Impact Factor ISSN: 1837-9664
J Cancer 2017; 8(1):111-116. doi:10.7150/jca.17060 This issue Cite
Research Paper
Association between expression of Carboxypeptidase 4 and stem cell markers and their clinical significance in liver cancer development
1. State Key Laboratory of Molecular Oncology, National Cancer Center/Cancer Hospital, Chinese Academy of Medical Sciences, Peking Union Medical College, Beijing, People's Republic of China.
2. Department of Pharmaceutical Sciences, University of Michigan, Ann Arbor, MI 48109, USA.
3. Department of abdominal surgical oncology, Cancer Institute (Hospital), Chinese Academy of Medical Sciences, Peking Union Medical College, Beijing 100021, People's Republic of China.
4. Department of Hematology and Oncology, Children's Hospital of Soochow University, Suzhou, Jiangsu, 215005, P.R. China.
Received 2016-8-1; Accepted 2016-9-30; Published 2017-1-1
Abstract
The development of liver cancer would undergo a sequential progression from chronic inflammatory liver disease, cirrhosis to neoplasia. During these pathophysiological changes, abnormal liver microenvironment might induce the hepatocytes to die, abnormally proliferate and initiate cancer stem cells. Metallocarboxypeptidases (MCPs) involved in multiple biological functions including inflammation, fibrosis and stem cell niche formation. This study aimed to evaluate the expression of carboxypeptidase 4 (CPA4) in hepatitis, liver cirrhosis and liver cancer tissues, and revealed its clinical significance in liver cancer progression. We firstly found that the CPA4 levels in tissues were significantly higher in liver cancer patients than those in other three groups. Then, elevated levels of CPA4 was observed in 57/100 (57%) liver cancer samples, and significantly correlated with Grade and Stage. We also identified a significant positive correlation between aberrant elevation of CPA4 and overexpression of stem cell markers including CD90, AFP and CD34 with follow-up data (n=100). Further Kaplan-Meier analysis confirmed that high levels of CPA4 and CD90 were significant predictors of poor overall survival. Multivariate Cox regression model showed that CPA4 was an independent prognostic factor for patients with liver cancer. This study demonstrated for the first time that high CPA4 expression was closely correlated with hepatocarcinogenesis, and might be used as an independent poor prognostic factor in liver cancer.
Keywords: Carboxypeptidase 4, CD90, marker, cirrhosis, neoplasia, hepatocarcinogenesis, prognosis.
Introduction
Liver cancer is one of most common malignant tumors, and is among a leading cause of cancer-related death worldwide[1]. It has been reported that liver cancer would undergo a sequential steps from chronic inflammatory liver disease, cirrhosis to neoplasia[2]. During these process, the hepatocytes are prone to die and abnormally proliferate, and this dysregulated feedback process always contribute to the formation of pro-cancer microenvironment [3, 4].
CarboxypeptidaseA4 (CPA4) belongs to a family of Zn-containing metallocarboxypeptidases. It is an important regulator of inflammation, and is speculated to be involved in creating tumor microenvironment to facilitate cancer progression. It has been reported that CPA4 was a strong candidate gene for prostate cancer aggressiveness[5]. Our previous studies indicated that CPA4 was significantly elevated in pancreatic cancer tissues and serum samples, which was closely associated with tumor progression and poor prognosis[6]. However, the expression of CPA4 in liver cancer tissues and its clinical significance still remains unclear.
In this study, we aimed at evaluating the expression of CPA4 and related factors in normal, inflammatory liver disease, liver cirrhosis, and liver cancer tissues by IHC analysis, and investigated their association with clinicopathological variables and clinical outcomes.
Materials and methods
Tissue microarray and immunohistochemistry
Two commercial tissue microarrays were constructed by Shanghai Biochip Co. Ltd., as described previously[7]. Briefly, the tissue microarrays including 100 liver cancer patients and 90 adjacent normal tissues were prepared from archival formalin-fixed, paraffin embedded tissue blocks. Clinicopathological information (age, gender, and pathology, differentiation, and TNM stage) and Follow-up information were available. Another slides contained 40 healthy controls, 10 inflammatory liver disease, 40 liver cirrhosis, and 110 liver cancer patients with clinicopathological information (age, gender, and pathology, differentiation, and TNM stage). A representative tumor area was carefully selected from a H&E-stain section. For all the specimens, standard Avidin-biotin complex peroxidase immunohistochemical staining was performed. Briefly, after deparaffinizationin xylene and graded alcohols, heated antigen retrieval was done in citrate buffer (10mmol/L pH 6.0) by water-bath kettle heating for 30min. Endogenous peroxidase was blocked in 0.3% hydrogen peroxide for 10 min. Nonspecific binding was blocked by incubation in 10% normal animal serum for 10min. Sections were incubated at 4°C for 24 h with primary antibodies including polyclonal antibody against anti-CPA4 (HPA021030, Sigma-Aldrich), Anti-CD90 antibody (ab92574), Anti-CD34 antibody (ab81289, Abcam), Anti-AFP antibody (ab46799, Abcam) and anti-Ki67 (ab833, Abcam). Next, biotinylated secondary antibodies and horseradish peroxidase labeled Avidin were incubated with samples. Color was developed using the DAB method.
