Impact Factor ISSN: 1837-9664
J Cancer 2018; 9(19):3620-3625. doi:10.7150/jca.26689 This issue Cite
Research Paper
Inhibition of Phospholipase D1 mRNA Expression Slows Down the Proliferation Rate of Prostate Cancer Cells That Have Transited to Androgen Independence
1. Department of Medical Laboratory, the First Affiliated Hospital of Wenzhou Medical University, Zhejiang Wenzhou, 325000, China
2. Liver Disease Center, the First Affiliated Hospital of Wenzhou Medical University, Zhejiang Wenzhou, 325000,China
*Co-first author: Keqing Shi with the same contribution to this work.
Received 2018-4-15; Accepted 2018-8-6; Published 2018-9-8
Abstract
To explore the role of phospholipase D1 (PLD1) mRNA in transition of prostate cancer (PCa) cells to androgen independence, we used Arraystar Human LncRNA Microarray V3.0 to detect and compare the differential expression of PLD1 and its signaling pathway-related gene in standard androgen dependence prostate cancer (ADPC) cell line LNCaP before and after the occurrence of androgen independence prostate cancer (AIPC) transition. In addition, we used the shRNA lentiviral vector to inhibit the PLD1 mRNA expression and observed its effect on LNCaP cell proliferation after AIPC transition by using MTS method. The results showed that the expression level of PLD1 mRNA was increased by 373-fold after AIPC transition (P<0.05); the PI3K/AKT signaling pathway-related gene expression was also elevated (P<0.05); the growth rate of LNCaP cells that had transited to androgen independence was reduced by about 30% when the PLD1 mRNA expression was inhibited by the shRNA lentivirus as compared with the negative control group (P<0.05). All these results suggest that PLD1 mRNA and the related PI3K/AKT signaling pathway may play an important role in AIPC. Down-regulating the expression of PLD1 mRNA could to some extent inhibit the proliferation rate of PCa cells after AIPC transition.
Keywords: phospholipase D1 (PLD1), prostate cancer, androgen independence, microarray, lentivirus
Introduction
Prostate cancer (PCa) is the leading cause of cancer-related death in men in European and American countries [1]. The incidence and mortality of PCa in China are also on the increase in recent years and has become a major disease endangering the health of old men [2]. In addition to surgery and radiotherapy, androgen deprivation therapy (ADT) is the standard treatment for advanced PCa. However, most PCa patients may experience the transition from androgen-dependent prostate cancer (ADPC) to androgen-independent prostate cancer (AIPC) within 2-3 years after receiving ADT, resulting in an extremely high mortality rate [3, 4]. Therefore, the mechanism underling the occurrence of AIPC has long been a hot point of research, mainly involving amplification of the androgen receptor (AR) gene, mutation of AR, and abnormality of the apoptosis-regulatory gene [5-9].
In our previous study, we used ultra-performance liquid chromatography-electrospray ionization-quadrupole-time of flight mass spectrometry (UPLC-ESI-Q-TOF MS) and gas chromatography-mass spectrometry (GC-MS) to detect metabolic products after LNCaP cells underwent AIPC transition and discovered 23 potential differentially expressed metabolic products, including 13 phosphatidyl cholines (PC), 2 amides, 2 lysophosphatidylcholines (LPC), and a sphingomyelin (SM), a phosphatidyl ethanolamine (PE), a phytosphingosine, and other undefined compounds. Of them, PCs was abnormally increased by (1.22-7.69) fold after LNCaP cell transition to AIPC [10]. Phospholipase D (PLD) is an important enzyme that regulates phospholipid metabolism in the organism [11, 12]. In mammals, PLD exists in two forms: PLD1 and PLD2 [13, 14]. Numerous studies have demonstrated that PLD1 plays an important role in tumor occurrence, differentiation, apoptosis and metastasis [15, 16]. The aim of the present study was to determine whether PLD1 also played an important role in PCa cell transition to AIPC.
