Impact Factor ISSN: 1837-9664
J Cancer 2018; 9(14):2415-2427. doi:10.7150/jca.24406 This issue Cite
Research Paper
ELTD1 Function in Hepatocellular Carcinoma is Carcinoma-Associated Fibroblast-Dependent
1. Department of Hepatobiliary Oncology, Sun Yat-sen University Cancer Center
2. Department of Experimental Research, Sun Yat-sen University Cancer Center
Received 2017-12-17; Accepted 2018-5-1; Published 2018-6-14
Abstract
Introduction: EGF, latrophilin, and seven transmembrane domain containing 1 (ELTD1) constitutes an orphan G-protein-coupled receptor (GPCR) of the adhesion family. High expression of ELTD1 is correlated with favorable prognosis of hepatocellular carcinoma (HCC). After silencing ELTD1 expression, however, tumor invasiveness is drastically reduced. The underlying mechanism of this apparent contradictory phenomenon is unknown. Because adhesion GPCRs couple extracellular adhesion to intracellular signaling, as a member of this family, ELTD1 function may be related to its tumor microenvironment. We therefore investigated the interaction between ELTD1 and the HCC tumor microenvironment.
Methods: ELTD1 expression was assessed by immunohistochemical analyses of tissue samples from two independent groups of 333 patients with HCC. Correlations between the ELTD1 expression and the clinicopathological values were examined. We also constructed ELTD1 overexpression and knockdown HCC cell lines and conducted a series of in vivo and in vitro ELTD1 functional assays. We further collected carcinoma associated fibroblast (CAF) culture supernatants to culture HCC cell lines and repeat the respective functional assays in comparison with the control group.
Results: Clinicopathologic correlations and in vivo models indicated ELTD1 as a tumor suppressor gene, whereas in vitro experiments suggested that ELTD1 could promote malignancy in HCC cell lines. Immunohistochemical staining of the generated ELTD1 overexpression xenograft tumors demonstrated that the CAF markers vimentin and α-SMA were highly expressed compared to the control group. This suggests that ELTD1 expression is correlated to CAF distribution. In addition, culturing with CAF supernatants inhibited HCC cell proliferation and invasion rates, confirming the correlation between CAF and ELTD1.
Conclusion: The results of this study indicated that ELTD1 regulation of HCC progression is CAF-dependent, suggesting that ELTD1 function is regulated by its tumor microenvironment. Further investigation is required to determine the underlying mechanisms.
Keywords: cancer associated fibroblast, ELTD1, hepatocellular carcinoma, therapeutics
Introduction
Hepatocellular carcinoma (HCC) is the most common neoplasm among all primary liver cancers and constitutes the second leading cause of cancer-related death [1]. Studies have shown that the crosstalk between cancer cells and their stromal microenvironment is crucial to the carcinogenesis and progression of HCC [2,3]. The modified stroma around the malignant cells is termed the tumor microenvironment; this is mainly composed of fibroblasts alongside with endothelial cells, pericytes, adipose cells, immune and inflammatory cells, and the extracellular matrix elements [4]. The interaction amongst these cells, which is mediated by G protein-coupled receptors (GPCRs), is essential to tumor microenvironment function. The GPCR superfamily comprises the largest family of integral transmembrane proteins in vertebrates and plays a dominant role in signal transduction [5]. Notably, over 100 GPCRs represent orphan receptors that do not have identified ligands, of which the largest family is the adhesion GPCRs [6]. Adhesion GPCRs feature unique subfamily-specific properties that couple (extracellular) cell adhesion to intracellular GPCR signaling. Specifically, their large extracellular N-terminal domains comprise multiple functional subdomains that facilitate interactions with extracellular proteins to facilitate cell-cell adhesion [7].
EGF, latrophilin and seven transmembrane domain containing 1 (ELTD1) is a relatively less studied orphan receptor of the adhesion GPCR family that was first reported to function as an important effector in the processes of angiogenesis [8]. High expression of ELTD1 has been shown to promote both normal and pathological angiogenesis [9]; however, the clinical relevance of ELTD1 expression in renal, head and neck, colorectal, and ovarian cancer demonstrates that higher expression of ELTD1 is correlated with favorable prognosis [10]. As considerable evidence supports the model that tumor proliferation, invasion, and metastasis depend on angiogenesis [11-13], the question of how ELTD1 can simultaneously promote tumor angiogenesis and be a favorable prognosis factor appears worthy further investigation. Considering that the adhesion GPCR family facilitates interactions with extracellular proteins, their functions are likely mediated by the tumor microenvironment. We therefore suspected that the apparent contradictory effect of ELTD1 may be caused by associated changes in the tumor microenvironment.
A growing body of evidence indicates that a sub-population of fibroblasts that comprise the tumor microenvironment can modulate cancer progression. The role of these cells, termed carcinoma-associated fibroblasts (CAFs) [14], in tumor progression is multifaceted. CAFs inhibit early stages of tumor progression, mainly through the formation of gap junctions between activated fibroblasts [15]. Subsequently, CAFs become activated by several tumor-secreted factors and promote both tumor growth and progression [16]. Notably, the initial gap junctions are most likely formed by adhesion GPCRs [17]. Thus, in the present study, we analyzed the frequent aberrant expression of ELTD1 in HCC tissues by immunohistochemical (IHC) staining, and examined possible correlations between ELTD1 and several clinicopathological variables. We then conducted a series of in vivo and in vitro experiments to verify the correlation between ELTD1 and the tumor microenvironment in HCC.
Materials and Methods
Patients and tissue specimens
This study was approved by the institutional review board of Sun Yat-sen University Cancer (SYSU). Written informed consent was obtained from all patients. Formalin-fixed, paraffin-embedded tissues were obtained from 333 patients who underwent curative resection for HCC at the Cancer Center of Sun Yat-sen University between 2004 and 2011. The inclusion criteria used for patient enrollment in the cohorts were an absence of anticancer therapies or distant metastasis before the operation; a lack of concurrent autoimmune disease, human immunodeficiency virus, or syphilis; and the availability of follow-up data. Patients with Child-Pugh class B or C were excluded from our study. Histological diagnosis was based on the World Health Organization Classification of Hepatocellular Carcinoma [18]. Tumor differentiation was defined in accordance with the criteria of Edmondson and Steiner [19]. Pathological staging was performed by the Department of Pathology, Sun Yat-sen University Cancer Center according to the Barcelona Clinic Liver Cancer (BCLC) staging classification [20]. Clinicopathological features were obtained from patient clinical records (Table 1).