Immunostaining analysis
Slides were independently evaluated by 2 two pathologists who were blinded to patients' clinical data. The levels were scored by staining intensity and the percentage of immunoreactive cancer cells. Staining intensity was arbitrarily scored on a scale of four grades: 0 (no staining of cancer cells), 1 (weak staining), 2 (moderate staining), and 3 (strong staining), and the percentage of positive cells was scored as follows: 0 (0%), 1 (1% to 30%), 2 (31% to 50%), and 3 (>50%). The staining positivity was determined using the following formula: overall score=positive percentage score×intensity score. For CPA4, CD90 and AFP, a score of 0 to≤2 was defined as “0”, and >2 as “1”. For Ki67, a score of 0 to≤4 was defined as “0”, and >4 as “1”. For CD34, a score of 0 to≤1 was defined as “0”, and >1 as “1”.
Statistical Analysis
The SPSS 15 software package (SPSS, Inc., Chicago, IL) was used for statistical analysis. The association between the immunoreactive markers and clinicopathologic features was analyzed using χ2-test or two-sided t-test as appropriate. The survival rates were assessed by the Kaplan-Meier method and compared by the log-rank test. To investigate the prognostic significance, we included all variables having p < 0.05 in multivariate survival analysis using a Cox regression model. Spearman's rank correlation coefficient and Fisher's exact test were used to explore the association among CPA4 and other clinicalpathological features. All comparisons were two-tailed, and p < 0.05 was considered statistically significant.
Results
Expression of CPA4 in normal, chronic liver disease and hepatocellular carcinoma tissues by immunohistochemistry
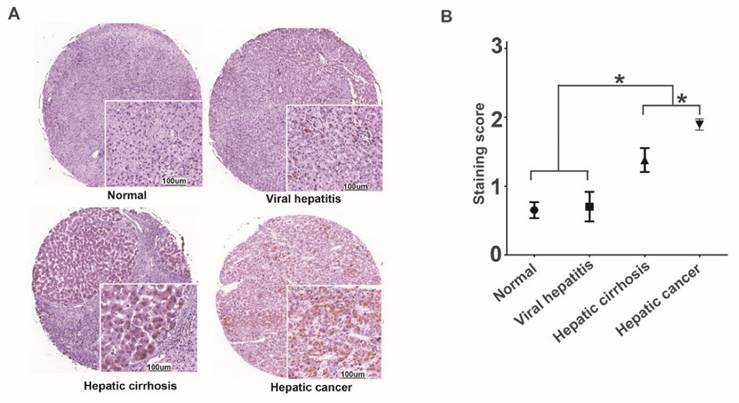
We performed immunohistochemistry to examine the CPA4 level, and found that the expression of CPA4 was gradually elevated, and the levels of CPA4 were significantly higher in liver cancer tissues than that in the cirrhosis, viral hepatitis and normal liver tissues (Figure 1). There was no statistical significance between healthy and hepatitis ones. These data suggested that CPA4 was closely associated with liver cancer development.
CPA4 expression in chronic inflammatory liver disease and liver cancer tissues were determined by immunochemistry. A. Positive expression of CPA4. B. Data represent the mean ± standard error of 4 groups.