Materials and Methods
Instruments and reagents
The instruments and reagents used in this study were standard ADPC cell line LNCaP (Shanghai Cell Bank of the Chinese Academy of Sciences, Shanghai, China); F12, phenol red-free DMEM/F12, common fetal bovine serum (FBS) and charcoal stripped FBS (Gibco, USA); total RNA extraction reagent TRIzol (Invitrogen, USA); Revert Aid First Strand cDNA synthesis kit (Fermentas, Lithuania); ABI SYBR Green PCR Master Mix (ABI, USA); Arraystar Human LncRNA Microarray V3.0 (PT Solution Biotechnology CO., LTD., Shanghai, China); CCK-8 cell proliferation-toxicity test kit (Dojindo China Co., Ltd.); CO2 incubator (Thermo, USA); ABI 7500 real-time fluorescence quantitative PCR (ABI, USA).
Cell culture
ADPC LNCaP cells were cultured in F12 containing 10% FBS. AIPC LNCaP-AI cells that had been successfully induced in our laboratory in our prior work were cultured in phenol red-free DMEM/F12 containing 10% fetal bovine serum (FBS) treated with activated carbon absorption in a 37°C 5% CO2 incubator. The medium was replaced 3-4 days after cell passaging, and about 7 days later confluent cells were digested with 0.25% trypsin for passage culture.
mRNA microarray
Total RNA was extracted from LNCaP and LNCaP-AI cells, the quality and quantity of which were verified by agarose gel electrophoresis. mRNA was isolated by using the mRNeasy Plus Mini Kit, labeled, hybridized, washed, and finally scanned with a laser confocal scanner. After quantile normalization and subsequent treatment of the original data by using GeneSpring GX v12.1 Software, mRNAs with more than 2-fold differential expression were screened out.
Verification of mRNA expression by quantitative real-time PCR
Cells were collected to extract total RNA, which was then reverse transcribed to cDNA. qPCR was performed by using cDNA (1μL) as the template and addition of 1.5μL upper- and down-stream primers (10μmol/L), 6μL RNase-free H2O and 10μL ABI SYBR Green PCR Master Mix, totaling 20μL under the condition of 95°C 15 min, 95°C 10 s, 58°C 40 s, totaling 40 cycles. The samples were stored at 4°C. Each sample was tested twice and the mean value was used for comparison. Using GAPDH as housekeeping gene of internal reference, the relative expression level of various target genes in LNCaP and LNCaP-AI cells, as well as the fold increase of the target genes in LNCaP-AI cells relative to LNCaP cells was calculated using the equation: ΔCt(target gene)=Ct(target gene)-Ct(GAPDH of target cell), and ΔΔCt=ΔCt(LNCaP-AI)-ΔCt(LNCaP).
Construction of the lentiviral vector and detection of cell proliferative activity by MTS assay
According to human PLD1 mRNA (Accession: NM_002662), gene sequences were screened by software to design two specific shRNA interference sequences, to both ends of which appropriate restriction-enzyme cutting sites were added to complete the construction of the vector. In addition, a termination signal (TTTTT) was added to the positive 3' end, and a termination signal compensatory sequence was added to the reverse 5' end (Table 3). Clones were cultured by connecting competent cells (DH5α) of product transformation, from which plasmids were extracted for enzyme digestion identification and sequencing, followed by transfection of 293T cells with Lipofectamine TM2000. The lentiviral RNA interference (RNAi) vector contained green fluorescence protein (GFP) in all groups. LNCaP-AI cells were infected with the lentivirus. Sequencing, viral packaging and titer determination were performed by Shanghai Genechem Co., Ltd (Shanghai, China). Knowing that the knockout efficiency of the PC positive targeted virus on the target gene is more than 70%; the target gene product plays an important role in the transcriptional process of mammalian cells; and silencing of this target gene can markedly inhibit cell proliferation in multiple tumor cell lines, it is often used as a positive reference in cell proliferation-related experiments to demonstrate the stability of the test system. Optical density was measured at 570nm (OD570) by MTS method in the positive control group, LNCaP-AI PLD1 mRNA shRNAi group, and negative control group for 5 consecutive days to map out cell growth curves.