Correlation between ELTD1 expression and clinicopathological characteristics in patients with hepatocellular carcinoma
| Characteristic | No. of patients | ELTD1 expression (%) | P-value | |
|---|---|---|---|---|
| high | low | |||
| Gender | 0.847 | |||
| Female | 37 | 18 (48.6) | 19 (51.4) | |
| Male | 296 | 139 (47.0) | 157 (53.0) | |
| Age (years) | 0.682 | |||
| ≤ 50 | 180 | 83 (46.1) | 97 (53.9) | |
| > 50 | 153 | 74 (48.4) | 79 (51.6) | |
| HBsAg | 0.838 | |||
| Negative | 33 | 15 (45.5) | 18 (54.5) | |
| Positive | 300 | 142 (47.3) | 158 (52.7) | |
| AFP (ng/mL) | 0.078 | |||
| ≤ 200 | 176 | 91(51.7) | 85 (48.3) | |
| > 200 | 157 | 66 (42.0) | 91 (58.0) | |
| GGT (U/L) | 0.099 | |||
| ≤ 100 | 256 | 127 (49.6) | 129 (50.4) | |
| > 100 | 77 | 30 (39.0) | 47 (61.0) | |
| Liver cirrhosis | 0.148 | |||
| No | 192 | 84 (43.8) | 108 (56.2) | |
| Yes | 141 | 73 (51.8) | 68 (48.2) | |
| Tumor diameter (cm) | 0.000 | |||
| ≤ 5 | 173 | 105 (60.7) | 68 (39.3) | |
| > 5 | 160 | 52 (32.5) | 108 (67.5) | |
| Tumor number | 0.000 | |||
| Single | 278 | 143 (51.4) | 135 (48.6) | |
| Multiple | 55 | 14 (25.5) | 41 (74.5) | |
| Tumor capsule | 0.235 | |||
| None/incomplete | 225 | 101 (44.9) | 124 (55.1) | |
| Complete | 108 | 56 (51.9) | 52 (48.1) | |
| Tumor differentiation | 0.676 | |||
| I-II | 221 | 106 (48.0) | 115 (52.0) | |
| III-IV | 112 | 51 (45.5) | 61 (54.5) | |
| Vascular invasion | 0.004 | |||
| No | 289 | 145 (50.2) | 144 (49.8) | |
| Yes | 44 | 12 (27.3) | 32 (72.7) | |
| TNM stage | 0.000 | |||
| I | 242 | 132 (54.5) | 110 (45.5) | |
| II | 30 | 9 (30.0) | 21 (70.0) | |
| III | 61 | 16 (26.2) | 45 (73.8) | |
| BCLC stage | 0.000 | |||
| 0 | 43 | 25 (58.1) | 18 (41.9) | |
| A | 115 | 75 (65.2) | 40 (34.8) | |
| B | 131 | 45 (34.4) | 86 (65.6) | |
| C | 44 | 12 (27.3) | 32 (72.7) | |
| Early recurrence | 0.000 | |||
| No | 196 | 111 (56.6) | 85 (43.4) | |
| Yes | 137 | 46 (33.6) | 91 (66.4) | |
| MVD | 0.000 | |||
| Low | 164 | 59 (36.0) | 105 (64.0) | |
| High | 169 | 98 (58.0) | 71 (42.0) | |
AFP: alpha-fetoprotein; GGT: gamma-glutamyl transferase; TNM: tumor, lymph node, metastasis; BCLC: Barcelona-Clinic Liver Cancer; MVD: microvessel density.
Patient follow-up
HCC recurrence within the first two years following surgery is mainly due to intrahepatic metastasis, whereas late recurrence usually results from multicentric occurrence [21]. Using two years as a cutoff, tumor recurrence was classified as either early recurrence or late recurrence [22]. During follow-up, serum alpha-fetoprotein (AFP) levels were monitored and three phase dynamic computed tomography scanning or magnetic resonance imaging was performed every one to three months after surgery. Overall survival (OS) was defined as from the date of liver resection to the date of death or last follow-up. Recurrence-free survival (RFS) was measured from the date of surgery until the date of relapse, metastasis, or last follow-up. Curative resection was defined as complete resection of all tumor nodules with clear microscopic resection margins and no residual tumors on computed tomography scans performed one month after surgery.
Isolation and culture of CAFs and preparation of CAF supernatants (CSNs)
The generation of CAFs was performed as previously described [3]. Paired fresh HCC and distant liver tissues were obtained from two patients with HCC undergoing partial hepatic resection. Tissues were finely minced with a scalpel in a tissue culture dish and enzymatically dissociated in 1 mg/mL collagenase type IV (Sigma-Aldrich, St Louis, MO, USA) at 37°C for 1 h with frequent shaking. The activity of collagenase was blocked using phosphate buffered saline (PBS) supplemented with 10% fetal bovine serum. The single cell suspension was washed and filtered through a 40-μm cell strainer (BD Biosciences, Milan, Italy). Cells were plated on a 100-mm dish in MyeloCult® medium (StemCell Technologies, Vancouver, BC, Canada) in the presence of 1 μM hydrocortisone sodium succinate and 1% antibiotics. Cells were maintained in a controlled CO2 incubator at 37°C, 95% humidity, and 5% CO2 with medium change every three days, and sub-cultured with 0.05% trypsin in PBS when they reached 80%-90% confluence. Morphological homogeneity of the cells was observed along subcultures. The primary cells from HCC nodules were identified as CAFs whereas cells from distant non-tumoral tissues were classified as non-tumoral fibroblasts.
CSNs were prepared as previously described [23] by plating 5 × 106 tumor cells in 10 mL complete medium in 100-mm dishes for 24 h, after which the medium was changed to complete Dulbecco's modified Eagle medium (DMEM; Gibco, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum. After 48 h, the supernatants were harvested, centrifuged, and stored in aliquots at -80°C.
Cell lines and culture conditions
Human HCC cell lines SMMC-7721 and Huh7, and one normal hepatic cell line, L02, were obtained from the National Cancer Centre Singapore (NCCS). All cell lines were cultured in DMEM supplemented with 10% fetal bovine serum (Gibco) or CSN. The cells were incubated in a humidified incubator supplied with 5% CO2 at 37°C.
Plasmid constructs and transfection
Human ELTD1 cDNA (GeneCopoeia, Inc., Rockville, MD, USA) was cloned into the pEZ-Lv201 vector (GeneCopoeia, Inc., Rockville, MD, USA) and then transfected into the 293FT packaging cell line using a Lenti-Pac™ HIV Expression Packaging Kit (GeneCopoeia, Inc.) according to the manufacturer's instructions. Virus-containing supernatants from 293FT cells were collected and filtered using 0.45-μm filters. The filtered supernatant was added to 70% confluent cells in the presence of 8 μg/mL polybrene (Sigma). After 48 h, the cells were incubated with fresh complete medium containing the appropriate concentration of puromycin for stable transduced cells. In addition, the control vector pEZ-Lv201 alone was also transfected in the same manner.
