Impact Factor ISSN: 1837-9664
J Cancer 2016; 7(12):1740-1746. doi:10.7150/jca.15620 This issue Cite
Research Paper
MiR-193a-5p Targets the Coding Region of AP-2α mRNA and Induces Cisplatin Resistance in Bladder Cancers
1. College of Life Science, Hunan Normal University, Changsha 410081, Hunan, China;
2. Hunan Provincial People's Hospital, Changsha 410005, Hunan, China
3. Hunan Cancer Hospital, Changsha 410013, Hunan, China.
Received 2016-3-23; Accepted 2016-6-29; Published 2016-8-7
Abstract
Transcription factor AP-2 alpha (AP-2α or TFAP2A) is a newly identified prognostic marker of chemotherapy; its expression is positively correlated with chemosensitivity and survival of cancer patients. Using computational programs, we predicted that the coding region of AP-2α gene contains a potential miRNA response element (MRE) of miR-193a-5p, and the single nucleotide polymorphism (SNP) site (c.497A>G, rs111681798) resides within the predicted MRE. The results of luciferase assays and Western blot analysis demonstrated that miR-193a-5p negatively regulated the expression of AP-2α proteins, but have no influence on the mutant AP-2α (c.497A>G). Infection with lentiviral AP-2α gene or miR-193a-5p inhibitor in the bladder cancer cells decreased migration and cisplatin resistance, while knockdown of AP-2α gene or overexpression of miR-193a-5p in the urothelial cell line SV-HUC-1 increased migration and cisplatin resistances. We concluded that miR-193a-5p induced cisplatin resistance by repressing AP-2α expression in bladder cancer cells.
Keywords: miR-193a-5p, AP-2α, cisplatin resistance, bladder cancer, single nucleotide polymorphism.
Introduction
Bladder cancer is the fourth most common cancer in men, and the patients with muscle-invasive and metastatic bladder cancer are at high risk of morbidity and mortality [1]. Current treatment strategy for bladder cancer is a combination of surgery with chemotherapy and/or radiotherapy. Cisplatin-based combination chemotherapy is the gold standard of treatment for metastatic bladder cancer [2].
However, only a subset of patients is sensitive to cisplatin-based therapy. Galsky et al's investigated the effectiveness of neoadjuvant therapy for muscle-invasive bladder cancer and found that pathologic complete response rate was about 30% in the patients who received cisplatin-based combination chemotherapy [3]. Therefore, it is necessary to identify reliable markers for assessing chemotherapeutic response. The molecular mechanism of cisplatin therapy is linking to its ability to cause DNA damage, and subsequently induce apoptosis in cancer cells [4]. Therefore, most apoptosis-related proteins are found to affect the sensitivity to cisplatin. Usually, anti-apoptotic proteins (i.e., Bcl-2) are linked to poorer survival in patients with invasive bladder cancer treated with chemotherapy, while pro-apoptotic proteins are the predictive markers for a good response and survival. Up to now, many proteins have been identified to be associated with the response to cisplatin based chemotherapy in bladder (reviewed in [5]). Among these, transcription factor AP-2 alpha (AP-2α or TFAP2A) is a newly identified prognostic marker of chemotherapy [5]. It is a member of the transcription factor AP-2 family, functions as a tumor suppressor by regulating the expression of various cancer-related genes, such as p21 [6], MMP9 [7], E-cadherin [7], Bcl-2 [8] and Bax [8]. AP-2α expression was positively associated with chemosensitivity in breast [9], bladder [10], endometrium [11] and pancreas [12] cancers. In cancer cells, AP-2α is usually downregulated by promoter hypermethylation [13, 14] or miRNAs [11], leading to chemoresistance [11].
miRNAs are a class of short single-stranded RNAs, which suppress gene expression by binding to the 3′-unstranslated region (UTR) of the target mRNA to inhibit translation and/or promote mRNA degradation. Recently, an increasing number of studies demonstrated that miRNAs also target the 5′-UTR [15] and coding region [16, 17] of mRNAs. miRNAs have been involved in various cell processes, including chemoresistance. For example, miR-203 enhances chemosensitivity to cisplatin in bladder cancer by targeting Bcl-w and Survivin [18]. Xu et al [19] comprehensively investigated the relationship of miRNA expression with the progression and prognosis of muscle-invasive bladder cancer, and found that the patients with higher let-7c, mir-125b-1, mir-193a, and mir-99a levels exhibited shorter survival intervals. Our previous study demonstrated that miR-200b/200c/429 induced cisplatin resistance by repressing AP-2α expression in endometrial cancer cells.
