Impact Factor ISSN: 1837-9664
J Cancer 2017; 8(14):2704-2712. doi:10.7150/jca.20665 This issue Cite
Research Paper
IDH1 R132H Mutation Is Accompanied with Malignant Progression of Paired Primary-Recurrent Astrocytic Tumours
1. Department of Neurosurgery, the Fourth Affiliated Hospital, Harbin Medical University, Harbin, China;
2. Department of Geriatrics, the Second Affiliated Hospital, Harbin Medical University, Harbin, China;
3. Department of Neurosurgery, the First Affiliated Hospital, Harbin Medical University, Harbin, China;
4. Department of Neurosurgery, Harbin Medical University Cancer Hospital, Harbin, China;
5. Department of Neurosurgery, the First Hospital of Harbin, Harbin, China;
6. Department of Pathology, the Second Affiliated Hospital, Harbin Medical University, Harbin, China;
7. Department of Pathology, the First Affiliated Hospital, Harbin Medical University, Harbin, China.
Received 2017-4-20; Accepted 2017-6-27; Published 2017-8-22
Abstract
IDH1 R132H mutation is an important marker of survival in patients with gliomas. Although there are many changes of genes in tumour malignant progression, IDH1 R132H mutation status in glioma progression remained unclear. Here, an in-depth characterization of IDH1 R132H mutations were assessed by immunohistochemistry in 55 paired primary-recurrent astrocytomas tissues, including 5 paired primary pilocytic astrocytoma (pPA, WHO grade I), 35 paired primary low grade astrocytoma (pLGA, WHO grade II and III) and 15 paired primary high grade astrocytoma (pHGA/ Glioblastoma, WHO grade IV). Meanwhile, the DNA was isolated from paired samples, and PCR amplification was used for IDH1 exon4 sequencing. Nonparametric test, KM and Cox models were used to examine the statistical difference and survival function. We found that the percent of IDH1 R132H mutation was 68.6% (24/35) in pLGA group, but no IDH1 mutation was found in pPA and pHGA groups. Meanwhile, the results from immunohistochemistry and DNA sequencing showed that, compared with primary astrocytoma, there was no change of IDH1 status in recurrent astrocytoma whatever tumour pathological grade raise or indolent. The pPA group has the longest recurrence-free period (RFP) and overall survival (OS) in three groups (p<0.01), while the pHGA group has the shortest ones (p<0.01). In pLGA group, the IDH1 R132H mutation subgroup has longer RFP than IDH1 wild type subgroup (p<0.01), but the OS has no statistical difference between two subgroups (p>0.6). Additionally, IDH1 R132H mutation independently predicted a long RFP in patients with pLGA (HR 1.073, 95% CI 0.151-0.775, p<0.01).
Keywords: Astrocytoma, IDH1 R132H mutation, Progression, Primary-recurrence.
Introduction
Gliomas are the most common central nervous system (CNS) tumours, which are classified according to histological criteria into groups including Astrocytic tumours, Oligodendroglial tumours, Oligoastrocytic tumours and other tumours [1]. Astrocytomas, arise from astrocytes are the most common type of gliomas, and representing 64% of human CNS malignances [2]. Although the wide application of treatment combines microsurgical tumorectomy with standard radiotherapy and chemotherapy, majority of gliomas remain relapse and their prognoses remain poor.
As is known, malignant tumour progression is a dynamic process which is accompanied by an altered molecular phenotype [3] including emergence of novel molecular alterations or the loss of previously alterations, such as gene mutation, DNA promoter methylation and chromosome codeletion. Whether there was new occurrence of IDH1 R132H mutation in astrocytoma recurrence? And some pLGA patients' pathological grade rise when tumour recurrence, while others' remain unchanged, whether IDH1 mutation influences this process? In present study, we investigated the change of IDH1 R132 status in the malignant progression of astrocytoma. Meanwhile, the survival analysis among IDH1 mutation, tumour progression and clinical features were also evaluated.
Materials and Methods
Patient Population
This study included 55 patients who underwent surgery twice at the same hospital from 2005 to 2014. The inclusion criteria included the following: older than 18 years old at the time of the primary diagnosis, a histopathologic diagnosis of astrocytoma (WHO grade I-IV), and the availability of formalin fixed, paraffin-embedded (FFPE) tumour tissue block. Those patients were divided into three groups:(1) primary pilocytic astrocytoma (PA, ICD-O 9421/1, WHO grade I) group: PA in primary surgery and LGA in second surgery respectively, 5 patients (10 samples); (2) primary low grade astrocytoma (LGA, ICD-O 9400/3 and ICD-O 9401/3, WHO grade II and III) group: LGA in primary surgery and LGA or HGA in second surgery respectively, 35 patients (70 samples); (3) primary high grade astrocytoma (HGA, ICD-O 9440/3, WHO grade IV) group: glioblastoma (GBM) in primary and second surgeries, 15 patients (30 samples). None of the 55 patients received any treatments before the primary surgery. Most of them received standard radiotherapy (60 Gy), and some of the patients received concomitant-adjuvant chemotherapy (Temozolomide/ other) after the primary tumour resection. The recurrence-free period (RFP) was defined as the time from the date of the primary surgery to the first MRI-confirmed tumour recurrence. All of patients underwent second surgery after imageological diagnosis of tumour recurrence. Overall survival (OS) was defined as the interval between primary surgery and death or last follow-up, while recurrence survival (RS) was calculated from the second surgery to the date of death or last follow-up. The mean follow-up period was 1425.4 days (range 289-3520 days), during which 44 (80%) patients died of astrocytoma and 11 (20%) patients remained alive. No patients were lost to follow-up. Tumour size was calculated as follows based on enhanced MR or CT images (on the layer with the maximum amount of tumour): long diameter (cm) X wide diameter (cm) X thickness (cm) X 1/2. All specimens were selected after histopathological review to ensure that we would have sufficient tissue for analysis. We prepared 4 adjacent tissue sections from each selected block: the first tissue section was for hematoxylin and eosin staining; the second tissue section was for anti-IDH1 R132H mutation immunostaining; the third tissue section was for anti-IDH1 immunostaining; and the fourth tissue section was for genetic analysis of IDH1. Slides for histology and immunohistochemistry were 4 μm thick, and slides for genetic analysis were 10 μm thick. Treatment information was obtained for the time from the primary surgery, including any adjuvant treatment. All research protocols were reviewed and approved of the Committee on Human Research at the Harbin Medical University, China.