Fold increase of gene mRNAs including PLD1 and AKT3 in LNCaP-AI relative to LNCaP group
| mRNA name | ΔCt(LNCaP) | ΔCt(LNCaP-AI) | ΔΔCt | fold increase | P |
|---|---|---|---|---|---|
| AKT3 | 15.16±0.96 | 7.83±0.84 | -(7.33±1.03) | 182.43±120.37 | 0.0084 |
| AR | 1.65±0.65 | -(3.90±0.43) | -(5.55±0.11) | 46.92±3.55 | 0.0046 |
| BAD | 1.72±0.22 | 3.41±0.25 | 1.69±0.03 | 0.31±0.01 | 0.0195 |
| C-raf | 8.27±0.27 | 5.76±0.14 | -(2.51±0.57) | 5.93±2.28 | 0.0156 |
| K-ras | 7.09±0.09 | 3.24±0.12 | -(3.84±0.02) | 14.30±0.22 | 0.0011 |
| mTOR | 0.48±0.17 | -(0.56±0.26) | -(1.28±0.26) | 2.43±0.84 | 0.0217 |
| PI3K | 1.28±0.34 | 0.23±0.21 | -(1.24±0.27) | 2.04±0.22 | 0.0383 |
| PLD1 | 18.31±8.31 | 9.78±0.41 | -(8.53±0.29) | 373.44±75.80 | 0.0044 |
| S6K1 | 6.81±0.64 | 4.49±0.51 | -(2.32±0.13) | 5.00±0.91 | 0.0238 |
Note: ΔCt(target gene)=Ct(target gene)-Ct(GAPDH of target cell), ΔΔCt=ΔCt(LNCaP-AI)-ΔCt(LNCaP);
Ct(GAPDH of LNCaP)=24.47±0.21, Ct(GAPDH of LNCaP-AI)=25.36±0.16.
Design of two shRNA sequences specific to human PLD1mRNA as the template
| 5' added base group | STEM | Loop | STEM | 3' added base group | |
|---|---|---|---|---|---|
| sequence 1 | |||||
| a | CCGG | AACTGGAAGATTACTTGACAA | CTCGAG | TTGTCAAGTAATCTTCCAGTT | TTTTTG |
| b | AATTCAAAAA | AACTGGAAGATTACTTGACAA | CTCGAG | TTGTCAAGTAATCTTCCAGTT | |
| sequence 2 | |||||
| a | CCGG | AAGAGGTGGCTGGTGGTGAAG | CTCGAG | CTTCACCACCAGCCACCTCTT | TTTTTG |
| b | AATTCAAAAA | AAGAGGTGGCTGGTGGTGAAG | CTCGAG | CTTCACCACCAGCCACCTCTT | |
Data analysis
Statistical treatment was conducted by using SPSS 13.0. Measurement data were treated by independent-sample t test. Values of P<0.05 were considered statistically significant.
Results
Results of Arraystar Human LncRNA Microarray V3.0
Changes in mRNA differential expression in LNCaP cells before and after AIPC transition were analyzed by using Arraystar Human LncRNA Microarray V3.0 (containing 30,586 lncRNAs and 26,109 mRNAs). It was found that PLD1 mRNA in LNCaP cells was increased after AIPC transition, and at the same time its downstream Ras-PI3K-AKT was activated and related molecule mRNA was increased markedly. We therefore used the result of RT-PCR to verify the reliability of the microarray result and found that the corresponding gene mRNAs were increased (Table 1, 2 and Fig. 1), confirming that the results obtained from the two methods were consistent.