In present study, we found that miR-193a-5p inhibits the expression of AP-2α by binding its coding region, and then induces cisplatin resistance in bladder cancer cells.
Materials and methods
Computational prediction of miRNA targets
The miRNAs targeting the 3′-UTR of AP-2α gene were predicted as described previously [11]. RegRNA 2.0[20] (http://regrna2.mbc.nctu.edu.tw) and DIANA-microT-CDS [21] (http://diana.cslab.ece.ntua.gr/micro-CDS) were used to search for the miRNAs targeting the CDS of AP-2α gene.
Plasmids
The dual-luciferase miRNA target expression vector pmirGLO was purchased from Promega Corporation (Madison, WI, USA). The vector pmirCDS was modified from pmirGLO using seamless cloning kit (Sinobio, Shanghai, China). The difference between them is that the multiple cloning site (MCS) of pmirGLO is located downstream of firefly luciferase gene, but the MCS of pmirCDS is located within the 3′-end of the coding sequence (CDS) of firefly luciferase gene. The sequence cloned into pmirCDS will be expressed as fusions to the C-terminus of firefly luciferase. The oligonucleotide pairs that contain the binding site of miR-193a-5p were synthesized, annealed and ligated into the vector pmirGLO and pmirCDS as the method described previously [11], these constructs are denoted UTR-MRE, mUTR-MRE, CDS-MRE, mCDS-MRE, respectively. Overlapping PCR [22] was performed using Myc-tagged AP-2α expression construct Myc-AP-2α as a template, to generate two Myc-tagged AP-2α mutant (Myc-AP2α497G, with a substitution of c.497A>G; Myc-AP2α498T, with a substitution of c.498C>T).
Cell culture and transfection
Human embryonic kidney cell line 293T, human urothelial cell line SV-HUC-1 and bladder cancer line UM-UC-3 (from ATCC, Manassas, VA, USA) were cultured in MEM or F12K media that was supplemented with glutamine, antibiotics and 10% fetal bovine serum at 37 ℃ and 5% CO2. Transfection was performed using Lipofectamine 2000 according to the manufacturer's instructions (Invitrogen, Carlsbad, CA, USA).
Luciferase assay
The dual-luciferase reporter plasmids were co-transfected with miRNA mimics (GenePharma, Shanghai, China) into 293T cells. At 24 h post-transfection, cells were harvested and assayed for luciferase activity using the Dual-Glo Luciferase Assay System (Promega) according to the manufacturer's instructions. The firefly luciferase activities were normalized to Renilla luciferase activity. For each transfection, the luciferase activity was averaged from three replicates.
RNA preparation, cDNA synthesis and real-time PCR
Total RNA was extracted from various cell lines using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). The expression of miRNA-193a-5p was determined using All-in-One miRNA qRT-PCR Detection Kit (GeneCopoeia, Guanzhou, China). Briefly, Poly-A polymerase is used to add poly-A tails to the 3′ end of miRNAs, and then miRNAs were reverse transcribed into cDNA using M-MLV RTase and a oligodT adaptor primer. SYBR green-based realtime PCR was performed using miRNA-specific primer and universal adaptor primer on an ABI 7900 thermocycler (Thermo Fisher Scientific). The U6 gene was used as a normalization control. The above primers and enzymes were purchased from GeneCopoeia.
Lentivirus and transduction
All the lentiviruses expressing AP-2α gene, AP-2α shRNA (target sequence: 5′-TTTCTCAACCGACAACATT-3′), has-miR-193a-5p precursor and has-miR-193a-5p inhibitor and their empty vector lentivirus (GV341, GV248, GV309, GV280, respectively) were purchased from GeneChem (Shanghai, China). Lentiviruses were transduced into cells at a multiplicity of infection (MOI) of 10 in the presence of polybrene.