Histological Analysis and Immunohistochemistry
A systematic neuro-pathological review was performed based on the 2007 WHO classification guidelines for tumours of the CNS [4]. All of the paired samples of slides were reviewed to determine their histological categories. Samples were then selected for immunohistochemical staining. Antigen retrieval was performed by submitting the tissues to heat treatment in 10 mmol/L sodium citrate buffer (pH 6.0) for 3 min (for IDH1) or in EDTA (pH 9.0) for 2 min (for IDH1 R132H mutation). Primary antibodies against IDH1 (N-terminal, dilution 1:150, Abcam, USA) and IDH1 R132H mutation [5] (clone H09, dilution 1:50; Maixin-BIO, China) were added to the tissues, and they were incubated overnight at 4oC. The samples were then incubated with horseradish peroxidase-labelled secondary antibodies using an immunohistochemical kit (KIT-5930, Maixin-bio, China) for 40min at room temperature. Each staining batch included a negative control that was processed without the primary antibody.
The data analyses were performed under a light microscope (Leica SP2, Leica Optical Co. Ltd., Germany). Two experienced pathologists who were blinded to the clinical background and the results of genetic analyses were recorded as the final result. We used the following 2-tiered system: negative, no tumour cells stained; positive, partly or all identifiable tumour cells stained.
Gene Expression Analysis
To generate high quality Genomic DNA from FFPE samples, GeneRead DNA FFPE Kit (QIAGEN, Carlsbad, USA) was used to extract DNA from paired samples. Introns and intron/exon boundaries of exon4 of the IDH1 gene were amplified by polymerase chain reaction using JumpStart RED Taq Ready Mix (Sigma Aldrich, St Louis, USA), as described previously [6]. Polymerase chain reaction products were sequenced on a 3730 Genetic Analyzer (Applied Biosystems, Foster City, USA) following standard procedures. Primers used for polymerase chain reaction amplification and sequencing were IDH1 exon4F (ACGGTCTTCAGAGAAGC), IDH1 exon4R (GGTGTAGATACCAAAAGATAAGAAT) [7]. The data were analysed by Chinese national human genome center.
Statistical Analysis
The raw data were tabulated using Microsoft Excel (Microsoft Corp.). Kaplan-Meier (KM) survival analysis was used to determine the difference of RFP, RS and OS, and the results were analysed with the log-rank test. To adjust for potential confounders, Cox proportional hazards models were used to evaluate hazard ratio (HR) of recurrence or death according to the clinical features in present investigation. Statistical analyses were performed with SPSS 13.0 (SPSS Inc. Chicago, USA) and GraphPad Prism 6.0 (GraphPad Software Inc. California, USA). For all statistical investigations, the tests used to determine the level significance were 2-sided. A (2-tailed) P value threshold of 0.05 was considered to be statistically significant.
Result
Patient Characteristics
The clinical features of the patients in the three groups were shown in Table 1 and Figure 1A. A total of 49 patients (89.1%) underwent gross total resection. 52 patients (94.5%) received standard radiotherapy (60 Gy), and 54.5% of the patients received concomitant-adjuvant chemotherapy (Temozolomide/other) after the primary tumour resection. Patients in pPA group have the lowest age at the time of primary or second surgery in three groups (p<0.05, Figure 1B), while the pLGA patients with IDH1 R132H mutation have lower age than patients with IDH1 wild type (p<0.05, data not shown). The tumour size at the time of primary surgery in pPA group was smaller than other two groups (p<0.05, Figure 1C). There were no significant differences in gender, race, or the laterality, location, extent of the resection, or tumour size at the time of second surgery across the groups (data not shown). The KM survival analysis indicated that the pPA group has the longest RFP (median, 1730 days, 95% CI 1494-1966 days) and OS (median, 3118 days, 95% CI 3065-3170 days), while the pHGA group has the shortest RFP (median, 275 days, 95% CI 138-411 days) and OS (median, 480 days, 95% CI 368-591 days) (p<0.01, Figure 1D and E). Additionally, the RS were also significantly different among the three groups (p<0.05, Figure 1F).