High expression of CPA4 was associated with clinicopathological parameters
To further investigate the prognostic significance of CPA4, we evaluated the expression of CPA4 in liver cancer tissues with clinicopathological information (age, gender, grade, differentiation, and TNM stage) and Follow-up information. IHC results indicated that the incidence of CPA4 expression in liver cancer cases was 57% (57/100), and there was weak or no staining of CPA4 in the normal liver tissues. Further analysis also showed that over-expression of CPA4 was significantly associated with Grade (P=0.009), Depth of invasion (p=0.032) and clinical stage (P=0.013), while there was no significant correlation between CPA4 expression and tumor size, differentiation, age and gender. (Table 1)
Correlation between CPA4 expression and clinicopathological characteristics in 100 liver cancer cases.
| CPA4 | |||
|---|---|---|---|
| negative | positive | p-value | |
| Gender (Male: Female) | 37:6 | 52:5 | 0.412 |
| Age | 54.58±13.16 | 51.74±10.14 | 0.225 |
| Tumor size(cm) | 0.059 | ||
| < 10 cm3 | 34 | 35 | |
| > 10 cm3 | 9 | 22 | |
| Grade | 0.009 | ||
| 1 | 5 | 2 | |
| 2 | 29 | 27 | |
| 3 | 9 | 28 | |
| Depth of invasion | 0.032 | ||
| T1 | 12 | 5 | |
| T2 | 13 | 19 | |
| T3 | 18 | 29 | |
| T4 | 0 | 4 | |
| Lymph node involvement | 0.458 | ||
| N0 | 42 | 54 | |
| N1 | 1 | 3 | |
| Stage | 0.013 | ||
| Ⅰ | 25 | 19 | |
| Ⅱ+Ⅲ+Ⅳ | 18 | 38 | |
| M | 0.220 | ||
| M0 | 42 | 55 | |
| M1 | 0 | 2 | |
| AFP level | 0.001 | ||
| negative | 31 | 22 | |
| positive | 12 | 35 | |
| CD90 level | 0.002 | ||
| negative | 22 | 12 | |
| positive | 21 | 45 | |
| CD34 level | 0.044 | ||
| negative | 17 | 12 | |
| positive | 26 | 45 | |
| Ki67 level | 0.077 | ||
| negative | 28 | 27 | |
| positive | 15 | 30 | |
Correlation between CPA4 and CD90, AFP, Ki67 and CD34 in liver cancer tissues
It has been speculated that chronic liver disease contribute to the development of liver cancer stem cell by regulating cell proliferation, apoptosis and vasculature reconstruction. Then we detected the related markers including CD90, AFP, Ki67 and CD34 in the same tissue array by immunohistochemistry. CD90 was a possible candidate CSC marker in liver cancer, and we observed that positive staining of CD90 was detected in 66% (66/100) liver cancer samples. Overexpression of Ki67 was detected in 45% (45/100) liver cancer samples, and positive stain of AFP was found in 47% (47/100) specimens, and the positive rate of CD34 was 71% (71/100). Further analysis indicated that aberrant expression of CPA4 was significantly associated with CD90 (P=0.002), AFP (P=0.001) and CD34 (P=0.044). Spearman's rank correlation analysis revealed that aberrant expression of CPA4 was positively associated with AFP (P=0.001), CD34(P=0.044), and CD90 (P=0.001) in liver cancer (Table 2). Taken together, these observations demonstrated that CPA4 was closely associated with the existence of liver cancer stem cell.
Correlation between the Expression of CPA4 and AFP, CD34, CD90, Ki67.
| AFP | CD90 | CD34 | Ki67 | ||
|---|---|---|---|---|---|
| CPA4 | Correlation Coefficient | .332** | .315** | .202* | .177 |
| Sig. (2-tailed) | .001 | .001 | .044 | .079 | |
| N | 100 | 100 | 100 | 100 |
**. Correlation is significant at the 0.01 level (2-tailed).
*. Correlation is significant at the 0.05 level (2-tailed).