Sequences of gene primers including PLD1 and AKT3
| Seq ID | Gene | Forward primer | Reverse primer |
|---|---|---|---|
| NM_001206729 | AKT3 | GTTACCTTTCTACAACCAGGACCATG | GGTCCTCCACCAAGGCGTTTA |
| NM_000044 | AR | CGGCATGGTGAGCAGAGTGC | CAAAACATGGTCCCTGGCAGTC |
| NM_004322 | BAD | TTCCAGATCCCAGAGTTTGAGC | CCCATCCCTTCGTCGTCCT |
| NM_002880 | c-raf | CTTCTTTGACTATGCGTCGTATGC | GGGCTGAAGGTGAGGCTGATT |
| NM_004985 | K-ras | GTAGTTGGAGCTGGTGGCGTAG | CCTCATGTACTGGTCCCTCATTG |
| NM_004958 | mTOR | TTAGAGGACAGCGGGGAAGGC | GCAGGTCCGGTTCCAAGCATC |
| NM_005027 | PI3K | ACAGAGCGTGGGCTGGATG | ACTGCTCGGGCAAGTCGG |
| NM_001130081 | PLD1 | GCCTATGGAAGGTGGGACGACA | CATTGCGGCAGGTGGGAGG |
| NM_003161 | S6K1 | TGGACCAGCCAGAGGACGC | CCTTTACCAAGTACCCGAAGTAGC |
The signaling pathway with PLD1 participation was activated in LNCaP cells in the androgen-free environment.
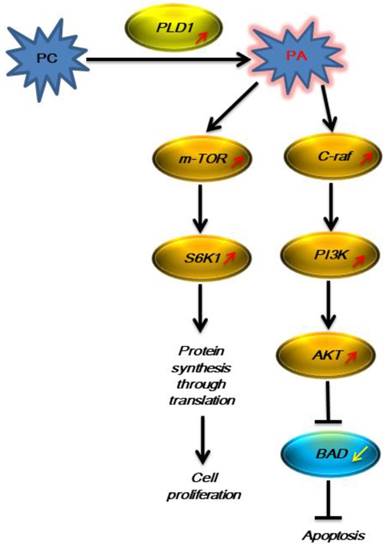
Impact of PLD1 gene silencing on the proliferation of LNCaP-AI cells
The knockout efficiency of the PC positive targeted virus on the target gene was more than 70%, confirming that the test system was stable. shCtrl refers to normal target cell+ negative control virus infection group; shPC refers to normal target cell+ positive gene shRNA virus infection group; shPLD1 refers to normal target cell+ PLD1 gene shRNA virus infection group (Table 3, Fig. 2). OD570/fold refers to OD570 multiples during day 1-5 relative to day 1 in all groups, representing the daily fold change. The OD570/fold value at day 5 was used to judge fold change of cell proliferation in the shCtrl (negative control) group relative to the experimental group. Fold change≥1.2 means that the proliferation rate in the experimental group was slowed down as compared with the shCtrl group, based on which the RNAi lentivirus-specific target gene of the experimental group was judged as the proliferation-related positive gene.
Change in the growth rate of LNCaP-AI cells after lentiviral transfection. A. Evaluation of the lentivirus transduction rate, which was more than 80% as calculated by cellular enumeration using fluorescence and light microscopy, showing that the lentiviral vector stably expressing shRNA targeting PLD1 was successfully constructed (×100). B. The cell growth state of LNCaP cells within 5 consecutive days after PLD1 mRNA shRNA transfection (GFP labeling, ×10). shPLD1: Fluorescence expression in the group transfected with PLD1 mRNA shRNA lentivirus. shCtrl: Fluorescence expression in the negative control group. shPC: Fluorescence expression in the positive control group.