Demographic and Clinical Characteristics of Study Patients
| Characteristic | Pilocytic astrocytoma (PA)WHO grade I (n=5) | Low grade astrocytoma (LGA)WHO grade II and III (n=35) | High grade astrocytoma (HGA)WHO grade IV (n=15) |
|---|---|---|---|
| Age at primary surgery, years | |||
| Mean±SE | 27.2±5.3 | 42.7±9.9 | 45.9±12.9 |
| Range | 22-34 | 30-66 | 22-68 |
| Age at recurrent surgery, years | |||
| Mean±SE | 32.2±4.4 | 45±9.6 | 46.4±13.8 |
| Range | 28-38 | 31-68 | 23-69 |
| Sex | |||
| Male | 2 (40%) | 10 (28.6%) | 6 (40%) |
| Female | 3 (60%) | 25 (71.4%) | 9 (60%) |
| Race/ethnicity | |||
| Han | 5 (100%) | 34 (97.1%) | 5 (100%) |
| Mongolian | 0 | 1 (2.9%) | 0 |
| Laterality | |||
| Left | 2 (40%) | 24 (68.6%) | 11 (73.3%) |
| Right | 3 (60%) | 11 (31.4%) | 4 (26.7%) |
| Tumor Location | |||
| Frontal | 1 (20%) | 16 (45.7%) | 6 (40%) |
| Temporal | 3 (60%) | 12 (34.3%) | 2 (13.3%) |
| Parietal | 0 | 5 (14.3%) | 4 (26.7%) |
| Occipital | 1 (20%) | 2 (5.7%) | 3 (20%) |
| IDH1 R132H mutation | |||
| 0 | 24 (68.6%) | 0 | |
| Tumor size in primary surgery, cm3 | |||
| Mean±SE | 14.8±3.7 | 39±16.3 | 40.9±11.5 |
| Range | 8.9-20.2 | 12-83.6 | 18.6-64.1 |
| Tumor size in recurrent surgery, cm3 | |||
| Mean±SE | 38.3±18.1 | 39.9±17.5 | 50.4±19.9 |
| Range | 21-54.5 | 15-85 | 30-110 |
| Extent of resection in primary surgery | |||
| Gross total resection | 3 (60%) | 32 (91.4%) | 14 (93.3%) |
| Subtotal resection (50-99%) | 2 (40%) | 3 (8.6%) | 1 (6.7%) |
| Adjuvant therapy after primary surgery | |||
| Radiotherapy only | 2 (40%) | 18 (51.4%) | 2 (13.3%) |
| Chemotherapy only | 0 | 0 | 0 |
| Radiotherapy plus Chemotherapy | 0 | 17 (48.6%) | 13 (86.7%) |
| First presenting symptom | |||
| Headache | 2 (40%) | 15 (42.9%) | 8 (53.3%) |
| Mental status change | 0 | 2 (5.7%) | 0 |
| Motor or movement change | 0 | 7 (20%) | 4 (26.7%) |
| Seizure/Epilepsy | 0 | 6 (17.1%) | 0 |
| Sensory or visual change | 0 | 5 14.3%) | 3 (20%) |
| Physical examination | 3 (60%) | 0 | 0 |
| Mortality rate | |||
| 2 (40%) | 28(80%) | 14 (93.3%) | |
| Recurrence-Free Period(RFP), days | |||
| Mean±SE | 1894±489.9 | 845.8±541.9 | 386.3±273.3 |
| Range | 1337-2774 | 113-2417 | 175-1150 |
| Recurrent survival(RS), days | |||
| Mean±SE | 1004.6±501 | 722.7±768.6 | 240.77±189.82 |
| Range | 373-1748 | 143-2871 | 70-598 |
| Overall survival(OS), days | |||
| Mean±SE | 2898.6±480.7 | 1568.6±821.8 | 627±378.2 |
| Range | 1993-3118 | 355-3520 | 309-1654 |
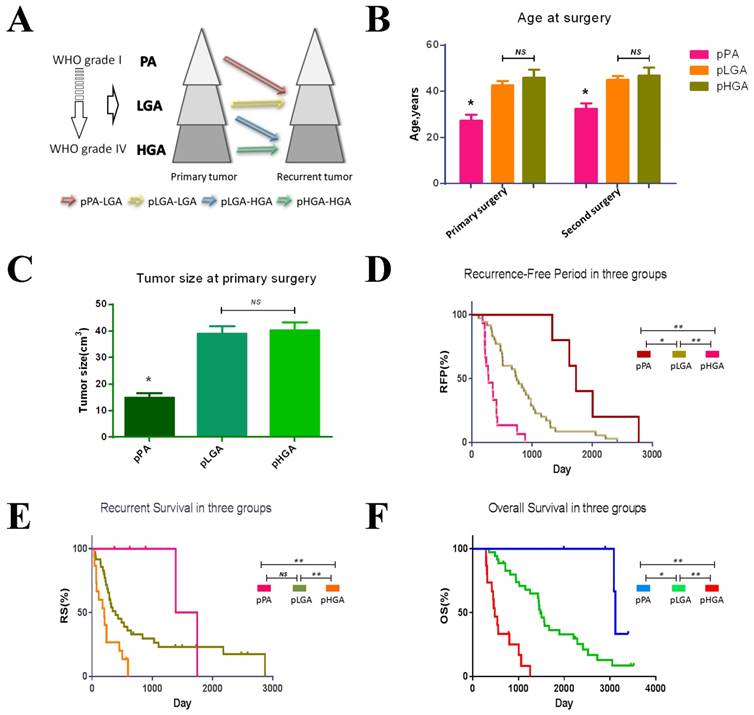