Short interfering RNA (siRNA) transfection
SiRNA targeting ELTD1 was purchased from Ribo Bio tech (Guangzhou, Guangdong, China) and transfected into HCC cells using Lipofectamine™ 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions.
IHC analysis and assessment of IHC staining
Paraffin-embedded samples were cut into 4-μm sections and processed for IHC. Tissue sections prepared for antigen retrieval by microwave treatment in citrate buffer (pH 6.0) were incubated with anti-ELTD1 (Proteintech Group, Chicago, IL, USA), anti-αSMA (Cell Signaling Technology, Danvers, MA, USA), anti-vimentin (Cell Signaling Technology), and anti-CD34 (Santa Cruz, Dallas, Texas, USA) primary antibodies. Immunostaining was performed using the Envision System with diaminobenzidine (Dako Cytomation, Glostrup, Denmark). Images were viewed and assessed using a microscope (Eclipse 80i, Nikon, Tokyo, Japan). Assessments of the ELTD1 staining were scored by two experienced pathologists blinded to patient identity and clinical status. The staining analysis technique was performed as previously described [24]. In particular, the proportion score was assigned based on the proportion of positive cells and the intensity of staining. The proportion of positive cells was scored from 0% to 100%. The intensity of staining was scored from 0 to 3 (0, none; 1, weak; 2, intermediate; 3, strong). The final quantitation of each staining was obtained by multiplying the two scores. For patient survival analyses, the median was set as the cut-off to distinguish strong and weak staining of ELTD1.
Evaluation of microvessel density (MVD)
As ELTD1 has been classified as an endothelial marker in both normal and pathological angiogenesis, we investigated the correlation between ELTD1 and MVD using CD34 staining for MVD evaluation of HCC as previously described [11]. Screening of the tumor was first performed at a low power (40×) to identify areas of MVD coordinating with the highest expression of ELTD1. Five areas of highest MVD were selected for counting under 200× magnification. Any brown-stained endothelial cell or endothelial cell cluster that was clearly separated from adjacent microvessels, tumor cells, and other connective tissues was considered a single, countable blood vessel regardless of whether a vessel lumen was seen. Image analysis for MVD counting was performed by using interactive software developed in our laboratory. The software allows the application of standardized computational algorithms as well as review and refinement of the result of those algorithms by human operators. The mean value of the vessel counts in the selected spots was retained as the final MVD count. For patient survival analyses, the median was set as the cut-off to distinguish high and low values of MVD.
Protein extraction and western blot analysis
Cells were lysed in the presence of 50 mM Tris (pH 7.5), 150 mM NaCl, and 0.5% NP-40 on ice. Protein lysates were separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and then transferred to a polyvinylidene fluoride membrane. After the membranes were blocked by 5% bolvine serum albumin, they were incubated with various antibodies at 4°C overnight. Then, the membranes were incubated with horseradish peroxidase-conjugated antibodies at room temperature for 45 min. Protein signals were detected using enhanced chemiluminescence (Pierce, Rockford, IL, USA).
RNA extraction and quantitative real-time polymerase chain reaction (PCR)
Total RNA was isolated from cell lines using TRIzol Reagent (Invitrogen Life Technologies). RNA (2 µg) was reverse-transcribed using a SuperScript® III First-Strand Synthesis System (Invitrogen Life Technologies) according to the manufacturer's instructions. For the real-time PCR assay, cDNA was subjected to PCR amplification using SYBR Green (Toyobo, Kita-ku, Osaka, Japan) and a Roche LightCycler 480 System (Madison, WI, USA). GAPDH was used as an internal control. The primers were as follows: GAPDH: Forward: GGT ATG ACA ACG AAT TTG GC, Reverse: GAG CAC AGG GTA CTT TAT TG; ELTD1: Forward: GAA ATT TAA CTC AGT CCT GTG G, Reverse: GTC CCA AAC TAC AAA TGT ATC C.
Cell colony formation assay
For colony formation assays, 1 × 103 cells were seeded in each well of 6-well plates and then cultured with DMEM supplemented with 10% fetal bovine serum for 10 days. The colonies were washed twice with PBS, fixed in methanol for 15 min, and stained with crystal violet for 15 min at room temperature. After the stain was washed out, the number of colonies was counted. Colony formation efficiency was measured as the ratio of the number of colonies formed to the total number of cells plated.
Cell invasion assay
Cell invasion assays were performed in Transwell chambers (8-μm pore size; Corning, Armonk, NY, USA). A total of 5 × 105 cells were seeded in the upper chamber of the insert with 1:8 diluted Matrigel (BD Biosciences, San Jose, CA, USA). The upper chamber was supplemented with serum-free medium, and medium with 5% fetal bovine serum was added to the lower chamber as a chemoattractant. These tumor cells were cultured at 37°C for 24 h. Five random fields were counted in triplicate assays.
In vivo tumorigenesis and invasion assays
All the animal experiments were performed in accordance with the guidelines of the Laboratory Animal Ethics Committee of Sun Yat-Sen University. For the in vivo tumorigenesis model, 5 × 106 cells were injected subcutaneously into BALB/C-nu/nu athymic nude mice. After four weeks, the mice were euthanized and the subcutaneous tumors were resected, fixed in phosphate-buffered neutral formalin, sectioned serially, and stained with hematoxylin and eosin. Then, IHC analysis was performed. Another orthotopic tumor model for HCC was established by intrahepatic tumor cell injection. Each group consisted of 6 male BALB/C-nu/nu athymic nude mice. Briefly, the mice were anesthetized with 1% pentobarbital and a midline incision was made to expose the liver. 7721-ELTD1 or 7721-Vector cells resuspended in Matrigel (1:1) were slowly injected under the hepatic capsule into the liver. After eight weeks, the livers and lungs of the two groups were collected after cervical dislocation to euthanise the mice.
Statistical analysis
All statistical calculations were performed using SPSS 19.0 statistical software (SPSS Inc., Chicago, IL, USA). For continuous variables, the data are expressed as the means ± standard error of the mean. The significance of differences between values was determined using the Student t test. The chi-squared test was applied to examine the correlation between ELTD1 expression and clinical pathological parameters. Survival curves for patients were calculated using the Kaplan-Meier method and analyzed using the log-rank test. Prognostic factors were examined by univariate and multivariate analyses using the Cox proportional hazards model. All differences were deemed significant at P < 0.05.