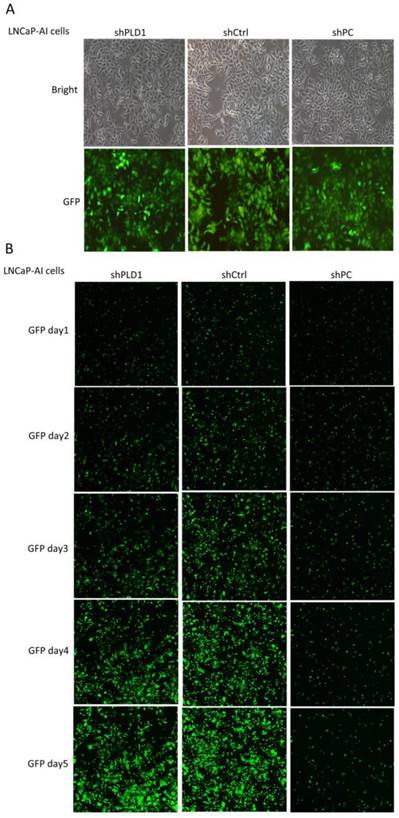
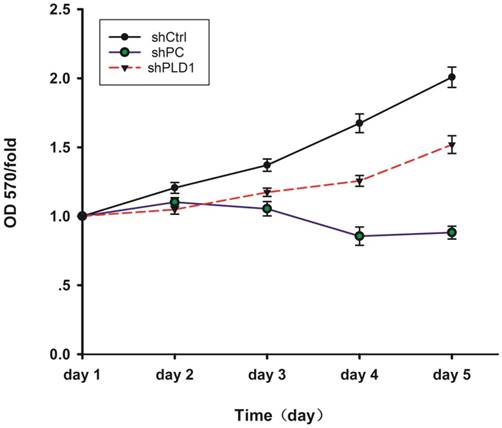
The result of MTS showed that cell proliferation in the shCtrl group was normal and inhibited in the shPC group, indicating that the test system of the present study was normal. Fold change of cell proliferation in the shCtrl group versus the shPLD1 group was 1.3051 at day 5, indicating that the proliferation rate of LNCaP-AI cells was reduced by 30.51% after inhibition of the PLD1 mRNA expression (Fig. 3).
MTS detection of OD570 changes in LNCaP-AI cells at different days after PLD1 mRNAi. The figure shows the cell proliferation curves in the three groups. The cell proliferation abilitywas assessed by the cell growth curve. shPLD1 delayed LNCaP-AI cell growth in the shPLD1 group in comparison with the shCtrl group.
Discussion
Although numerous studies in recent years have demonstrated that PLD is closely associated with cell proliferation and tumor formation, and PLD1 expression and activation were found to be up-regulated in multiple malignant tumors, the regulatory mechanism remains unclear [17-19]. In our prior work [20], we used natural ADPC cell line LNCaP as the control group and LNCaP-AI as the experiment group (induced by gradual tapering of the androgen concentration in the medium). When using UPLC-ESI-Q-TOF MS and GC-MS methods to detect the metabolic products after transition of LNCaP cells to AIPC, we accidentally found that PC was remarkably elevated. Knowing that PLD is an important enzyme participating in this metabolic process [10], we speculated that PLD might also participate in the process of AIPC transition.
In the present study, we used Arraystar Human LncRNA Microarray V3.0 to detect differential expression of PLD1 mRNA and its signaling-related mRNA before and after LNCaP cells underwent AIPC transition. The results of the present microarray and our previous UPLC-ESI-Q-TOF MS/GC-MS showed that the PC level was elevated dramatically after LNCaP cell transition to AIPC, though the exact mechanism remains unclear; the expression of PLD1 mRNA was up-regulated by 373 (373.44±75.80) fold; and the expression of the key molecule mRNA on its downstream Ras-PI3K-AKT and mTOR pathway was increased substantially (Fig 1), suggesting that in the androgen-free environment PCa cells could initiate some mechanism to increase the generation of the metabolic product PC, which in turn activated and increased the expression of its corresponding metabolic enzyme PLD1 mRNA, thus promoting PC transition to phosphatidic acid (PA). PA is a second messenger that plays a central role in the important cancer-associated signaling pathway network, which further activates the downstream Ras-PI3K-AKT and mTOR pathway as represented by the increased mRNA expression of the key enzymes such as PIK3R2, AKT3, C-raf and K-ras [21, 22]. It was reported in the literature that AKT exerted its anti-apoptosis effect by phosphorylating the target protein via multiple downstream pathways [23, 24], and on the other hand the activated AKT regulated the cell function, proliferation and differentiation by phosphorylating multiple enzymes, kinases, transcription factors and other downstream factors[25, 26]. Therefore, activation of this pathway could partially explain why PCa cells can continue to survive in the androgen-free environment.