Clinical features in three groups. (A) Tumour progression in three groups. (B) Age of primary and recurrent tumours in three groups. The patients with pPA have the younger age than other groups in twice surgeries (p<0.05). (C) Tumour size at primary surgery. Tumour size in pPA group was smaller than other groups (p<0.05). (D) Recurrence Free Period in three groups. The pPA group has the longest RFP, and the pHGA group has the shortest RFP. (E) Recurrent Survival in three groups. The pHGA group has the shortest RS (p<0.05), but the pPA group and pLGA have no statistic different. (F) Overall Survival in three groups. The pPA group has the longest OS, and the pHGA group has the shortest OS. * p<0.05, ** p<0.01, NS p>0.05
Immunohistochemical Analysis upon Tumour Progressions
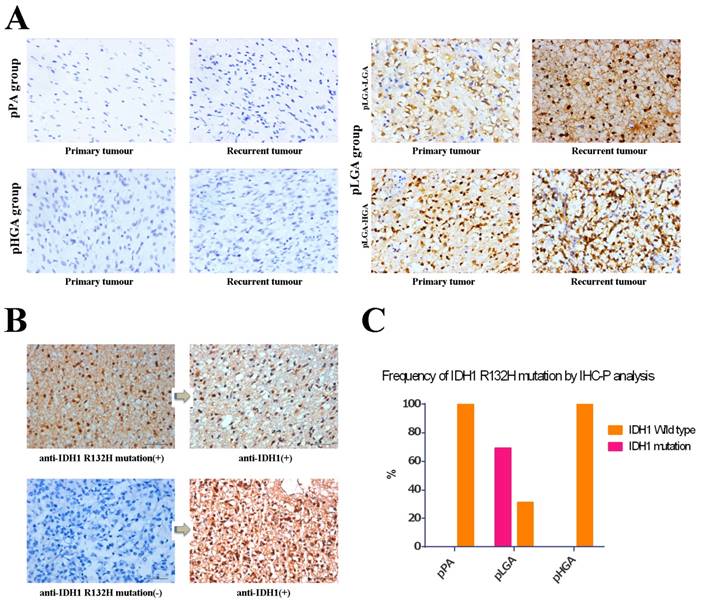
We used anti-IDH1 R132H mutation antibodies detected paired primary-recurrent samples in different status of malignant tumour progression, and the anti-IDH1 R132H mutation antibodies labeled both nuclei and cytoplasm of the tumour cells (Figure 2A). We observed periaxonal, perivascular, and subpial arrangements of IDH1 R132H mutation immunolabeled cells in the infiltration zone of tumours, but there were no IDH1 R132H mutation positive cells in normal brain tissues (the donor had died of myocardial infarction). Furthermore, we used anti IDH1 antibody to re-stain the samples which anti-IDH1 R132H mutation immunostaining as negative, however, the results showed anti-IDH1 positive (Figure 2B). In all, we found anti-IDH1 R132H mutation immunostaining positive in 24 (68.6%) of the 35 patients in pLGA group, but there were no patients with IDH1 R132H mutation in pPA and pHGA groups (Figure 2C). Meanwhile, there were concordant immunostaining results compared primary tumour with recurrent tumour in all 55 patients of three groups (data not shown).
IDH1 and IDH1 R132H mutation expression were analysed using standard IHC-P staining (x400). (A) IDH1 R132H mutation immunohistochemical staining between primary sample and recurrent sample in three groups. (B) anti-IDH1 vs. anti-IDH1 R132H mutation immunohistochemical staining in the same samples. Left and right pictures are the same samples, anti-IDH1 R132H mutation (H09) immunostaining is in the left while anti-IDH1 immunostaining is in the right respectively. (C) Frequency of anti-IDH1 R132H mutation immunostaining positive in three groups.
Genetic Analysis and Comparison with Immunohistochemical Analysis
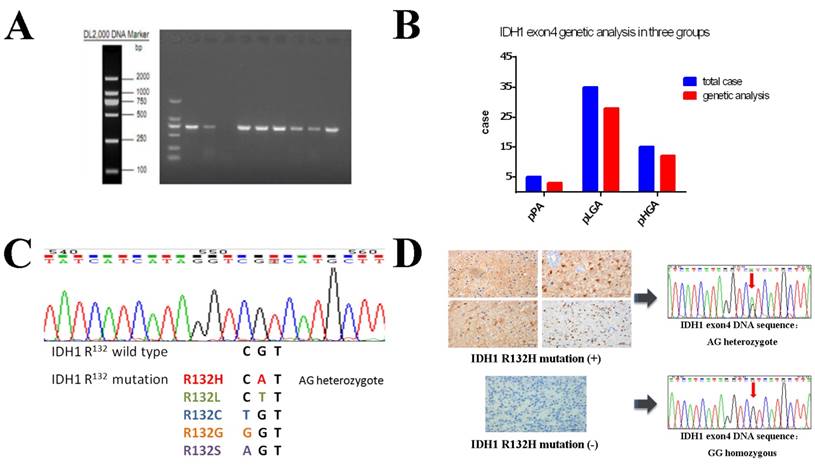
High quality genomic DNA samples were extracted from 43 (78.2%) paired FFPE samples (Figure 3A), and genetic expression analysis were successful in 3 patients of the pPA group, 28 patients of the pLGA group and 12 patients of the pHGA group (Figure 3B). IDH1 mutations were detected in 19 (44.2%) of the 43 paired cases, and all of them were R132H mutations (AG heterozygote), while no other types of IDH1 R132 mutations were found (e.g. R132L, R132C, R132G or R132S); in other 24 (55.8%) paired cases, we detected IDH1 status as wild type, because all of them were GG homozygous (Figure 3C).
