurl2 revealed a close correlation in human lung adenocarcinoma (Figure 4B). At last, the surviving curve of Twsit2 and Neurl2 were also analysed through OncoLnc. Lower level of Twist2 and higher level of Neurl2 conferred higher survival rate (Figure 4C). These results suggest that Twsit2 may promote lung CSC self-renewal by inhibiting Neurl2 in human lung adenocarcinoma, and Neurl2 may be explored as a new marker for lung cancer prognosis.
The analysis of correlation between Twist2 and Neurl2 in clinical TCGA data. A. (i) and (ii). the analysis of mRNA expression of Twist2 and Neurl2 in human lung adenocarcinoma from TCGA database, T1, T2, T3 and T4 were the TMN classification, n was the number of human lung adenocarcinoma samples. B. the analysis of correlation between Twist2 and Neurl2 in the human lung adenocarcinoma, n was the number of human lung adenocarcinoma samples. C. (i) and (ii) the survival curve of human lung adenocarcinoma patients were classified as the expression of Twist2 and Neurl2.
Discussion
Targeting cancer stem cells (CSCs) is an appealing approach for the cure of cancer [19-21]. The ability of CSCs to maintain self-renew as well as to undergo differentiation - though not to completion, bears resemblance to normal stem cells (NSCs). However, mechanisms underlying CSC self-renewal have been largely unknown. In this study, by using the LLC-SE CSC model we developed from the LLC-P model, we characterized the importance of EMT TF Twist2 in regulating the stemness of LLC-SE (Figure 2). Our findings in the adenomacarcinoma of the lung model are consistent with the literature report of Twist2 on regulating self-renewal of CSCs in the liver and the breast cancer [6, 22]. More importantly, we provided a new mechanism that underlies Twist2 regulation of stemness, i.e, the direct Twist2-Neurl2 transcriptional interaction (Figure 3F). The clinical relevance of Twist2-Neurl2 interaction was supported by the analysis of TCGA dataset, showing the potential of Neurl2 as a favorable prognostic marker for lung cancer (Figure 4).
Prior to this study, the role of Twist2 in lung cancer progression was explored in the context of EMT [9]. Its role in regulating lung CSC self-renewal has not been studied in lung cancer. In some recent reports, Twist2 was reported to regulate the stemness of hepatocellular carcinoma through CD24-STAT3-Nanog pathway [6]. Twist2 could increase the CD44high/CD24low CSC sub-population content in breast cancer [22]. In our study, the conventional stemness makers of normal stem cells (Oct4, Sox2) and lung CSCs CD133, CD44 and ALDH1 were not decreased after Twist2 knockdown (data not shown). Other studies also reported the unaltered expression of stenmness markers by EMT TFs [23]. In our previous study, we showed that Zeb1, another EMT TF, could regulate the symmetric division of mouse Lewis lung carcinoma stem cells through Numb mediated by miR-31[14]. Here, we provided an alternative mechanism CSC stemness regulation by Twist2 through its transcriptional inactivation of asymmetric geen Neurl2. Thus, EMT TFs can elicit robust regulation of lung CSC stemness by augmenting SD and inhibiting ASD division.
Symmetric division (SD) and asymmetric division (ASD) are unique forms of cell division that are essential for the maintenance of stemness of normal stem cells as well as cancer stem cells[15]. Recent study showed that some EMT TFs could regulate the self-renew of CSCs through targeting asymmetric division in colon cancer[24-26]. In our previous, SD and ASD cell lines were obtained from the LLC-SE in which the expression of EMT TFs were higher in SD than ASD, and the the relevance of Twist2 to ASD was confirmed by Twist2 knockdown in LLC-SE[14, 18]. Neurl2 was shown to underly Twist2 regulation of neurl cells differentiation [27]. Our study demonstrates that the transcriptional interaction between Twist2 and Neurl2 in normal stem cells also functions in lung CSCs. Moreover, we provided the first clinical evidence showing the clinical relevance of the interaction between Twist2 and Neurl2, and the potential of exploring Neurl2 as a favorable prognostic marker for human lung cancer.
In summary, our study indicated that EMT TF Twist2 can regulate the stemness of lung CSCs by transcriptional inactivation of the asymmetric division factor Neurl2.