CPA4 expression was associated with poor overall survival in liver cancer patients
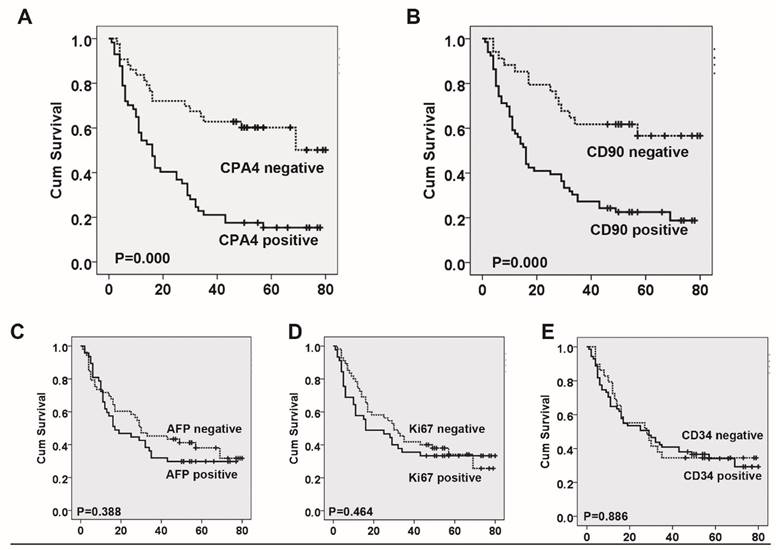
Based on the association between CPA4 and other cancer marker (AFP, CD90, ki67 and CD34), we analyzed their impact on liver cancer patient's survival. Using Kaplan-Meier survival analysis, we found that both CPA4 and CD90 overexpression were significantly correlated with poor overall survival of liver cancer patients, while high levels of Ki67, CD34 or AFP had no statistically significant association with poor survival (Figure 3). Next, Cox multivariate survival analysis revealed that CPA4 level (HR=2.381; 95% CI: 1.260-4.501; P=0.008) and CD90 (HR=1.968; 95% CI: 1.020-3.797; P=0.043) were statistically independent predictive factors of poorer prognosis for liver cancer (Table 3).
Survival curves for liver cancer using the Kaplan-Meier method and the log-rank test. A. Overall survival curves for patients with negative CPA4 expression (dotted line) and patients with positive CPA4 (full line); B. Overall survival curves for patients with negative CD90 expression (dotted line) and patients with positive CD90 (full line); C. Overall survival curves for patients with negative AFP expression (dotted line) and patients with positive AFP (full line). D. Overall survival curves for patients with negative Ki67 expression (dotted line) and patients with positive Ki67 (full line); E. Overall survival curves for patients with negative CD34 expression (dotted line) and patients with positive CD34 (full line).
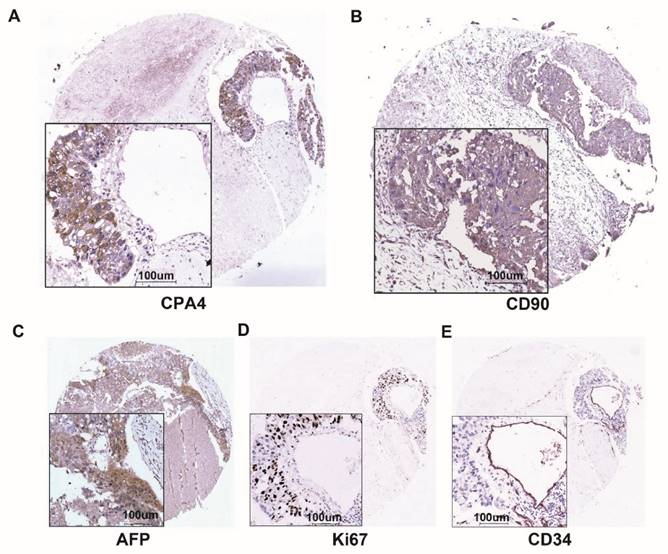
CPA4, CD90, AFP, Ki67 and CD34 expression in same liver cancer tissues were determined by immunochemistry. A. The expression of CPA4. B. The expression of CD90 C. The expression of AFP. D. The expression of Ki67. E. The expression of CD34.
Multivariate analysis by Cox Proportional Hazards Model in liver cancer.
| Characteristics | B | SE | Wald | df | Sig. | Exp(B) | 95.0% CI for Exp(B) | |
|---|---|---|---|---|---|---|---|---|
| Lower | Upper | |||||||
| CPA4 level | .868 | .325 | 7.132 | 1 | .008 | 2.381 | 1.260 | 4.501 |
| CD90 level | .677 | .335 | 4.077 | 1 | .043 | 1.968 | 1.020 | 3.797 |
| AFP level | -.186 | .265 | .493 | 1 | .483 | .830 | .494 | 1.396 |
| Ki67 level | .522 | .293 | 3.180 | 1 | .075 | 1.686 | .950 | 2.992 |
| Grade | -.239 | .262 | .829 | 1 | .363 | .788 | .471 | 1.317 |
| Stage | .358 | .501 | .511 | 1 | .475 | 1.430 | .536 | 3.814 |
| Depth of invasion | .470 | .290 | 2.629 | 1 | .105 | 1.600 | .906 | 2.826 |
Discussion
The development of liver cancer is a sequential process involving chronic hepatitis infection and cirrhosis[8]. In chronic inflammatory states, a series of factors including cytokines, reactive oxygen species (ROS) and angiogenesis factor could cause a permissive hepatic microenvironment for transition of damaged hepatocytes into liver cancer as well as regulation of CSCs [9]. Cancer stem cells (CSCs) has been regarded as the origin of many types of solid tumor including liver cancer, and CD90 is a potential marker for liver CSCs[10].