IDH1 R132 mutation status was analysed using gene sequencing. (A) PCR amplification products of DNA samples with IDH1 exon4. The 43 of 55 paired samples were successful analysed in three groups. (B) Proportion of IDH1 gene expression analysis in three groups: 3 of 5 in pPA group, 28 of 35 in pLGA group and 12 of 15 in pHGA group. (C) The results of wild⁃type IDH1 and R132H mutant sequencing. A total of 38 codon mutations among 86 cases (35.56% ) were found and all of mutated cases had CGT→CAT transition change, a single amino acid from arginine to histidine (R132H). (D) Comparison of gene sequencing results with corresponding immunohistochemical staining expression
We compared the results of gene sequencing with immunohistochemical staining: In all 19 paired cases that were detected as IDH1 R132H mutation positive by gene analysis, immunostaining of IDH1 R132H mutation antibody (clone H09) showed concordant results (Figure 3D up). All 24 paired cases that were assessed as negative on immunohistochemical staining showed IDH1 wild type by gene sequencing (Figure 3D down).
Survival Analysis based on Tumour Progression and IDH1 R132H mutation status
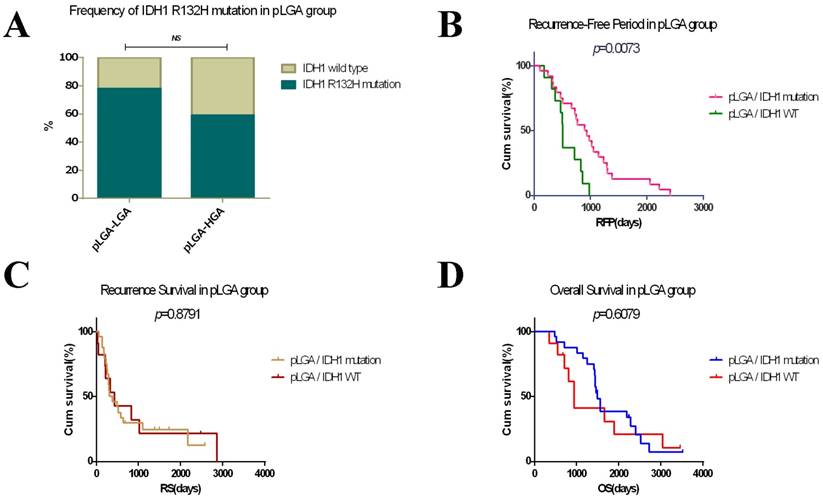
In the progression of astrocytomas, the sequels of pLGAs were dissimilitude. Some of pLGAs transformed into glioblastomas when tumour recurrences, while others remain pure. So we next divided pLGAs into two subgroups by the pathological grade of recurrent tumours: pLGA-LGA subgroup and pLGA-HGA subgroup (Figure 1A). To determine the prognostic effects of the IDH1 R132H mutation, we analysed the frequency of IDH1 R132H mutation in the two subgroups, the pLGA-LGA subgroup has a higher proportion of IDH1 mutation than pLGA-HGA subgroup, but no statistical significant difference has been obtained in primary and recurrent tumour between them (Figure 4A). Meanwhile, a KM survival analysis indicated that the IDH1 R132H mutation subgroup has a longer RFP (median, 898 days, 95% CI 615-1180 days) than IDH1 wild type subgroup (median, 508 days, 95% CI 467-549 days) in all patients with pLGA (p<0.05, Figure 4B). However, there was no significant difference in RS and OS between the two subgroups (p>0.6, Figure 4C and 4D). A Cox regression analyses demonstrated that the IDH1 R132H mutation was an independent predictor of lengthen RFP (HR 1.073, 95% CI 0.151-0.775, p<0.01). However, no independent predictor of OS or RS was identified in this study.
Survival analyses in patients with pLGA. (A) Frequency of IDH1 R132H mutation in primary/recurrent tumour between pLGA-LGA and pLGA-HGA subgroups (p>0.05). (B) The IDH1 R132H mutation subgroup has a longer RFP than IDH1 wild type subgroup in patients with pLGA (p<0.05). (C) There was no significant difference in RS between the two subgroups (p>0.05). (D) There was no significant difference in OS between the two subgroups (p>0.05).