Materials and methods
Cell culture
The mouse Lewis lung carcinoma parental cell line (LLC-P) was purchased from the cell bank of the Shanghai Institute of life sciences, Chinese Academy of Sciences. LLC-P cells were cultured with DMEM culture medium contained 1% Amphotericin B, 1% penicillin-streptomycin and 10% FBS. The Lewis Lung Carcinoma-Spheroid Enrichment cells (LLC-SE) and the Lewis Lung carcinoma-Symmetric Division (LLC-SD) cells were cultured in the DMEM culture medium with 2% B27, 1% Amphotericin B and 1% penicillin-streptomycin. And the LLC-ASD cells were cultured in this culture medium containing DMEM ,1% Amphotericin B, 1% penicillin-streptomycin and 5% FBS. All cells were kept in the incubator, with 5% CO2 and constant humidity and at 37℃.
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
Total RNAs were isolated with Trizol agent. In brief, 1mL Trizol agent were added into the EP tube containing cells, incubated on the ice for 10 minutes, followed addition of 200 uL chloroform and oscillation. Thereafter, the mixture was centrifuged in 4℃ for 15 minutes followed by addition of Isoamyl alcohol and incubation on the ice for 10 minutes. At last, the mixture was centrifuged in 4℃ for 10 minutes followed addition of 70% ice-cold ethanol and preceded with another centrifugation in 4℃ centrifuge for 10 minutes. The pellets were dissolved using 20-30 ul DEPC water. RT-PCR was conducted using PrimeScript RT Master Mix (Takara) according to the manufacturer's instructions.
Real-time Quantitative PCR
The reaction system of QPCR was 10µL, containing 5µL mastermix agent, 0.8µL Cdna, 0.8µL primer and 3.4µL DEPC water. Triplicates were set up for every sample. PCR was conducted according to the manufacturer's instructions. The sequences of PCR primers are listed in Table 1.
PCR Primer Sequence
| Gene name | Forward primers | Reverse primers |
|---|---|---|
| Tbp | CCGTGAATCTTGGCTGTAAAC | CAGTTGTCCGTGGCTCTCTT |
| Bmi1 | ATCCCCACTTAATGTGTGTCCT | CTTGCTGGTCTCCAAGTA ACG |
| Nanog | TTAGAAGCGTGGGTCTTGGT | CCCTCA AACTCCTGGTCCTT |
| Klf4 | GGACCACCTTGCCTTACACA | GACTTGCTGGGAACTTGACC |
| Zeb1 | CTCCCAGTCAGCCACCTTTA | TAACTCCATCCAGCGGTAGG |
| Zeb2 | TTTCCTGCCCTCTTTGTAGC | CCTTGGGTTAGCATTTGGTG |
| Snail1 | CACACGCTGCCTTGTGTCT | GGTCAGCAA AAGCACGGTT |
| Snail2 | TGGTCAAGAAACATTTCAACGCC | GGTGAGGATCTCTGGTTTTGGTA |
| Snail3 | ACAGCGAAC TGG ACA CACAC | GGGTAAAGGAGAGTGGAG TGG |
| Twist1 | GGACAAGCTGAGCAAGAT TCA | CGGAGA AGGCGT AGCTGAG |
| Twist2 | GTCTCAGCTACGCCTTCTCC | AGATGTGCAGGT GGGTCCT |
| E47E2a | GCAAACGAAGGCAGAGTAGG | TGGTCAAGAGACACCTCA AGC |
| Foxc2 | AACCCAACAGCAAACTTCCC | GCGTAGCTCGATAGGGCAG |
| Gsc1 | AACGCCGAGAAGTGGAACAAG | CCGAGTCCAAATCGCTTT TACC |
| Gsc2 | CGTGGAGGTCTGGTTCAAG | CGGACAGCATCA ACA ACTCT |
Western Blot
12% of resulving gel and 6% of condensing gel were used in this study. The voltage of electrophoresis was 120V while the current for member transfer was 350 mA. After transfer, the membrane was blocked with 5% of BSA, followed by incubation with the primary antibody (11752-1-AP, Proteintech) at 4℃ for 12h and completed with incubation with the secondary antibody (SA00001-2, Proteintech) at room temperature for 1 hour. The results were acquired using automatic analysis system of electrophoresis gel imaging.
Subcutaneous tumor transplantation in nude mice
BALB/cA-nude mice were purchased from Beijing Hua Gao Kang Bio Polytron Technologies Inc and were examined by the Institute of medical laboratory animal research, Chinese Academy of Medical Sciences. 103 or 104 cells were injected to each side of the hinder leg of each mouse. After tumor cell transplantation, tumor volume was measured every two days between Day 11 day and Day 23 and was plotted.