Western blot
Cells were lysed in RIPA buffer [50 mM Tris-HCl (pH 7.2), 150 mM NaCl, 1% (v/v) Triton X-100, 1% (w/v) sodium deoxycholate, 0.1% (w/v) SDS] with protease inhibitors. Proteins were separated on 10% SDS-polyacrylamide gel and transferred to PVDF membranes. A prestained protein ladder was used as a molecular marker. A single membrane was cut into two parts at the 45-kD band and incubated with anti-AP-2α (Abcam, Cambridge, UK) and anti-GAPDH (or anti-β-Actin) (Santa Cruz, CA, USA) primary antibodies, respectively. The protein was detected using an HRP-conjugated secondary antibody and a ChemiLucent ECL Detection System (Millipore, Billerica, MA, USA).
Cisplatin treatment and MTT assay
At 72h after transduction, cells were seeded on 96-well plates and treated with different concentration of cisplatin (Selleck Chemicals, Houston, TX, USA) for the indicated time. Then, the cells were cultured on media with 3-(4, 5-methylthiazol-2-yl)-2, 5-diphenyl-tetrazolium bromide (MTT) (0.5mg/mL) for 4 h. Media were removed and 100 μL dimethyl sulfoxide (DMSO) was added to solubilize formazon crystals. Absorbance was measured at 490 nm in a Biotek Synergy 2 plate reader (Biotek; Winooski, VT, USA). Based on the result of MTT assay, the IC50 (half maximal inhibitory concentration) of cisplatin was calculated using Microsoft Excel.
Wound healing assay
Cells were cultured on a 24-well plate until 100% confluence and then wounded with a 100 μL pipette tip. Migration photos were captured at 0, 24, and 48h hours after scratching. All experiments were repeated in triplicate independently.
Statistical analysis
Microsoft Excel was used for statistical analysis. Student's t-test was performed to evaluate the significance of difference between samples.
Results
miR-193a-5p targets the coding region of AP-2α mRNA and rs111681798 affects the binding of miR-193a-5p
We used several computational programs to predict the miRNAs regulating AP-2α gene, and found a binding site (miRNA response element, MRE) of miR-193a-5p in the coding region of AP-2α gene (Fig. 1A). Then, by searching the Ensembl database, we identified an SNP(c.497A>G, rs111681798) within the MRE. To determine whether the predicted MRE can be recognized by miR-193a-5p, we generated a Myc-tagged wild-type and two mutants (c.497A>G and c.498C>T) AP-2α expression plasmids, and co-transfected with miR-193a-5p mimics into 293T cells. At 24 h post-transfection, cell lysates were prepared and subjected to Western blotting assay using anti-Myc antibody. The result showed that the wild-type AP-2α expression can be inhibited by miR-193a-5p, but 497A>G or 498C>T substation can disrupt the suppressive role of miR-193a-5p on AP-2α expression (Fig. 1B).
The miR-193a-5p targets the coding region (CDS) of AP-2α mRNA. (A) Diagram of the binding between miR-193a-5p seed sequence and the AP-2α CDS. (B) Myc-tagged wild-type or mutant AP-2α expression plasmids were co-transfected with miR-193a-5p mimics into 293T cells. At 24 h post-transfection, cell lysates were prepared and subjected to Western blotting assay using anti-Myc antibody. (C) Each luciferase construct was co-transfected with miR-193a-5p mimics into 293T cells. At 24 h post-transfection, the luciferase activity was examined. The firefly luciferase activity was normalized to Renilla luciferase activity. Data are shown as the mean+SD of three independent experiments. *, p<0.05; **, p<0.01.