Discussion
As an incurable malignant tumour, patients with astrocytomas will experience a period of “tumour-silence” after tumorectomy, and then tumour recurrence will occur. Tumours recur in the resection margin in nearly 90% of malignant CNS neoplasm after primary surgery and additional therapies [8]. However, during the dynamic process of malignant progression, immune responses, biological behaviors, genetic alterations and clinical outcomes are constantly changing. Recently, IDH1 mutation has been recognized as an important biomarker and a strong prognostic factor in brain tumours [1, 9, 10]. Although several studies have evaluated the frequency of IDH1 R132H mutation in glioma [11-15], they have all evaluated data in stationary and isolated tumour stages (e.g., tumours with a low or high clinicopathological grade). To overcome this barrier, in this study, we investigated the dynamic changes that occur in IDH1 gene status and other relevant clinical factors during astrocytoma recurrence in tumours that advanced from pPA to LGA, pLGA to LGA/HGA and pHGA to HGA.
Preusser et al. [5] found that clone H09 immunostaining produced better results with less nonspecial background staining and more distinct labeling of tumour cells than imab-1 antibody. So we in this study use the antibody (H09) to label the IDH1 R132H mutation positive cells. In paired samples that successful detected with immunohistochemical antibody (clone H09) and DNA sequencing, all of the immunostaining results showed consistent IDH1 R132 status as the corresponding results of gene sequencing. Our data confirm anti-IDH1 R132H mutation immunostaining as a reliable and suitable method to evaluate IDH1 status, especially use of antibody clone H09. Compared with genetic analysis, IDH1 R132 status could be easily, fast and powerful detected by immunohistochemical staining. Furthermore, when there is a tiny biopsy or tumour sample mixed with normal brain tissue, immunohistochemistry may be more sensitive method for the detection of IDH1 R132H mutation than genetic analysis.
Previous studies showed that IDH1 R132H mutation was present in 80% of WHO grade II and III low grade glioma (LGG) but less than 5% of primary high grade glioma (HGG/ glioblastoma) [7, 15-19]. Because of the high frequencies in LGG, IDH1 R132H mutation was identified as a common early and important event during tumour development [7]. Furthermore, after dissected multiple biopsies from the same patients, and detected the status of the IDH1 mutation, TP53 mutation and loss of 1p/19q, Watanabe et al. [16] also suggested that IDH1 mutation was early genetic event in the development of glioma. In our study, we found 68.6% pLGA patients have IDH1 R132H mutation, but there was no IDH1 R132H mutation has been found in pPA patients and there was no new mutation in the progression from PA to LGA. Those results may suggest that compared with LGA harboring IDH1 R132H mutation which was an early molecular event, the LGA without IDH1 mutation originated from PA and have the different molecular origin. Further study with a larger cohort would be needed to evaluate this mutation in patients with pPA.
Tumour progression is an inescapable phenomenon in patients suffering from gliomas. A very recent research by Cai et al. [20] showed that IDH1 R132H mutation occurred stably in matched primary-recurrent LGGs and HGGs only by immunohistochemical staining method. After evaluating the IDH1 R132 status of paired primary-recurrent astrocytic tumours, we found that not only in LGA and HGA groups but also in PA group, the concordance rate of the paired assays was 100% by immunohistochemical staining and gene sequencing. This result further suggests that it was not IDH1 mutation, but other molecular aberrations accumulate during astrocytoma malignant transformation [21]. But Lass and colleagues [22], with the method of directly gene sequencing, showed that there were 3/47 paired gliomas (1 astrocytoma and 2 oligodendrogliomas) changed its IDH1 status in recurrence, and in all of the 3 instances the IDH1 R132H mutation were observed in the primary tumour but not be detected in the respective recurrent tumours. Meanwhile, the loss of previously detected molecular alterations also affecting TP53, isolated LOH1p, isolated LOH19q and combined LOH1p/19q was also observed at tumour recurrence. The limited number of tumour tissues which used for DNA extraction or the co-presence of contaminating normal brain tissues maybe two feasible reasons that could explain his results.
In pLGA, during tumour progression, the sequelae of LGA are unpredictable. We therefore compared the primary tumours in pLGA-LGA subgroup with those in pLGA-HGA subgroup. However, there was no significant difference in the proportions of IDH1 R132H mutation between the two subgroups. Together with our results that all primary tumours keep constant IDH1 R132 status in respectively recurrent tumour, we concluded that there was no association between IDH1 R132H mutation and pLGA malignant transformation.
Evidence has accumulated in the reports regarding the relationship between IDH1 R132H mutation and survival in different WHO grade or histopathological type of gliomas, particularly high grade gliomas [6, 17, 20, 23-25]. Nevertheless, the prognostic significance of IDH1 R132H mutation in LGA and PA was more debatable. Dubbink et al. [26] investigated the IDH1 R132 status in 49 LGA patients who treated with temozolomide at the time of progression after radiotherapy and demonstrated the association between IDH1 mutation and improved OS. Iwadate et al. [27] found that IDH1 mutation was significant for OS in patients with diffuse astrocytomas (belongs to LGA, WHO grade II), in contrast, for oligodendrogliomas (belongs to low grade oligodendroglial tumours, WHO grade II), IDH1 status were not significant for progression free survival (PFS) or OS. Waqar et al. [28] and Jungk et al. [29] also found that IDH1 R132H mutation was an independent positive prognosticator for OS in patients with WHO grade II astrocytomas. On the other hand, Ahmadi et al. evaluated 100 diffuse astrocytomas patients who did not receive adjuvant treatment and found no correlation between IDH1 mutation and clinical outcome in terms of OS and PFS [30]. Meanwhile, due to the relatively small size and the lack of samples with IDH1 R132H mutation in PA and HGA groups, survival analysis of IDH1 mutations were not performed in the two groups. We only evaluated the prognostic impact of IDH1 mutation in Chinese patients with pLGA. We found that patients with pLGA harboring IDH1 R132H mutation have a longer RFP than ones with IDH1 wild type, but the OS in the two subgroups has no significant difference. In addition, we also identified IDH1 R132H mutation as an independent prognostic marker in lengthen RFP of LGAs after adjusting for age, gender, extent of resection tumour size and chemotherapy.