Orthotopic tumor cell transplantation in C57/BL6 mice
C57/BL6 mice were purchased from the Laboratory Animal Center of Chongqing medical university, and the protocol was approved by the animal ethics committee. After being anaesthetized with 1% pentobarbital, the left chest was exposed and the skin was disinfected with 75% alcohol. A longitudinal incision was made at the cardiopulmonary junction, 1x105 cells were injected. 3 weeks later, the extent of metastasis to the left lung, the right lung and the thymus as well as to the walls of the chest were examined and recorded.
Immunofluorescence
The cells were treated with BrdU for at least seven days, followed by BrdU withdrawal for at least two days. Cells were harvested, fixed with ice-cold 70% ethanol for 30 minutes, and ttreated with hydrochloric acid for 1 hour. Cells were washed with PBS for three times, the slides were blocked with 10% BSA solution in the shaker at 37℃ for 1 hour. Thereafter, the cells were incubated with the first antibody (anti-BrdU, Millipore) at 4℃ for 12 hour with gentle agitation, followed by incubation with the secondary antibody (Proteintech) at 37℃ for 1 hour. Images were taken by OLYMPUS inverted fluorescence microscope.
Chromatin immunoprecipitation (ChIP) assay
The ChIP assay was performed with the Chromatin immunoprecipitation (ChIP) Assay Kit of Beyotime Biotechnology according to manufacturer's instructions. Firstly, cells were cross-linked in 300 µl formaldehyde for 10 min at room temperature and the reaction was stopped by incubation with 500 µl 2.5mol/L Glycine solution at room temperature for 5 min. Then, the mixture was poured out and the PBS with 10x PMSF buffer was added into the precipitation and centrifuged at 2000g for 4 min. Thereafter, 1 ml SDS/lysis buffer was added into the precipitation and the mixture was broken via sonication, followed by incubation with the first antibody (Abcam) at 4℃ for 12 hour. Then 60ul protein A Agarose were added and incubated with gentle oscillation at 4℃. The mixture was then centrifuged and washed with Low salt solution, high salt solution and LiCl solution sequnetially. Finally, the precipitates were washed twice wth TE buffer. DNA was extracted using a Extraction Kit from Tiangen company. At last, QPCR assay was performed using the freshly extracted DNA.
Lentivirus packaging and infection
Stable knockdown of Twist2 was achieved with infection of cells with pLL3.7 in the presence of 8 μg/mlpolybrene (Genepharma, China) for 12 hours, and the medium was changed at 72 hours post infection.
TCGA Data Analysis
The mRNA expression of Twist2 in lung adenocarcinoma was evaluated by TCGA Research Network (http://cancergenome.nih.gov). To analyze the survival of patients with lung adenocarcinoma, patient samples were analysed by OncoLnc (http://www.oncolnc.org).
Statistical analysis
Data were analyzed by one-way analysis of variance using GraphPad Prism software and presented as mean ± SD or SEM. Differences were considered statistically significant when P < 0.05.
Acknowledgements
We thank the members in the Laboratory of Translational Cancer Stem Cell Research who are not listed as the authors for their encouragement. This work was supported by the National Natural Science Youth Fund (Grant No.81602596), the National Natural Science Fund (Grant No.81672908) and the Science and Technology Project Affiliated to the Education Department of Chongqing (Grant No. KJQN201800426).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Clarke MF, Dick JE, Dirks PB, Eaves CJ, Jamieson CH, Jones DL. et al. Cancer stem cells-perspectives on current status and future directions: AACR Workshop on cancer stem cells. Cancer research. 2006;66:9339-44
2. Reya T, Morrison SJ, Clarke MF, Weissman IL. Stem cells, cancer, and cancer stem cells. Nature. 2001;414:105-11
3. Jen J, Tang YA, Lu YH, Lin CC, Lai WW, Wang YC. Oct4 transcriptionally regulates the expression of long non-coding RNAs NEAT1 and MALAT1 to promote lung cancer progression. Molecular cancer. 2017;16:104
4. Shan J, Shen J, Liu L, Xia F, Xu C, Duan G. et al. Nanog regulates self-renewal of cancer stem cells through the insulin-like growth factor pathway in human hepatocellular carcinoma. Hepatology (Baltimore, Md). 2012;56:1004-14