Carboxypeptidase A4 (CPA4) is a zinc-containing exopeptidase that catalyze the release of carboxy-terminal amino acids[11]. Several studies have indicated that CPA4 was closely associated with cell growth, differentiation and aggressiveness[12].
In breast cancer, O-glycosylated CPA4 could be secreted at a higher level by MCF-7 cells comparing to human mammary epithelial cells (HMEpC), and it may play key roles in breast cancer progression[13]. CPA4 was also found to be elevated in the squamous cell carcinoma of the head and neck tissues by loss of imprinting, and overexpression of CPA4 was closely associated with poor survival[14]. Our previous study indicated that CPA4 level was significantly elevated in pancreatic cancer tissues as well as serum samples, and was closely associated with tumor progression[6]. In NSCLC, CPA4 overexpression is associated with an unfavorable prognosis, and serum CPA4 level combining with serum CYFRA21-1 level could be used to aid early detection of NSCLC[15].
In this study, we firstly evaluated the CPA4 level in hepatitis, cirrhosis and liver cancer tissues. The results demonstrated that CPA4 was significantly higher in liver cirrhosis or liver cancer patients than in the healthy controls and hepatitis patients. These results revealed that CPA4 played key roles in hepatocarcinogenesis. To further study the prognostic value of CPA4 in liver cancer, we also collected the liver cancer tissue samples (n=100) with follow-up information. IHC results indicated that CPA4 was specifically elevated in 57% (57/100) cases, and high level of CPA4 was significantly associated with histological Grade and clinical stage. It has reported that CPA4 played key roles in establishing a micro-environment, which could increase the proportion of cancer stem cells and promote tumor angiogenesis. Hepatic cancer stem cell marker CD90 is a glycophosphatidylinositol (GPI)-anchored protein expressed in many normal cells including neurons, endothelial cells and fibroblast[10, 16]. As a specific biomarker for liver cancer, Alpha-fetoprotein (AFP) combing with EpCAM was used to define cancer stem cell[17]. CD34 has been widely used as a marker for hematopoietic stem cells and endothelial cells[18]. Ki67 is a proliferative and independent prognostic marker for cancer progression[19]. Based on the above information, we evaluated the expression of CD90, AFP, Ki67 and CD34 in the same tissue array by immunohistochemistry. we evaluated the expression of in the same tissue array by immunohistochemistry. The results showed that staining of CD90 was detected in 66% (66/100) liver cancer samples. The results revealed that aberrant expression of CPA4 was significantly associated with CD90 (P=0.002) in liver cancer. The results demonstrated that AFP, Ki67 and CD34 were positive in 47 (47%), 45 (45%) and 71(71%) in liver cancer samples, respectively. Further analysis also showed that aberrant expression of CPA4 was positively associated with AFP, Ki67 and CD90 in liver cancer tissues. Kaplan-Meier survival analysis also revealed a correlation between high levels of CPA4 or CD90 and shorter overall survival times. Multivariate Cox regression model demonstrated that CPA4 and CD90 level were statistically independent predictive factors of poor prognosis for liver cancer.
In conclusion, our studies for the first time suggested that CPA4 was highly expressed in liver cancer tissues. Overexpression of CPA4 can be used as an independent poor prognostic factor in liver cancer.
Acknowledgements
Supported by grant from National High-tech R&D Program of China for Young Scholars (No: 2014AA020537), Beijing Talents Fund (No: 2015000021223ZK23), Beijing Nova Program (No: Z1511000003150121), Beijing Gao Chuang Ji Hua (No.G02060050), PUMC Novel Star Program.
Conflicts of Interest
No potential conflicts of interest were disclosed.