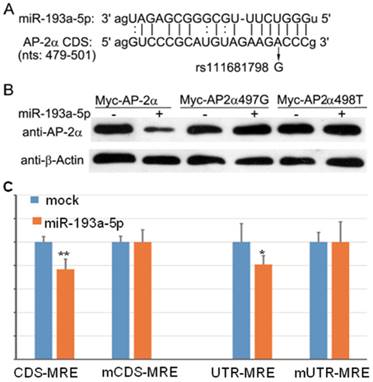
To confirm the above result, we inserted wild-type and mutant MRE in-frame with the firefly luciferase gene of reporter vector pmirCDS, respectively, leading to that MRE can express as a fusion to the C-terminus of firefly luciferase. These constructs (CDS-MRE and mCDS-MRE) were transfected into 293T cell with or without miR-193a-5p mimics. At 36h post-transfection, luciferase activities were determined. The result (Fig. 1C) showed that miR-193a-5p suppressed the firefly luciferase activities of CDS-MRE, but had no influence on mCDS-MRE, indicating that the predicted MRE is really recognized by miR-193a-5p.
Then, we cloned wild-type and mutant MRE into 3′-UTR of firefly luciferase and transfected into 293T cell with or without miR-193a-5p mimics. As shown in Figure 1C, luciferase activity of UTR-MRE can be repressed by miR-193a-5p as CDS-MRE, indicating that the MRE in the 3′-UTR functions as efficiently as it in the CDS.
miR-193a-5p is upregulated in bladder cancer cells and inhibits the expression of AP-2α
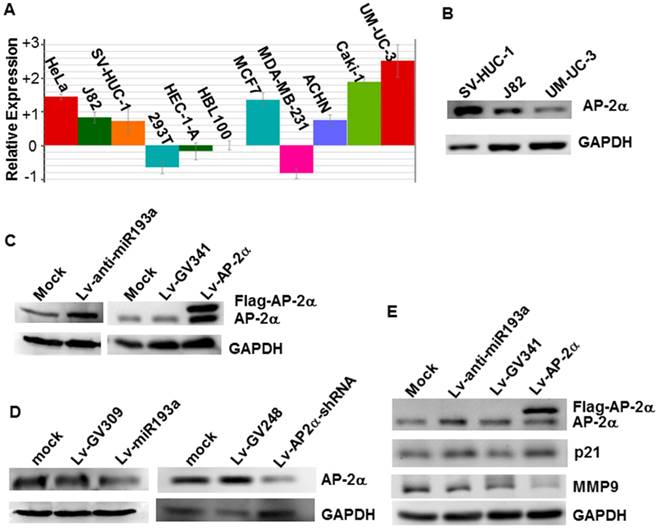
We investigated the miR-193a-5p expression in several cell lines by realtime PCR, and found that the miR-193a-5p expression in the bladder cancer line UM-UC-3 is the highest in the cells studied, far higher than that in the normal human urothelial cells SV-HUC-1 (Fig.2A). Western blotting showed that AP-2α expression in the SV-HUC-1 is much stronger than that in the UM-UC-3 (Fig.2B). Then, to investigate whether endogenous AP-2α expression was inhibited by miR-193a-5p, we transduced the lentiviral miR-193a-5p mimics and inhibitors into SV-HUC-1 and UM-UC-3, respectively, and then examined the level of AP-2α mRNA and protein. As shown in Figure 2C & D, overexpression of miR-193a-5p suppressed the AP-2α protein expression, while knock-down of miR-193a-5p increased the level of AP-2α protein. However, miR-193a-5p had no significant influence on AP-2α mRNA expression (data not shown). These results suggested that AP-2α expression is negatively regulated by miR-193a-5p.
Influence of miR-193a-5p on AP-2α expression. (A)Quantitative analysis of miRNA-193a-5p expression on various cell lines by realtime PCR. (B) Expression of AP-2α protein in three cell lines using Western blotting. (C)(D) Expression of AP-2α protein in UM-UC-3 cells (C) and SV-HUC-1 cells (D). (E) Influence of miR-193a-5p on AP-2α downstream targets. All the cells were infected with lentivirus, and harvested for Western blot at 72 h after infection.