Following differences perhaps explained such discrepancy in relationship between IDH1 R132H mutation and survival. Firstly, methodology of IDH1 status detecting, most of studies use DNA sequencing or immunohistochemical staining to evaluate IDH1 R132H mutation, but we in this research used the two methods at the same time to enhance the accuracy of detecting. Secondly, the range of cases included in studies were different, due to the difference in molecular and clinical behaviors, LGA was defined as WHO grade II and III astrocytic tumours [9]. However, patients in Iwadate, Waqar, Jungk and Ahmadi's studies were only those who have been diagnosed as WHO grade II astrocytoma. Thirdly, adjuvant treatments may potentially influence the evaluation of prognostic impact of IDH1 R132H mutation in LGAs. Contrast to other studies, all of patients with LGA in our study received early standard radiotherapy after the first surgery, and 48.6% patients received concomitant adjuvant chemotherapy. Consistent with us, Juratli [31] also showed that in LGA patients who had early radiation therapy, cases with mutated IDH1 had a significantly longer PFS but no improved OS than those who had no IDH1 R132H mutation. Additionally, in contrast to these studies which only used primary samples and defined sign of radiological progression, death or last follow-up as terminal events of PFS which could not exact evaluate the time of tumour progression, we studied paired primary-recurrent gliomas and all of the patients have the accurate date of RFP.
Conclusions
In summary, we used immunohistochemistry and DNA sequencing to detect the status of IDH1 in paired primary-recurrent samples, and found that there is no change of IDH1 R132 status when glioma recurrence occurred. Furthermore, in patients with LGA, there was no association between IDH1 R132H mutation and LGA malignant transformation, but tumour with this mutation had a longer RFP than wild type, and it is an independent prognostic factor for a favourable RFP.
Acknowledgements
This research was supported by the Heilongjiang Natural Science Foundation (H2016027, H2016034), the Health and Family Planning Commission of Heilongjiang Province (No. 2014-313), Harbin medical university scientific research innovation fund (2016LCZX06).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK. et al. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta neuropathologica. 2016;131:803-20
2. Ostrom QT, Gittleman H, Fulop J, Liu M, Blanda R, Kromer C. et al. CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2008-2012. Neuro-oncology. 2015;17(Suppl 4):iv1-iv62
3. Alessandrini F, Ceresa D, Appolloni I, Marubbi D, Malatesta P. Noninvasive Monitoring of Glioma Growth in the Mouse. Journal of Cancer. 2016;7:1791-7
4. Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A. et al. The 2007 WHO classification of tumours of the central nervous system. Acta neuropathologica. 2007;114:97-109
5. Preusser M, Wohrer A, Stary S, Hoftberger R, Streubel B, Hainfellner JA. Value and limitations of immunohistochemistry and gene sequencing for detection of the IDH1-R132H mutation in diffuse glioma biopsy specimens. Journal of neuropathology and experimental neurology. 2011;70:715-23
6. Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P. et al. An integrated genomic analysis of human glioblastoma multiforme. Science (New York, NY). 2008;321:1807-12
7. Balss J, Meyer J, Mueller W, Korshunov A, Hartmann C, von Deimling A. Analysis of the IDH1 codon 132 mutation in brain tumors. Acta neuropathologica. 2008;116:597-602
8. Hamard L, Ratel D, Selek L, Berger F, van der Sanden B, Wion D. The brain tissue response to surgical injury and its possible contribution to glioma recurrence. Journal of neuro-oncology. 2016;128:1-8
9. Brat DJ, Verhaak RG, Aldape KD, Yung WK, Salama SR, Cooper LA. et al. Comprehensive, Integrative Genomic Analysis of Diffuse Lower-Grade Gliomas. The New England journal of medicine. 2015;372:2481-98
10. Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer cell. 2010;17:98-110
11. Lewandowska MA, Furtak J, Szylberg T, Roszkowski K, Windorbska W, Rytlewska J. et al. An analysis of the prognostic value of IDH1 (isocitrate dehydrogenase 1) mutation in Polish glioma patients. Molecular diagnosis & therapy. 2014;18:45-53
12. Myung JK, Cho HJ, Park CK, Kim SK, Phi JH, Park SH. IDH1 mutation of gliomas with long-term survival analysis. Oncology reports. 2012;28:1639-44
13. Riehmer V, Gietzelt J, Beyer U, Hentschel B, Westphal M, Schackert G. et al. Genomic profiling reveals distinctive molecular relapse patterns in IDH1/2 wild-type glioblastoma. Genes, chromosomes & cancer. 2014;53:589-605