References
1. Iakova P, Timchenko L, Timchenko NA. Intracellular signaling and hepatocellular carcinoma. Seminars in cancer biology. 2011;21:28-34
2. Ramakrishna G, Rastogi A, Trehanpati N, Sen B, Khosla R, Sarin SK. From Cirrhosis to Hepatocellular Carcinoma: New Molecular Insights on Inflammation and Cellular Senescence. Liver Cancer. 2013;2:367-83
3. He G, Karin M. NF-κB and STAT3 - key players in liver inflammation and cancer. Cell Research. 2011;21:159-68
4. Schwartsburd PM. Age-promoted creation of a pro-cancer microenvironment by inflammation: pathogenesis of dyscoordinated feedback control. Mechanisms of ageing and development. 2004;125:581-90
5. Ross PL, Cheng I, Liu X, Cicek MS, Carroll PR, Casey G. et al. Carboxypeptidase 4 gene variants and early-onset intermediate-to-high risk prostate cancer. BMC Cancer. 2009;9:69
6. Sun L, Burnett J, Guo C, Xie Y, Pan J, Yang Z. et al. CPA4 is a promising diagnostic serum biomarker for pancreatic cancer. American Journal of Cancer Research. 2016;6:91-6
7. Zhang Z, Wang J, Ji D, Wang C, Liu R, Wu Z. et al. Functional Genetic Approach Identifies MET, HER3, IGF1R, INSR Pathways as Determinants of Lapatinib Unresponsiveness in HER2-Positive Gastric Cancer. Clinical Cancer Research. 2014;20:4559-73
8. Wallace MC, Friedman SL. Hepatic fibrosis and the microenvironment: fertile soil for hepatocellular carcinoma development. Gene expression. 2014;16:77-84
9. Kocabayoglu P, Friedman SL. Cellular basis of hepatic fibrosis and its role in inflammation and cancer. Frontiers in bioscience. 2013;5:217-30
10. Yang ZF, Ho DW, Ng MN, Lau CK, Yu WC, Ngai P. et al. Significance of CD90+ Cancer Stem Cells in Human Liver Cancer. Cancer Cell. 2008;13:153-66
11. Tanco S, Zhang X, Morano C, Avilés FX, Lorenzo J, Fricker LD. Characterization of the Substrate Specificity of Human Carboxypeptidase A4 and Implications for a Role in Extracellular Peptide Processing. The Journal of Biological Chemistry. 2010;285:18385-96
12. Kayashima T, Yamasaki K, Yamada T, Sakai H, Miwa N, Ohta T. et al. The novel imprinted carboxypeptidase A4 gene ( CPA4) in the 7q32 imprinting domain. Hum Genet. 2003;112:220-6
13. Tan A-A, Phang W-M, Gopinath SCB, Hashim OH, Kiew LV, Chen Y. Revealing Glycoproteins in the Secretome of MCF-7 Human Breast Cancer Cells. BioMed Research International. 2015;2015:453289
14. Hsu CM, Lin PM, Lin HC, Lai CC, Yang CH, Lin SF. et al. Altered Expression of Imprinted Genes in Squamous Cell Carcinoma of the Head and Neck. Anticancer research. 2016;36:2251-8
15. Sun L, Wang Y, Yuan H, Burnett J, Pan J, Yang Z. et al. CPA4 is a Novel Diagnostic and Prognostic Marker for Human Non-Small-Cell Lung Cancer. Journal of Cancer. 2016;7:1197-204
16. Yang ZF, Ngai P, Ho DW, Yu WC, Ng MN, Lau CK. et al. Identification of local and circulating cancer stem cells in human liver cancer. Hepatology. 2008;47:919-28
17. Yamashita T, Ji J, Budhu A, Forgues M, Yang W, Wang H-Y. et al. EpCAM-positive hepatocellular carcinoma cells are tumor initiating cells with stem/progenitor cell features. Gastroenterology. 2009;136:1012-24
18. Nielsen JS, McNagny KM. Novel functions of the CD34 family. J Cell Sci. 2008;121:3683-92
19. Dudderidge TJ, Stoeber K, Loddo M, Atkinson G, Fanshawe T, Griffiths DF. et al. Mcm2, Geminin, and KI67 define proliferative state and are prognostic markers in renal cell carcinoma. Clin Cancer Res. 2005;11:2510-7
Author contact
Corresponding author: Lichao Sun, PhD. State Key Laboratory of Molecular Oncology, Cancer Hospital, Chinese Academy of Medical Sciences, Peking Union Medical College, Beijing, 100021, P. R. China. Prof_sunlichaocom. Duxin Sun, PhD. Department of Pharmaceutical Sciences, University of Michigan, Ann Arbor, MI 48109. duxinsedu.

 Global reach, higher impact
Global reach, higher impact