Results
ELTD1 staining by IHC in HCC tissues is inversely correlated with poorer clinicopathologic features
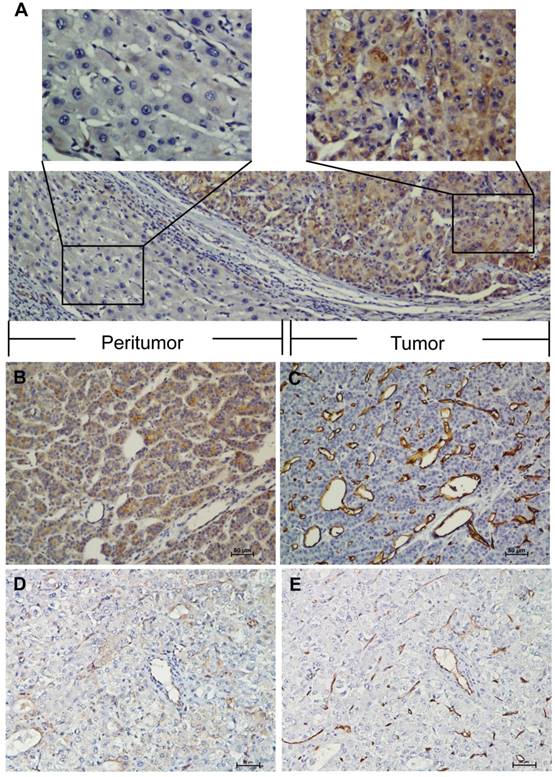
IHC analysis indicated that ELTD1 was primarily localized in the cytoplasm of the tumor cells (Figure 1A). The median of 0.5 was set as a cut-off. ELTD1 expression was classified as strong staining if the score was higher than the cut-off (Figure 1B), and weak if the score was 0.5 or less (Figure 1D). A total of 157 (47.1%) primary HCC specimens were detected as strong- and 176 (52.9%) as weak-staining specimens. The correlation between ELTD1 expression and clinicopathological characteristics is shown in Table 1. ELTD1 expression in cancer cells was correlated with tumor diameter (P = 0.000), tumor number (P = 0.000), vascular invasion (P = 0.000), TNM stage (P = 0.000), BCLC stage (P = 0.000), early recurrence (P = 0.000), and MVD (P = 0.000) (Table 1). Weak intensity of staining in the cancer cells tended to be associated with poorer characteristics than the other staining group.
IHC characteristics of ELTD1 in HCC specimens. ELTD1 was mainly expressed in the tumor cell cytoplasm. A small portion of endothelial cells of blood vessels were positive for ELTD1 expression in both tumor tissues. (A) Representative images of positive ELTD1 expression in the tumor cytoplasm (×200). The upper panel shows an enlargement of the indicated area (×400). (B) Representative staining of positive ELTD1 expression in the tumor cell cytoplasm (×200). (C) Staining of strong CD34 expression in the endothelial cells representing high MVD (×200). (D) Representative staining of negative ELTD1 expression in the tumor cell cytoplasm (×200). (E) Staining of weak CD34 expression in the endothelial cells representing low MVD (×200).
ELTD1 expression correlates with HCC angiogenesis as determined by MVD
MVD was categorized by counting the amount of vessels stained with CD34 by IHC. The mean value of MVD ranged from 5 to 400 microvessels/mm2; the median 88 microvessels/mm2 was set as the cut-off. MVD was classified as low density if the count was lower than the median; if it received a higher count, the case was classified as high density. A total of 169 (50.8%) primary HCC specimens were detected as high density compared to 164 (49.2%) low density specimens (Figure 1B, D). MVD classification was used to investigate whether ELTD1 expression is associated with angiogenesis in HCC (Figure 1C, E). Low MVD was significantly correlated with low ELTD1 expression (Table 1, P = 0.000).
ELTD1 expression correlates with enhanced survival in patients with HCC
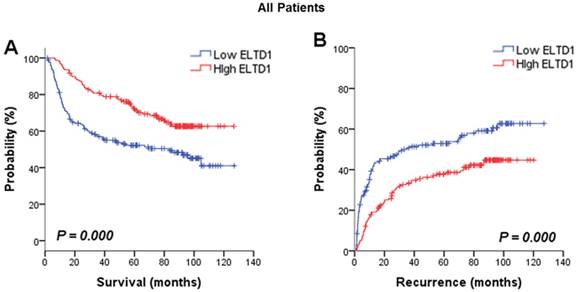
The association between ELTD1 expression in cancer cells and survival outcome was evaluated. Analysis of the prognostic significance of ELTD1 expression calculated by Kaplan-Meier survival analysis revealed that lower expression of ELTD1 was associated with a poorer survival rate (Figure 2A) and a higher possibility of recurrence (Figure 2B). In univariate analysis of OS and RFS in all patients, ELTD1 expression was significantly correlated with both OS and RFS (Table 2). The median OS in ELTD1 low-expression and ELTD1 high-expression subgroups was 39.0 and 74.9 months, respectively, whereas the median RFS was 22.2 and 57.2 months. These data indicated that the ELTD1 expression level in tumor cells represented an independent prognostic factor for HCC.
Positive ELTD1 expression in tumor cells is related to better prognoses of patients with HCC. Based on ELTD1 expression in the tumor cell cytoplasm analyzed by IHC, the patients were divided into two groups, ELTD1-low (negative expression) and ELTD1-high (positive expression). (A) Kaplan-Meier analysis for OS is displayed. (B) Kaplan-Meier analysis for RFS is displayed.