14. Takano S, Kato Y, Yamamoto T, Kaneko MK, Ishikawa E, Tsujimoto Y. et al. Immunohistochemical detection of IDH1 mutation, p53, and internexin as prognostic factors of glial tumors. Journal of neuro-oncology. 2012;108:361-73
15. Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W. et al. IDH1 and IDH2 mutations in gliomas. The New England journal of medicine. 2009;360:765-73
16. Watanabe T, Nobusawa S, Kleihues P, Ohgaki H. IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. The American journal of pathology. 2009;174:1149-53
17. Sanson M, Marie Y, Paris S, Idbaih A, Laffaire J, Ducray F. et al. Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 2009;27:4150-4
18. Kang MR, Kim MS, Oh JE, Kim YR, Song SY, Seo SI. et al. Mutational analysis of IDH1 codon 132 in glioblastomas and other common cancers. International journal of cancer. 2009;125:353-5
19. Bleeker FE, Lamba S, Leenstra S, Troost D, Hulsebos T, Vandertop WP. et al. IDH1 mutations at residue p.R132 (IDH1(R132)) occur frequently in high-grade gliomas but not in other solid tumors. Human mutation. 2009;30:7-11
20. Cai J, Zhu P, Zhang C, Li Q, Wang Z, Li G. et al. Detection of ATRX and IDH1-R132H immunohistochemistry in the progression of 211 paired gliomas. Oncotarget. 2016;7:16384-95
21. Cheng Y, Ng HK, Ding M, Zhang SF, Pang JC, Lo KW. Molecular analysis of microdissected de novo glioblastomas and paired astrocytic tumors. Journal of neuropathology and experimental neurology. 1999;58:120-8
22. Lass U, Numann A, von Eckardstein K, Kiwit J, Stockhammer F, Horaczek JA. et al. Clonal analysis in recurrent astrocytic, oligoastrocytic and oligodendroglial tumors implicates IDH1- mutation as common tumor initiating event. PloS one. 2012;7:e41298
23. Shibahara I, Sonoda Y, Kanamori M, Saito R, Yamashita Y, Kumabe T. et al. IDH1/2 gene status defines the prognosis and molecular profiles in patients with grade III gliomas. International journal of clinical oncology. 2012;17:551-61
24. van den Bent MJ, Dubbink HJ, Marie Y, Brandes AA, Taphoorn MJ, Wesseling P. et al. IDH1 and IDH2 mutations are prognostic but not predictive for outcome in anaplastic oligodendroglial tumors: a report of the European Organization for Research and Treatment of Cancer Brain Tumor Group. Clinical cancer research: an official journal of the American Association for Cancer Research. 2010;16:1597-604
25. Weller M, Felsberg J, Hartmann C, Berger H, Steinbach JP, Schramm J. et al. Molecular predictors of progression-free and overall survival in patients with newly diagnosed glioblastoma: a prospective translational study of the German Glioma Network. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 2009;27:5743-50
26. Dubbink HJ, Taal W, van Marion R, Kros JM, van Heuvel I, Bromberg JE. et al. IDH1 mutations in low-grade astrocytomas predict survival but not response to temozolomide. Neurology. 2009;73:1792-5
27. Iwadate Y, Matsutani T, Hirono S, Ikegami S, Shinozaki N, Saeki N. IDH1 mutation is prognostic for diffuse astrocytoma but not low-grade oligodendrogliomas in patients not treated with early radiotherapy. Journal of neuro-oncology. 2015;124:493-500
28. Waqar M, Hanif S, Brodbelt AR, Rathi N, Das K, Zakaria R. et al. Prognostic Factors in Lobar World Health Organization Grade II Astrocytomas. World neurosurgery. 2015;84:154-62
29. Jungk C, Scherer M, Mock A, Capper D, Radbruch A, von Deimling A. et al. Prognostic value of the extent of resection in supratentorial WHO grade II astrocytomas stratified for IDH1 mutation status: a single-center volumetric analysis. Journal of neuro-oncology. 2016;129:319-28
30. Ahmadi R, Stockhammer F, Becker N, Hohlen K, Misch M, Christians A. et al. No prognostic value of IDH1 mutations in a series of 100 WHO grade II astrocytomas. Journal of neuro-oncology. 2012;109:15-22
31. Juratli TA, Kirsch M, Robel K, Soucek S, Geiger K, von Kummer R. et al. IDH mutations as an early and consistent marker in low-grade astrocytomas WHO grade II and their consecutive secondary high-grade gliomas. Journal of neuro-oncology. 2012;108:403-10
Author contact
Corresponding authors: Qiang Gao (Department of Geriatrics, the Second Affiliated Hospital, Harbin Medical University, Baojian Road 157, Harbin, Heilongjiang 157011, P.R. China. Tel: +86-451-86605721; Fax: +86-451-86605721; E-mail: gaoqiangedu.cn) and Yuwen Song (Department of Neurosurgery, the Fourth Affiliated Hospital, Harbin Medical University, Yiyuan Street 37, Harbin, Heilongjiang 157001, P.R. China. Tel: +86-451-82576658; Fax: +86-451-82576658; E-mail: 350457918com)

 Global reach, higher impact
Global reach, higher impact