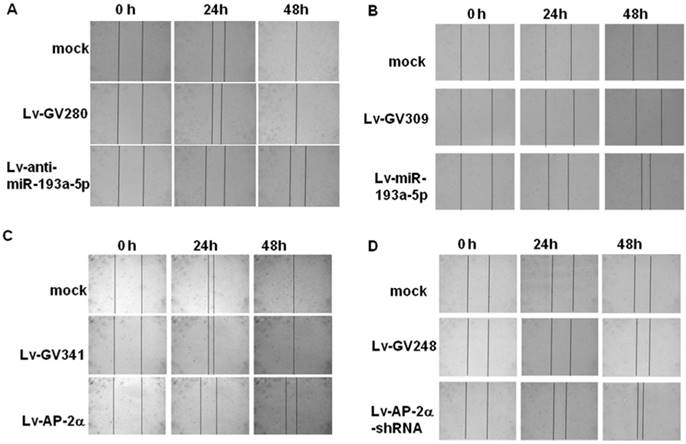
Representative pictures of wound healing dynamics. UM-UC-3 (A, C) and SV-HUC-1 cells (B, D) infected with the indicated lentivirus were cultured on a 24-well plate until 100% confluence and then wounded with a 100 μL pipette tip. Migration photos were captured at 0, 24, and 48h hours after scratching.

It has been shown that AP-2α up-regulated p21 expression [6] and down-regulated MMP9 expression [6]. To further confirm the influence of miR-193a-5p on AP-2α protein expression, we examined expression of p21 and MMP9 and found that knock-down of miR-193a-5p or over-expression of AP-2α increased the level of p21 protein, but suppressed the expression of MMP9 (Fig. 2E).
miR-193a-5p stimulates, but AP-2α suppresses cell migration
The wound healing assay (Fig. 3) showed that the SV-HUC-1 cells infected with lentiviral miR-193a-5p or AP-2α shRNA obtained faster closure of the scratched wound compared with control cells, while UM-UC-3 cells infected with lentiviral miR-193a-5p inhibitor or AP-2α gene exhibited lower wound closure rate than the cells infected empty vector. These result suggested overexpression of miR-193a-5p or inhibition of AP-2α gene increases cell migration.
miR-193a-5p decreases, but AP-2α increases cisplatin sensitivity
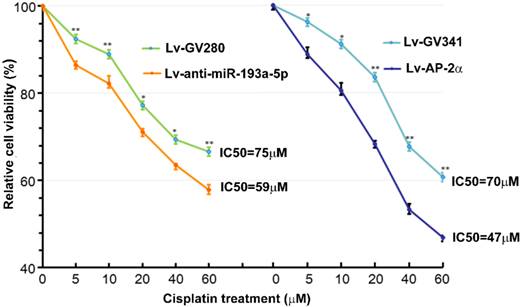
Our previous study showed that AP-2α expression was positively correlated with cisplatin sensitivity [11]. Therefore, we investigated effects of AP-2α and miR-193a-5p on cisplatin sensitivity of bladder cancers using MTT assay. The bladder cancer line UM-UC-3 cells were infected with lentiviral miR-193a-5p inhibitor or AP-2α gene, and then treated with different concentration of cisplatin for 36 h. As shown in Figure 4, the IC50s of cells transfected with lentiviral miR-193a-5p inhibitor or AP-2α gene is significantly lower than those of their control cells. This result suggests that knock down of miR-193a-5p or overexpression of AP-2α gene can increase cisplatin sensitivity of bladder cancer cells.
Discussion
The pre-miR-193a can generate two mature miRNAs, miR-193a-5p and miR-193a-3p. The role of miR-193a-3p has been well studied. It has been reported that miR-193a-3p inhibits chemoresistance in bladder cancers by targeting PSEN1 [23] or ING5 [24], while promotes chemoresistance by inhibiting HOXC9 [25] or LOXL4 [26]. But, the targets of miR-193a-5p remain to be unknown. In present study, we found that miR-193a-5p target the coding sequence of AP-2α mRNA, and suppresses its expression. To our knowledge, this is the first report about the target of miR-193a-5p.
Influence of miR-193a-5p and AP-2α on cisplatin sensitivity. UM-UC-3 cells were infected with lentiviral miR-193a-5p inhibitor or AP-2α gene. At 72h after transduction, cells were seeded on 96-well plates and treated with different concentration of cisplatin for 36 h. The cell viability was determined by MTT assay.