Univariate and multivariate analysis of ELTD1 association with survival and recurrence in patients with hepatocellular carcinoma
| Variable | OS | RFS | ||||||
|---|---|---|---|---|---|---|---|---|
| Univariate | Multivariate | Univariate | Multivariate | |||||
| P-value | P-value | HR | 95% CI | P-value | P-value | HR | 95% CI | |
| Gender (Female vs. Male) | 0.758 | 0.130 | ||||||
| Age, years (≤ 50 vs. > 50) | 0.631 | 0.721 | ||||||
| AFP (ng/mL) (≤ 200 vs. > 200) | 0.005 | 0.133 | 1.316 | 0.920-1.884 | 0.010 | 0.080 | 1.343 | 0.965-1.869 |
| HBsAg (Negative vs. Positive) | 0.558 | 0.277 | ||||||
| GGT (U/L) (≤ 100 vs. > 100) | 0.022 | 0.991 | 0.952 | 0.639-1.417 | 0.207 | |||
| Liver cirrhosis (No vs. Yes) | 0.001 | 0.000 | 1.900 | 1.353-2.669 | 0.126 | |||
| Tumor diameter (cm) (≤ 5 vs. > 5) | 0.000 | 0.078 | 1.501 | 0.955-2.360 | 0.000 | 0.053 | 1.540 | 1.007-2.355 |
| Tumor number (Single vs. Multiple) | 0.000 | 0.000 | 3.115 | 2.076-4.672 | 0.000 | 0.000 | 2.651 | 1.795-3.915 |
| Tumor capsule (No/ incomplete vs. Complete) | 0.001 | 0.058 | 0.819 | 0.666-1.007 | 0.147 | |||
| Tumor differentiation (I-II vs. III-IV) | 0.014 | 0.400 | 0.849 | 0.580-1.243 | 0.031 | 0.733 | 0.940 | 0.657-1.344 |
| Vascular invasion (No vs. Yes) | 0.000 | 0.000 | 2.938 | 1.893-4.561 | 0.000 | 0.000 | 2.184 | 1.425-3.347 |
| ELTD1 (Low versus High) | 0.000 | 0.009 | 0.623 | 0.437-0.888 | 0.000 | 0.046 | 0.723 | 0.521-1.004 |
| MVD (Low versus High) | 0.000 | 0.117 | 0.706 | 0.457-1.091 | 0.001 | 0.528 | 0.874 | 0.575-1.328 |
OS: overall survival; AFP: alpha-fetoprotein; GGT: gamma-glutamyl transferase; TNM: tumor, lymph node, metastasis; BCLC: Barcelona-Clinic Liver Cancer; MVD: microvessel density.
Lower ELTD1 expression generally correlates with worse prognosis of patients with HCC according to clinical (BCLC Stage) and pathological subgroup
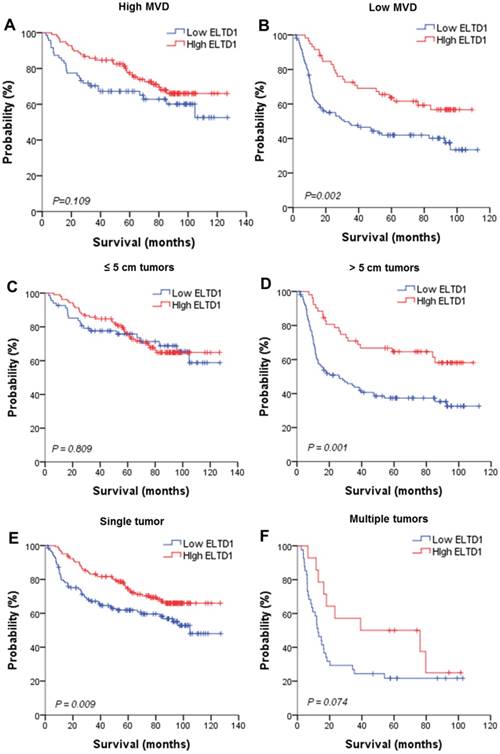
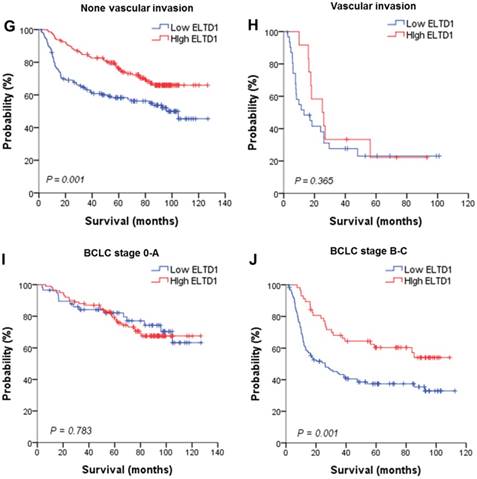
To further evaluate the prognostic value of ELTD1 in different subgroups, patients were stratified according to MVD staining (Figure 3A, B), tumor size (Figure 3C, D), tumor number (Figure 3E, F), vascular invasion (Figure 3G, H), and BCLC stage (Figure 3I, J). Kaplan-Meier survival analysis according to ELTD1 expression suggested ELTD1 to be of prognostic value particularly in patients with low MVD (Figure 3 A, B). The prognostic value of lower ELTD1 expression was maintained in predicting poorer OS and RFS in most of the other subgroups as well.
Effect of ELTD1 expression in tumor cells on the prognoses of patients stratified into subgroups. (A, B) Kaplan-Meier survival analysis of OS in patients exhibiting high (A) or low (B) MVD. The OS in the ELTD1-low group was significantly decreased compared with that in the ELTD1-high group (P = 0.002). (C) Patients with tumor diameter ≤ 5cm. (D) Patients with tumor diameter > 5cm. The OS in the ELTD1-low group was significantly decreased compared with that in the ELTD1-high group (P = 0.001). (E) Patients with a single tumor. The OS in the ELTD1-low group was significantly decreased compared with that in the ELTD1-high group (P = 0.009). (F) Patients with multiple tumors. (G) Patients without vascular invasion. The OS in ELTD1-low group was significantly decreased compared with that in the ELTD1-high group (P = 0.001). (H) Patients with vascular invasion. (I) BCLC stage 0-A patients. (J) BCLC stage B-C patients. The OS in the ELTD1-low group was significantly decreased compared with that in the ELTD1-high group (P = 0.001).
ELTD1 displays an oncogenic function in HCC in vitro
Because ELTD1 expression in tumor cells was inversely associated with malignant clinicopathological features, we explored the potential biological functions of ELTD1 in HCC tumorigenesis. First, we examined the ELTD1 expression pattern in HCC cell lines (SMMC-7721 and Huh7) and normal liver cells (L02). Notably, SMMC-7721 displayed lower ELTD1 messenger RNA and protein levels whereas Huh7 displayed significantly higher levels of ELTD1 than the normal liver cell line (Figure 4A, B).
ELTD1 expression in a normal liver cell line and HCC cell lines. (A) Western blotting and qPCR results show that SMMC-7721 cells exhibit low expression and Huh7 exhibits high ELTD1 expression compared to the normal liver cell line L02. (B) ELTD1-overexpressing SMMC-7721 cell line verified by western blotting and qPCR. (C) ELTD1-knockdown Huh7cell line verified by western blotting and qPCR.