To observe the effect of PLD1 mRNA expression on LNCaP cell proliferation after AIPC transition, we screened out two 19-21nt RNAi target sequences specific to PLD1 mRNA in our subsequent experiment, based on which we designed a shRNAi sequence to construct a GV115 vector (GFP labeling) and encapsulated it into lentiviral particles to interfere with the expression of PLD1 mRNA in LNCaP-AI cells. The results showed that the growth rate of LNCaP-AI cells that grew very fast in the androgen-free environment was decreased by 30% after PLD1 mRNAi (Fig. 3), indicating that PLD1 mRNA played a certain role in helping ADPC cells survive in the androgen-free environment. However, it is not the only factor affecting their survival, because interference only slowed down rather inhibited the rate of cell proliferation. In addition, the result of our microarrays showed that the expression of AR was up-regulated, suggesting PCa cells may continue to make use of the trace amount of androgen in the androgen-free environment to maintain their survival, or they may synthesize androgen intrinsically to maintain their viability [27-29].
In summary, our microarray and lentiviral interference have provided preliminary evidence that PLD1 mRNA and related PI3K/AKT and mTOR signaling pathway may play important roles in the process of AIPC transition, although it may not be the only pathway. Down-regulating the PLD1 mRNA expression could to some extent inhibit the proliferation rate of PCa cells after AIPC transition, which may provide a new way of thinking in clinical inhibition of PCa cell proliferation after AIPC transition.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (Grant No. 81202026); the Health and Family Planning Commission of Zhejiang Province (Grant No. 2015KYB243); and the Wenzhou Science & Technology Bureau (Grant No. Y20160505).
Abbreviation
PLD1: phospholipase D1; PCa: prostate cancer; ADPC: androgen dependence prostate cancer; AIPC: androgen independence prostate cancer.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Jemal A, Siegel R, Xu JQ. et al. Cancer Statistics, 2010. Ca-Cancer J Clin. 2010;60:277-300
2. Liu N, Liang WF, Ma XH. et al. Simultaneous and combined detection of multiple tumor biomarkers for prostate cancer in human serum by suspension array technology. Biosens Bioelectron. 2013;47:92-8
3. Humphreys MR, Fernandes KA, Sridhar SS. Impact of Age at Diagnosis on Outcomes in Men with Castrate-Resistant Prostate Cancer (CRPC). J Cancer. 2013;4:304-14
4. Azzouni F, Mohler J. Biology of castration-recurrent prostate cancer. The Urologic clinics of North America. 2012;39:435-52
5. Koivisto P, Visakorpi T, Kallioniemi OP. Androgen receptor gene amplification: a novel molecular mechanism for endocrine therapy resistance in human prostate cancer. Scandinavian journal of clinical and laboratory investigation Supplementum. 1996;226:57-63
6. Edwards J, Krishna NS, Mukherjee R. et al. Amplification of the androgen receptor may not explain the development of androgen-independent prostate cancer. BJU international. 2001;88:633-7
7. Li Y, Qu S, Li P. A novel mutation of the androgen receptor gene in familial complete androgen insensitivity syndrome. European review for medical and pharmacological sciences. 2015;19:4146-52
8. Ning Y, Zhang F, Zhu Y. et al. Novel androgen receptor gene mutation in patient with complete androgen insensitivity syndrome. Urology. 2012;80:216-8
9. Fiandalo MV, Wu W, Mohler JL. The role of intracrine androgen metabolism, androgen receptor and apoptosis in the survival and recurrence of prostate cancer during androgen deprivation therapy. Curr Drug Targets. 2013;14:420-40
10. Liu Y, Zhu X, Chen Y. et al. A preliminary study on metabonomics of androgen-dependent prostate cancer cell lines. J Anal Chem. 2011;39:305-11 (in Chinese)
11. Small EJ. Redefining Hormonal Therapy for Advanced Prostate Cancer: Results from the LATITUDE and STAMPEDE Studies. Cancer Cell. 2017;32:6-8