Additionally, we found that inhibition of miR-193a-5p or overexpression of AP-2α gene decreased cisplatin resistance in bladder cancer cell line UM-UC-3. These results are in accordance with previous studies [10, 19]. Xu et al comprehensively analyzed the microRNA (miRNA) expression profiles in muscle invasive bladder cancers, and found that miR-193a expression is negatively correlated with patient survival [19]. Conversely, AP-2α has been reported to be as a predictive marker for good response and survival after cisplatin-containing chemotherapy in several cancers [9-12]. Nordentoft et al demonstrated that high levels of AP-2α protein in bladder cancers were associated with good response to cisplatin, and AP-2α knockdown increased the proliferation of the SW780 bladder cells and reduced cisplatin and gemcitabine sensitivity [10]. These results indicate that miR-193a-5p induces cisplatin resistance by inhibiting AP-2α expression.
In conclusion, we found that miR-193a-5p is upregulated in bladder cancer cells, and inhibits the expression of AP-2α by binding its coding region. AP-2α gene down-regulates, but miR-193a-5p up-regulates cisplatin resistance in bladder cancer cells. This finding should be valuable toward understanding the mechanism of chemoresistance of bladder cancers.
Acknowledgements
This study was supported by National Natural Science Foundation of China (grant no. 81272318), Cooperative Innovation Center of Engineering and New Products for Developmental Biology of Hunan Province (grant no. 20134486), Health and Family Planning Commission of Hunan Province (grant no. B2014-090) and Hunan Provincial Science and Technology Department (grant no. 2015SK2044).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Solomon JP, Hansel DE. Morphologic and Molecular Characteristics of Bladder Cancer. Surg Pathol Clin. 2015;8:663-76
2. Sternberg CN, Bellmunt J, Sonpavde G, Siefker-Radtke AO, Stadler WM, Bajorin DF. et al. ICUD-EAU International Consultation on Bladder Cancer 2012: Chemotherapy for urothelial carcinoma-neoadjuvant and adjuvant settings. Eur Urol. 2013;63:58-66
3. Galsky MD, Pal SK, Chowdhury S, Harshman LC, Crabb SJ, Wong YN. et al. Comparative effectiveness of gemcitabine plus cisplatin versus methotrexate, vinblastine, doxorubicin, plus cisplatin as neoadjuvant therapy for muscle-invasive bladder cancer. Cancer. 2015;121:2586-93
4. Dasari S, Tchounwou PB. Cisplatin in cancer therapy: molecular mechanisms of action. Eur J Pharmacol. 2014;740:364-78
5. Ecke TH. Biomarker in Cisplatin-Based Chemotherapy for Urinary Bladder Cancer. Adv Exp Med Biol. 2015;867:293-316
6. Scibetta AG, Wong P-P, Chan KV, Canosa M, Hurst HC. Dual association by TFAP2A during activation of the p21cip/CDKN1A promoter. Cell Cycle. 2014;9:4525-32
7. Zhang Z, Zhang L, Jia L, Cui S, Shi Y, Chang A. et al. AP-2alpha suppresses invasion in BeWo cells by repression of matrix metalloproteinase-2 and -9 and up-regulation of E-cadherin. Mol Cell Biochem. 2013;381:31-9
8. Zhang L, Jia L, Cui S, Shi Y, Chang A, Wang P. et al. AP-2alpha-dependent regulation of Bcl-2/Bax expression affects apoptosis in the trophoblast. J Mol Histol. 2012;43:681-9
9. Wajapeyee N, Raut CG, Somasundaram K. Activator protein 2alpha status determines the chemosensitivity of cancer cells: implications in cancer chemotherapy. Cancer Res. 2005;65:8628-34
10. Nordentoft I, Dyrskjot L, Bodker JS, Wild PJ, Hartmann A, Bertz S. et al. Increased expression of transcription factor TFAP2alpha correlates with chemosensitivity in advanced bladder cancer. BMC Cancer. 2011;11:135