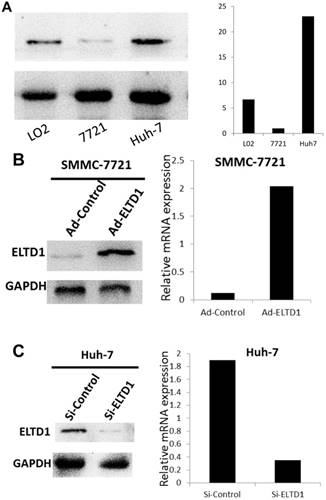
To further investigate the role of ELTD1 in malignancy, SMMC-7721 was transfected with an ELTD1 expression plasmid (pEZ-Lv201-ELTD1) and a control vector (pEZ-Lv201), whereas Huh7 cells were transfected with an ELTD1 knockdown siRNA and a control vector. The ectopic expression of ELTD1 in the cells was confirmed by quantitative real-time PCR and western blot analysis (Figure 4C, D). Functional assays (Figure 5, left) showed that overexpression of ELTD1 in HCC cells significantly promoted the cell invasion rate (P ˂ 0.05; Figure 5A) and growth (P ˂ 0.05; Figure 5C). Knockdown of ELTD1, on the other hand, inhibited cell invasion rate (P ˂ 0.05; Figure 5B) and growth (P ˂ 0.05; Figure 5D).
ELTD1 increases cell growth and migration directly but decreases cell growth and migration after culture with CAF supernatants in vitro. (A) Matrigel invasion assay showing the effect of ELTD1 overexpression (Ad-ELTD1 vs Ad-Control) on cell migration and invasion in SMMC-7721 cells. Representative images of invaded cells are shown in the left panels, and the results are summarized in the right panels. The results are expressed as the means of three independent experiments. (B) Matrigel invasion assay showing the effect of ELTD1 knockdown on cell migration and invasion in Huh7 cells (Si-ELTD1 VS Si-Control). (C) Cell colony formation assay showing the effect of ELTD1 overexpression on cell growth in SMMC-7721 cells. (D) Cell colony formation assay showing the effect of ELTD1 knockdown on cell growth in Huh7 cells.
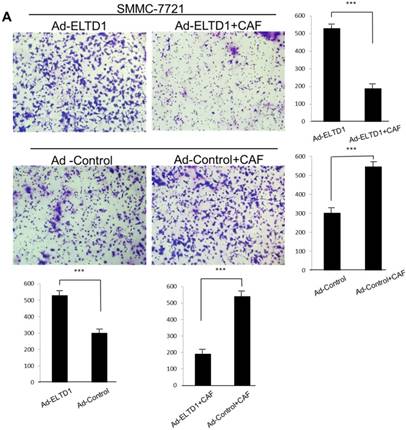
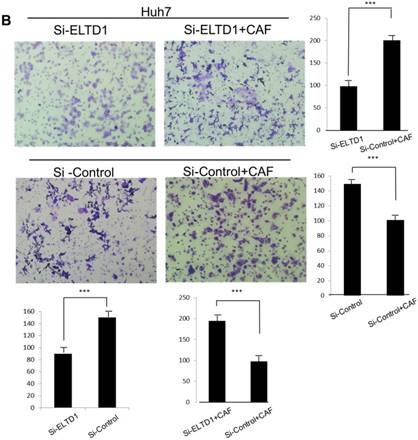
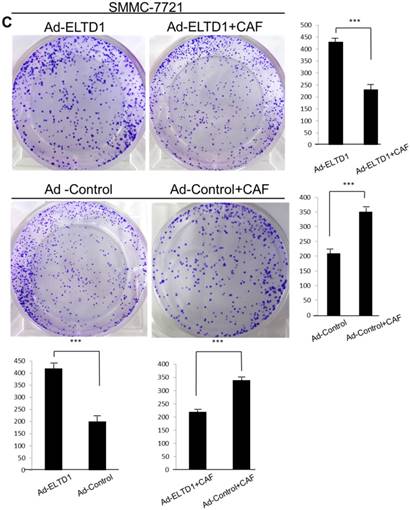
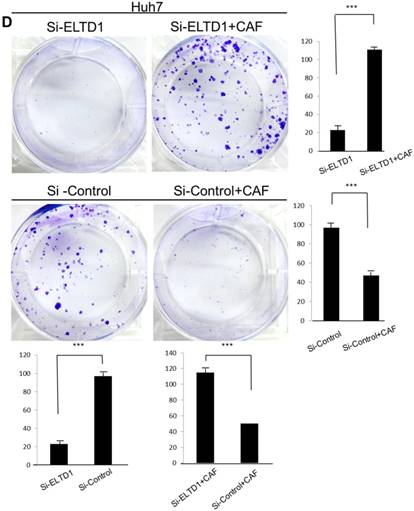
ELTD1 overexpression decreases whereas ELTD1 knockdown increases cell proliferation and tumor invasion after culture with CSN
To assess the influence of the tumor microenvironment on ELTD1 function, we next cultured ELTD1 overexpression SMMC-7721 cells and ELTD1 knockdown Huh7 cells with CSNs and repeated the functional assays (Figure 5, right). Notably, the cell colony and invasion were inhibited upon CSN culture in the ELTD1-overexpressimg HCC cells but were promoted in the ELTD1 knockdown cells (Figure 5).
ELTD1 expression inhibits tumorigenesis in in vivo mouse models
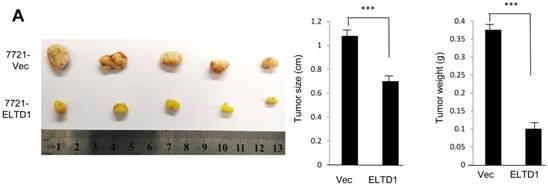
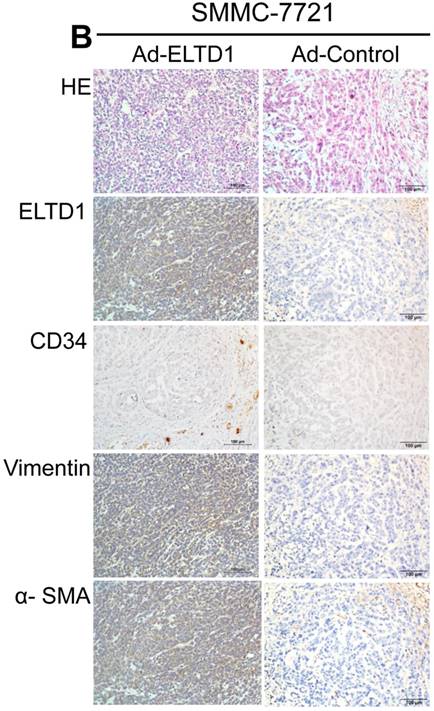
To demonstrate ELTD1 function in vivo, 7721-ELTD1 and 7721-Vector cells were subcutaneously injected into nude mice. After 4 weeks, the mice were sacrificed and the xenograft tumors were harvested and measured. The results showed that the xenograft tumors of the ELTD1 overexpression group were significantly smaller and lighter (P < 0.05) than those of the control group (Figure 6A). The corresponding images for the hematoxylin and eosin and IHC of ELTD1, CD34, vimentin, and α-SMA staining are shown in Figure 6B In addition, two groups of 6 mice each were used to confirm the effect of ELTD1 overexpression on hepatic tumor formation and metastasis by the orthotopic transplantation of 7721- ELTD1 or 7721-Vector cells into BALB/c mice. The liver tumors from the group of 7721- ELTD1 mice were significantly smaller than those from the 7721-Vector group (Figure 6C). This series of experiments suggested that ELTD1 affected HCC cells differently in vivo than in vitro.