12. Noble AR, Maitland NJ, Berney DM. et al. Phospholipase D inhibitors reduce human prostate cancer cell proliferation and colony formation. Br J Cancer. 2018;118:189-99
13. Han JM, Kim Y, Lee JS. et al. Localization of phospholipase D1 to caveolin-enriched membrane via palmitoylation: implications for epidermal growth factor signaling. Mol Biol Cell. 2002;13:3976-88
14. Colley WC, Sung TC, Roll R. et al. Phospholipase D2, a distinct phospholipase D isoform with novel regulatory properties that provokes cytoskeletal reorganization. Curr Biol. 1997;7:191-201
15. Zhang Y, Frohman MA. Cellular and physiological roles for phospholipase D1 in cancer. J Biol Chem. 2014;289:22567-74
16. Kang DW, Choi CY, Cho YH. et al. Targeting phospholipase D1 attenuates intestinal tumorigenesis by controlling beta-catenin signaling in cancer-initiating cells. J Exp Med. 2015;212:1219-37
17. Bruntz RC, Lindsley CW, Brown HA. Phospholipase D signaling pathways and phosphatidic acid as therapeutic targets in cancer. Pharmacol Rev. 2014;66:1033-79
18. Dhingra S, Rodriguez ME, Shen Q. et al. Constitutive activation with overexpression of the mTORC2-phospholipase D1 pathway in uterine leiomyosarcoma and STUMP: morphoproteomic analysis with therapeutic implications. Int J Clin Exp Pathol. 2010;4:134-46
19. Gozgit JM, Pentecost BT, Marconi SA. et al. PLD1 is overexpressed in an ER-negative MCF-7 cell line variant and a subset of phospho-Akt-negative breast carcinomas. Br J Cancer. 2007;97:809-17
20. Xu G, Mao Y, Chen B. et al. Establishment of the androgen-independent prostate cancer cell model induced by Flutamide in combination with the androgen-free environment. J Cell Biol. 2009;31:373-8 (in Chinese)
21. Lim HK, Choi YA, Park W. et al. Phosphatidic acid regulates systemic inflammatory responses by modulating the Akt-mammalian target of rapamycin-p70 S6 kinase 1 pathway. J Biol Chem. 2003;278:45117-27
22. Rodriguez-Gonzalez A, Ramirez de Molina A, Banez-Coronel M. et al. Inhibition of choline kinase renders a highly selective cytotoxic effect in tumour cells through a mitochondrial independent mechanism. Int J Oncol. 2005;26:999-1008
23. Tan XL, Guo L, Wang GH. Polyporus umbellatus inhibited tumor cell proliferation and promoted tumor cell apoptosis by down-regulating AKT in breast cancer. Biomed Pharmacother. 2016;83:526-35
24. Hung CM, Lin YC, Liu LC. et al. CWF-145, a novel synthetic quinolone derivative exerts potent antimitotic activity against human prostate cancer: Rapamycin enhances antimitotic drug-induced apoptosis through the inhibition of Akt/mTOR pathway. Chem Biol Interact. 2016;260:1-12
25. Ke K, Li Q, Yang X. et al. Asperosaponin VI promotes bone marrow stromal cell osteogenic differentiation through the PI3K/AKT signaling pathway in an osteoporosis model. Sci Rep. 2016;6:35233
26. Reddi S, Kumar N, Vij R. et al. Akt drives buffalo casein-derived novel peptide-mediated osteoblast differentiation. J Nutr Biochem. 2016;38:134-44
27. Maity SN, Titus MA, Gyftaki R. et al. Targeting of CYP17A1 Lyase by VT-464 Inhibits Adrenal and Intratumoral Androgen Biosynthesis and Tumor Growth of Castration Resistant Prostate Cancer. Sci Rep. 2016;6:35354
28. Bremmer F, Jarry H, Strauss A. et al. Increased expression of CYP17A1 indicates an effective targeting of the androgen receptor axis in castration resistant prostate cancer (CRPC). Springerplus. 2014;3:574
29. Zhou W, Jiang Y, Ji LL. et al. Expression profiling of genes in androgen metabolism in androgen-independent prostate cancer cells under an androgen-deprived environment: mechanisms of castration resistance. International Journal Of Clinical And Experimental Pathology. 2016;9:8424-31
Author contact
Corresponding author: Meijuan Zhang, Department of Medical Laboratory, the First Affiliated Hospital of Wenzhou Medical University, Zhejiang Wenzhou, 325000, China, Email: zhangmeijuanedu.cn

 Global reach, higher impact
Global reach, higher impact