11. Wu Y, Xiao Y, Ding X, Zhuo Y, Ren P, Zhou C. et al. A miR-200b/200c/429-binding site polymorphism in the 3' untranslated region of the AP-2alpha gene is associated with cisplatin resistance. PLoS One. 2011;6:e29043
12. Jonckheere N, Fauquette V, Stechly L, Saint-Laurent N, Aubert S, Susini C. et al. Tumour growth and resistance to gemcitabine of pancreatic cancer cells are decreased by AP-2alpha overexpression. Br J Cancer. 2009;101:637-44
13. Douglas DB, Akiyama Y, Carraway H, Belinsky SA, Esteller M, Gabrielson E. et al. Hypermethylation of a small CpGuanine-rich region correlates with loss of activator protein-2alpha expression during progression of breast cancer. Cancer Res. 2004;64:1611-20
14. Hallberg AR, Vorrink SU, Hudachek DR, Cramer-Morales K, Milhem MM, Cornell RA. et al. Aberrant CpG methylation of the TFAP2A gene constitutes a mechanism for loss of TFAP2A expression in human metastatic melanoma. Epigenetics. 2014;9:1641-7
15. Orom UA, Nielsen FC, Lund AH. MicroRNA-10a binds the 5'UTR of ribosomal protein mRNAs and enhances their translation. Mol Cell. 2008;30:460-71
16. Fang Z, Rajewsky N. The impact of miRNA target sites in coding sequences and in 3'UTRs. PLoS One. 2011;6:e18067
17. Hausser J, Syed AP, Bilen B, Zavolan M. Analysis of CDS-located miRNA target sites suggests that they can effectively inhibit translation. Genome Res. 2013;23:604-15
18. Zhang X, Zhang Y, Liu X, Fang A, Li P, Li Z. et al. MicroRNA-203 Is a Prognostic Indicator in Bladder Cancer and Enhances Chemosensitivity to Cisplatin via Apoptosis by Targeting Bcl-w and Survivin. PLoS One. 2015;10:e0143441
19. Xu Z, Yu YQ, Ge YZ, Zhu JG, Zhu M, Zhao YC. et al. MicroRNA expression profiles in muscle-invasive bladder cancer: identification of a four-microRNA signature associated with patient survival. Tumour Biol. 2015;36:8159-66
20. Chang TH, Huang HY, Hsu JB, Weng SL, Horng JT, Huang HD. An enhanced computational platform for investigating the roles of regulatory RNA and for identifying functional RNA motifs. BMC Bioinformatics. 2013;14(Suppl 2):S4
21. Reczko M, Maragkakis M, Alexiou P, Grosse I, Hatzigeorgiou AG. Functional microRNA targets in protein coding sequences. Bioinformatics. 2012;28:771-6
22. Zhou J, Fan C, Zhong Y, Liu Y, Liu M, Zhou A. et al. Genomic organization, promoter characterization and roles of Sp1 and AP-2 in the basal transcription of mouse PDIP1 gene. FEBS Lett. 2005;579:1715-22
23. Meng F, Qian L, Lv L, Ding B, Zhou G, Cheng X. et al. miR-193a-3p regulation of chemoradiation resistance in oesophageal cancer cells via the PSEN1 gene. Gene. 2016;579:139-45
24. Li Y, Deng H, Lv L, Zhang C, Qian L, Xiao J. et al. The miR-193a-3p-regulated ING5 gene activates the DNA damage response pathway and inhibits multi-chemoresistance in bladder cancer. Oncotarget. 2015;6:10195-206
25. Lv L, Li Y, Deng H, Zhang C, Pu Y, Qian L. et al. MiR-193a-3p promotes the multi-chemoresistance of bladder cancer by targeting the HOXC9 gene. Cancer Lett. 2015;357:105-13
26. Deng H, Lv L, Li Y, Zhang C, Meng F, Pu Y. et al. miR-193a-3p regulates the multi-drug resistance of bladder cancer by targeting the LOXL4 gene and the oxidative stress pathway. Mol Cancer. 2014;13:234
Author contact
Corresponding author: Jianlin Zhou, PhD, Professor, College of Life Science, Hunan Normal University, Changsha 410081, Hunan, China. Phone/Fax: +86-731-88872792, e-mail: jlzhouedu.cn.

 Global reach, higher impact
Global reach, higher impact