An experimental mouse model was used to evaluate the effect of ELTD1 overexpresison on tumor growth by subcutaneous injection of 7721-ELTD1 or 7721-Vector cells. The tumor diameter and weight are shown in the right panel. (B) Representative hematoxylin and eosin (HE), ELTD1, CD34, vimentin and α-SMA staining of tumor nodules are shown. (C) Another HCC model was created by direct intrahepatic injection of 7721-ELTD1 or 7721-Vector cells suspended in Matrigel. Representative orthotopic liver tumors and metastatic nodules are shown in the left panel. The right panel is representative of the intra- and extrahepatic metastasis of the HCC model.

Discussion
ELTD1 expression was first identified in cardiomyocytes and vascular smooth muscle cells. ELTD1 has been suggested to assist in the development of the heart and the maturation of vascular smooth muscle [8]. In addition, ELTD1 has been implicated in the angiogenesis of several types of cancer including gliomas [25], renal cancer [26], head and neck, colorectal, and ovarian cancer [10]. Although angiogenesis is believed to enhance tumor proliferation, invasion, and metastasis [11-13], previous studies demonstrated that high expression of ELTD1, conversely, was correlated with favorable prognosis [10]. This apparent contradictory effect may be due to the unique characteristic of ELTD1 as an adhesion GPCR, containing a novel long N-terminus including multiple domains that facilitate extracellular protein interactions [8,17]. Therefore, adhesion GPCRs play a dominant role in transmembrane signal transduction [5], especially in the tumor microenvironment, which is primarily composed of CAFs along with several other cell types [4]. The role of CAFs in tumor progression is based on their crosstalk with cancer cells. In particular, CAFs affect cancer cell responses; in turn, however, cancer cells trigger a reactive response in the tumor microenvironment [16]. We suspect that the interaction between CAFs and cancer cells might adjust the expression of ELTD1, causing apparent contradictory effects on cancer cells; therefore, in the present study, we explored the function of ELTD1 in HCC.
In our study, we investigated the interaction between CAFs and ELTD1 in HCC. Similar to other cancer types, the correlation between ELTD1 expression and MVD was significant, suggesting that ELTD1 facilitates HCC angiogenesis. However, the IHC results showed that the main distribution of ELTD1 was in the cytoplasm of HCC cells. To investigate this issue further, we next analyzed the correlation between ELTD1 expression and clinicopathological characteristics. The results demonstrated that high expression of ELTD1 was an independent predictor for better OS and RFS in HCC. After stratification into subgroups, ELTD1 maintained its prognostic value in single tumors, tumors larger than 5 cm, and tumors with no vascular invasion. These results suggested that ELTD1 expression correlates with favorable prognosis in patients with HCC, as had been reported in previous studies [10].
To further explore the biological functions of ELTD1 and clarify its dependence on CAFs, we isolated CAFs from HCC and harvested their supernatants for use in HCC cells line culture. In contrast, the control group was only cultured with 10% fetal bovine serum. In the control group, functional assays showed that overexpression of ELTD1 in HCC cells significantly promoted cell growth and invasion rate. Knockdown of ELTD1, however, inhibited cell growth and invasion rate. Conversely, after being cultured with CSN, overexpression of ELTD1 inhibited cell growth and invasion rate whereas knockdown of ELTD1 promoted HCC malignancy. In order to further examine the interaction between ELTD1 and CAF, xenograft tumors were constructed. We found that the xenografts of the ELTD1 overexpression group were significantly smaller and lighter than those of the ELTD1 knockdown group. The IHC results of the xenograft tumors showed that in the ELTD1 overexpression group, CAF-related markers vimentin and α-SMA were highly expressed, suggesting that the expression of ELTD1 is correlated to CAF distribution.
Previous studies reported that in HCC, CAF is essential for tumor growth and metastasis [27]. HCC cells can further stimulate the proliferation of CAFs [28]. However, different HCC cell lines responded differently after co-culture with CAFs [29]. This may be the result of different TGF-β expression. In breast cancer, CAFs that overexpress TGF-β promote the tumorigenic outgrowth of normal breast epithelium [30]. Moreover, it has been reported that the loss of TGF-β signaling in CAFs contribute in the development of prostate cancer [31, 32]. These observations support the importance of TGF-β in CAF functions. In the present study, we investigated the possibility of ELTD1 as a potential regulatory gene in the progression of HCC. Furthermore, ELTD1 function was regulated by the secreted factors of CAFs. The possibility therefore exists that ELTD1 may mediate the TGF-β signaling of CAFs.
In summary, our study showed that ELTD1 expression is increased in the cytoplasm of HCC cells compared to that of paratumor tissues. High expression of ELTD1 is correlated with favorable outcome in patients with HCC. Native in vitro studies showed that ELTD1 acted as a tumor promoting gene, whereas in vivo studies suggested otherwise. However, after culturing HCC cells with CSN, ELTD1 inhibited the proliferation and metastasis of HCC, demonstrating that the apparent contradictory effects of ELTD1 are regulated by CAFs and suggesting the likely importance of the tumor microenvironment in mediating the positive impact of ELTD1 expression on patient outcome in HCC. Moreover, as previous reports have suggested that TGF-β recruits CAFs and invokes the expression of ELTD1, our data raise the possibility that TGF-β signaling regulates the interaction between CAFs and ELTD1. However, this notion needs further investigation.
Acknowledgements
The authors would like to thank Prof. Dr. J.P. Yun and Dr. M.Y. Cai, Department of Pathology, Sun Yat-sen University Cancer Center, for providing and dissecting the tissue samples. We are also grateful to and thank all the patients who participated in this study.