5. Liu CW, Li CH, Peng YJ, Cheng YW, Chen HW, Liao PL. et al. Snail regulates Nanog status during the epithelial-mesenchymal transition via the Smad1/Akt/GSK3beta signaling pathway in non-small-cell lung cancer. Oncotarget. 2014;5:3880-94
6. Liu AY, Cai Y, Mao Y, Lin Y, Zheng H, Wu T. et al. Twist2 promotes self-renewal of liver cancer stem-like cells by regulating CD24. Carcinogenesis. 2014;35:537-45
7. Li P, Yang R, Gao WQ. Contributions of epithelial-mesenchymal transition and cancer stem cells to the development of castration resistance of prostate cancer. Molecular cancer. 2014;13:55
8. Chaffer CL, Marjanovic ND, Lee T, Bell G, Kleer CG, Reinhardt F. et al. Poised chromatin at the ZEB1 promoter enables breast cancer cell plasticity and enhances tumorigenicity. Cell. 2013;154:61-74
9. Puisieux A, Brabletz T, Caramel J. Oncogenic roles of EMT-inducing transcription factors. Nature cell biology. 2014;16:488-94
10. Martin A, Cano A. Tumorigenesis: Twist1 links EMT to self-renewal. Nature cell biology. 2010;12:924-5
11. Singh A, Settleman J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene. 2010;29:4741-51
12. Yu M, Smolen GA, Zhang J, Wittner B, Schott BJ, Brachtel E. et al. A developmentally regulated inducer of EMT, LBX1, contributes to breast cancer progression. Genes & development. 2009;23:1737-42
13. Gupta GP, Massague J. Cancer metastasis: building a framework. Cell. 2006;127:679-95
14. Wang J, Zhou T, Sun Z, Ye T, Zhou S, Li J. et al. Zeb1 Regulates the Symmetric Division of Mouse Lewis Lung Carcinoma Stem Cells through Numb mediated by miR-31. International journal of biological sciences. 2018;14:1399-410
15. Santoro A, Vlachou T, Carminati M, Pelicci PG, Mapelli M. Molecular mechanisms of asymmetric divisions in mammary stem cells. EMBO Rep. 2016;17:1700-20
16. Mukherjee S, Kong J, Brat DJ. Cancer stem cell division: when the rules of asymmetry are broken. Stem cells and development. 2015;24:405-16
17. Costa G, Harrington KI, Lovegrove HE, Page DJ, Chakravartula S, Bentley K. et al. Asymmetric division coordinates collective cell migration in angiogenesis. Nature cell biology. 2016;18:1292-301
18. Wang J, Sun Z, Liu Y, Kong L, Zhou S, Tang J. et al. Comparison of tumor biology of two distinct cell sub-populations in lung cancer stem cells. Oncotarget. 2017;8:96852-64
19. Lobo NA, Shimono Y, Qian D, Clarke MF. The biology of cancer stem cells. Annual review of cell and developmental biology. 2007;23:675-99
20. Dalerba P, Cho RW, Clarke MF. Cancer stem cells: models and concepts. Annual review of medicine. 2007;58:267-84
21. Tang DG. Understanding cancer stem cell heterogeneity and plasticity. Cell research. 2012;22:457-72
22. Fang X, Cai Y, Liu J, Wang Z, Wu Q, Zhang Z. et al. Twist2 contributes to breast cancer progression by promoting an epithelial-mesenchymal transition and cancer stem-like cell self-renewal. Oncogene. 2011;30:4707-20
23. Xu H, Mu J, Xiao J, Wu X, Li M, Liu T. et al. CD24 negative lung cancer cells, possessing partial cancer stem cell properties, cannot be considered as cancer stem cells. American journal of cancer research. 2016;6:51-60
24. Bu P, Chen KY, Chen JH, Wang L, Walters J, Shin YJ. et al. A microRNA miR-34a-regulated bimodal switch targets Notch in colon cancer stem cells. Cell stem cell. 2013;12:602-15
25. Liang R, Li Y, Wang M, Tang SC, Xiao G. MiR-146a promotes the asymmetric division and inhibits the self-renewal ability of breast cancer stem-like cells via indirect upregulation of Let-7. Cell Cycle. 2018;17:1445-56
26. Hwang WL, Jiang JK, Yang SH, Huang TS, Lan HY, Teng HW. et al. MicroRNA-146a directs the symmetric division of Snail-dominant colorectal cancer stem cells. Nature cell biology. 2014;16:268-80
27. Song R, Koo BK, Yoon KJ, Yoon MJ, Yoo KW, Kim HT. et al. Neuralized-2 regulates a Notch ligand in cooperation with Mind bomb-1. The Journal of biological chemistry. 2006;281:36391-400
Author contact
Corresponding author: Jianyu Wang, Mailing address: 1 Yi Xue Yuan Road, Yuzhong District Chongqing, Chongqing Medical University, 400016, P.R. China. Phone and Fax number: +86-023-63662443, +86-023-63662443; Email address: wjy2003123com

 Global reach, higher impact
Global reach, higher impact