Abbreviations
AFP: alpha-fetoprotein;
BCLC: Barcelona Clinic Liver Cancer;
CAF: carcinoma-associated fibroblasts;
CSN: carcinoma-associated fibroblast supernatant;
DMEM: Dulbecco's modified Eagle medium;
ELTD1: EGF, latrophilin and seven transmembrane domain containing 1;
GGT: gamma-glutamyl transferase;
GPCR: G-protein-coupled receptor;
HCC: hepatocellular carcinoma;
HE: hematoxylin and eosin;
IHC: immunohistochemical;
MVD: microvessel density;
OS: overall survival;
PBS: phosphate buffered saline;
PCR: polymerase chain reaction;
RFS: Recurrence-free survival;
siRNA: short interfering RNA;
TNM: tumor, lymph node, metastasis.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Llovet JM, Zucman-Rossi J, Pikarsky E. et al. Hepatocellular carcinoma. Nat Rev Dis Primers. 2016;2:16018
2. Li HC, Fan XL, Houghton JM. Tumor microenvironment: the role of the tumor stroma in cancer. J Cell Biochem. 2007;101:805-15
3. Sukowati CH, Anfuso B, Crocé LS. et al. The role of multipotent cancer associated fibroblasts in hepatocarcinogenesis. BMC Cancer. 2015;15:188
4. Liu J, Chen S, Wang W. et al. Cancer-associated fibroblasts promote hepatocellular carcinoma metastasis through chemokine-activated hedgehog and TGF-beta pathways. Cancer Lett. 2016;379:49-59
5. Krishnan A, Dnyansagar R, Almén MS. et al. The GPCR repertoire in the demosponge Amphimedon queenslandica: insights into the GPCR system at the early divergence of animals. BMC Evol Biol. 2014;14:270
6. Paavola KJ, Hall RA. Adhesion G protein-coupled receptors: signaling, pharmacology, and mechanisms of activation. Mol Pharmacol. 2012;82:777-83
7. Nijmeijer S, Vischer HF, Leurs R. Adhesion GPCRs in immunology. Biochem Pharmacol. 2016;114:88-102
8. Nechiporuk T, Urness LD, Keating MT. ETL, a novel seven-transmembrane receptor that is developmentally regulated in the heart. ETL is a member of the secretin family and belongs to the epidermal growth factor-seven-transmembrane subfamily. J Biol Chem. 2001;276:4150-7
9. Serban F, Artene SA, Georgescu AM. et al. Epidermal growth factor, latrophilin, and seven transmembrane domain-containing protein 1 marker, a novel angiogenesis marker. Onco Targets Ther. 2015;8:3767-74
10. Masiero M, Simões FC, Han HD. et al. A core human primary tumor angiogenesis signature identifies the endothelial orphan receptor ELTD1 as a key regulator of angiogenesis. Cancer Cell. 2013;24:229-41
11. Weidner N, Semple JP, Welch WR. et al. Tumor angiogenesis and metastasis-correlation in invasive breast carcinoma. N Engl J Med. 1991;324:1-8
12. Yao H, Liu N, Lin MC. et al. Positive feedback loop between cancer stem cells and angiogenesis in hepatocellular carcinoma. Cancer Lett. 2016;379:213-9
13. Wong PP, Bodrug N, Hodivala-Dilke KM. Exploring novel methods for modulating tumor blood vessels in cancer treatment. Curr Biol. 2016;26:1161-6
14. Tahmasebi Birgani M, Carloni V. Tumor microenvironment, a paradigm in hepatocellular carcinoma progression and therapy. Int J Mol Sci. 2017;18:E405
15. McAnulty RJ. Fibroblasts and myofibroblasts: their source, function and role in disease. Int J Biochem Cell Biol. 2007;39:666-71
16. Cirri P, Chiarugi P. Cancer associated fibroblasts: the dark side of the coin. Am J Cancer Res. 2011;1:482-97
17. Gupte J, Swaminath G, Danao J. et al. Signaling property study of adhesion G-protein-coupled receptors. FEBS Lett. 2012;586:1214-9
18. Hamilton SR, Aaltonen LA. Tumours of the liver and intrahepatic bile ducts. WHO International Histological Classification of Tumours, No. 8, 2nd edn. France: IARC. 2000
19. Edmondson HA, Steiner PE. Primary carcinoma of the liver: a study of 100 cases among 48,900 necropsies. Cancer. 1954;7:462-503
20. Llovet JM, Brú C, Bruix J. Prognosis of hepatocellular carcinoma: the BCLC staging classification. Semin Liver Dis. 1999;19:329-38
21. Shimada M, Hamatsu T, Yamashita Y. et al. Characteristics of multicentric hepatocellular carcinomas: comparison with intrahepatic metastasis. World J Surg. 2001;25:991-5
22. Imamura H, Matsuyama Y, Tanaka E. et al. Risk factors contributing to early and late phase intrahepatic recurrence of hepatocellular carcinoma after hepatectomy. J Hepatol. 2003;38:200-7
23. Kuang DM, Wu Y, Chen N. et al. Tumor-derived hyaluronan induces formation of immunosuppressive macrophages through transient early activation of monocytes. Blood. 2007;110:587-95
24. Vandeputte C, Docherty JR. Vascular actions of 3, 4-methylenedioxymethamphetamine in alpha (2A/D)-adrenoceptor knockout mice. Eur J Pharmacol. 2002;457:45-9
25. Towner RA; Jensen RL; Colman H. et al. ELTD1, a potential new biomarker for gliomas. Neurosurgery. 2013;72:77-90
26. Baldewijns MM, van Vlodrop IJ, Vermeulen PB. et al. VHL and HIF signalling in renal cell carcinogenesis. J Pathol. 2010;221:125-38
27. Tang XL, Wang Y, Li DL. et al. Orphan G protein-coupled receptors (GPCRs): biological functions and potential drug targets. Acta Pharmacol Sin. 2012;33:363-71
28. Mazzocca A, Fransvea E, Dituri F. et al. Down-regulation of connective tissue growth factor by inhibition of transforming growth factor beta blocks the tumor-stroma cross-talk and tumor progression in hepatocellular carcinoma. Hepatology. 2010;51:523-34
29. Lin ZY, Chuang YH, Chuang WL. Cancer-associated fibroblasts up-regulate CCL2, CCL26, IL6 and LOXL2 genes related to promotion of cancer progression in hepatocellular carcinoma cells. Biomed Pharmacother. 2012;66:525-9
30. Kuperwasser C, Chavarria T, Wu M. et al. Reconstruction of functionally normal and malignant human breast tissues in mice. Proc Natl Acad Sci USA. 2004;101:4966-71
31. Bhowmick NA, Neilson EG, Moses HL. Stromal fibroblasts in cancer initiation and progression. Nature. 2004;432:332-7
32. Bhowmick NA, Chytil A, Plieth D. et al. TGF-β signaling in fibroblasts modulates the oncogenic potential of adjacent epithelia. Science. 2004;303:848-51
Author contact
Corresponding author: Rong-ping Guo, Department of Hepatobiliary Oncology, Sun Yat-sen University Cancer Center, 651 Dongfeng Road, Guangzhou 510060, Guangdong, People's Republic of China. E-mail address: guorporg.cn; Tel.: +86-20-87342266

 Global reach, higher impact
Global reach, higher impact