Impact Factor ISSN: 1837-9664
J Cancer 2016; 7(12):1610-1620. doi:10.7150/jca.14923 This issue Cite
Research Paper
MicroRNA Expression Profile on Solid Subtype of Invasive Lung Adenocarcinoma Reveals a Panel of Four miRNAs to Be Associated with Poor Prognosis in Chinese Patients
1. Biorepository, State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, Chengdu, 610041, China
2. Department of Pathology, West China Hospital, Sichuan University, Chengdu, 610041, China
3. Department of Thoracic Oncology and Cancer Center, West China Hospital, Sichuan University, Chengdu, 610041, China
4. Institute for Genomic Medicine, University of California, San Diego, San Diego, CA 92093, USA
5. Molecular Medicine Center, West China Hospital, Sichuan University, Chengdu, 610041, China
6. Department of Ophthalmology, University of California, San Diego, San Diego, CA 92093, USA.
*These two authors contribute to this article equally.
Received 2016-1-8; Accepted 2016-5-23; Published 2016-7-18
Abstract
According to the reclassification of lung adenocarcinoma (LAC) proposed in 2011, solid predominant lung adenocarcinoma (SPA) has been associated with poor outcomes for LAC patients. However, the prognostic value of the presence of solid subtype remains unclear. Besides, there is little data about the roles of microRNA (miRNA) in solid subtype of LAC. In this study, 243 LAC patients were classified into solid subtype positive and negative groups (S+ LAC, n=134 and S- LAC, n=109) according to whether the solid subtype was more than 5% of the tumor component or not. We analyzed the relationship between solid subtype and patients' outcome by univariate and multivariate analyses. Solid subtype was proved to be significantly associated with the 5-year overall survival and played as an independent prognostic factor for stage I-III invasive LAC patients. Then miRNA microarray was used to identify differentially expressed miRNAs in solid subtype, resulting in 31 differential miRNAs. Quantitative reverse transcription-PCR (QRT-PCR) was used to validate 4 key miRNAs (miR-133b, miR-155-5p, miR-124-3p, miR-145-5p). Further, CCK-8 and transwell assays were performed to validate the impact of one dysregulated miRNA (miR-133b) on LAC cell function. Interestingly, while miR-133b could significantly inhibit the proliferation of A549 and SPC-A1, it showed no effect on the migration or invasion of LAC cell lines. These results suggest that solid subtype can exert independent prognostic impact on LAC patients, and 4 important dysregulated miRNAs in solid subtype of LAC may be involved in the malignancy of S+LAC, thus may further have clinical perspective for S+ LAC in the future.
Keywords: Lung adenocarcinoma, stage I-III, Solid subtype, Prognosis, microRNA (miRNA).
Introduction
Lung cancer is a deadly disease with the highest incidence and mortality among all cancers worldwide, and lung adenocarcinoma (LAC) is the most common histologic type [1, 2]. Five main subtypes of LAC are defined according to reclassification system proposed by the International Association for the Study of Lung Cancer (IASLC)/American Thoracic Society (ATS)/European Respiratory Society (ERS) in 2011 [3]. Of these subtypes, solid predominant adenocarcinoma (SPA) and micropapillary predominant adenocarcinoma (MPA) have been reported to be associated with poor prognosis [4-9]. However, no consensus exists regarding the true impact of these subtypes [10-13]. While most reports focus on the overall prognosis of LAC, we decide to investigate the relationship between the presence of the solid subtype and patient survival as it remains controversial whether the solid subtype component has independent prognostic value for patients with LAC [7, 11, 14].
The molecular heterogeneity of LAC is well documented, with a growing number of reports addressing the intra-tumor heterogeneity of LAC driver mutations among different predominant subtypes, including mutations in KRAS, EGFR, HER2, ALK, ROS1, and RET [5, 15-19]. However, little is known to date about the role of microRNAs (miRNAs) in invasive LAC. Significant effort is required to elucidate the possible role of miRNAs in the subtypes of LAC and their value for risk stratification and prognosis [3]. MiRNAs are a group of small (22-25bp), endogenous, and conserved noncoding RNAs that regulate gene transcription through binding to the 3'UTR or 5'UTR of their target mRNAs [20, 21] and they have been demonstrated to be involved in various human cancers [22-24]. Notably, differential miRNA expression has been reported to be significantly associated with mutational status [25], subtypes of non-small cell lung cancer (NSCLC) [26, 27], and clinicopathologic features [26], thus has the potential to be of great value in the diagnosis and prognosis of lung cancer [28, 29].
In this study, we analyzed the independent prognostic impact of the presence of solid subtype on the 5-year overall survival (5-year OS) of stage I-III Han Chinese patients with LAC, as well as the correlation between the solid subtype and clinicopathologic features such as smoking status and stage. Next, miRNA microarray and QRT-PCR studies were performed on solid subtype tumors and paired normal tissue samples to identify 4 significantly dysregulated miRNAs that may play key roles in the pathogenesis of solid subtype. The contribution to the malignant phenotype of one dysregulated miRNA, miR-133b, was validated in LAC cell lines using cellular assays. As far as we know, this is the first study to identify dysregulated miRNAs in solid subtype of invasive LAC.
Materials and Methods
Patients
We retrospectively selected 255 primary LAC patients (stage I-III) who underwent surgical resection from 2009 to 2012 at West China Hospital, Sichuan University. All patients had no chemo- or radiotherapy before surgery. Corresponding clinicopathologic features were collected including sex, age at diagnosis, smoking status, clinical stage (according to the 7th edition TNM classification by IASLC), maximum tumor diameter (MTD), bronchial stump (positive or negative).
Histologic Evaluation
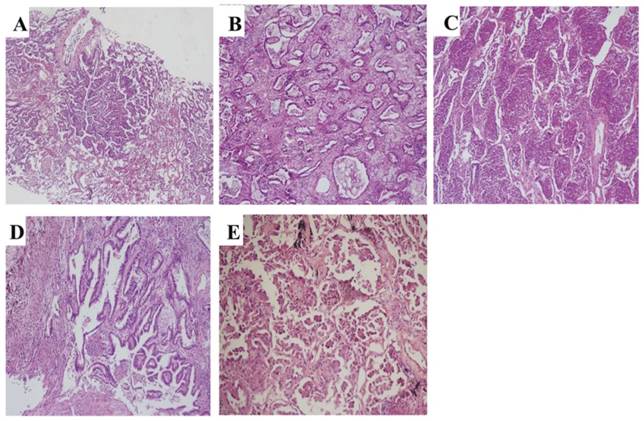
All cases were histologically reclassified based on the criteria established by IASLC/ATS/ERS in 2011. The percentage of each subtype (lepidic, acinar, solid, papillary, micropapillary, and variants, Figure 1) was semi-quantitatively assessed and recorded in 5% increments. The predominant subtype was defined as the subtype with the largest percentage. 243 cases were classified into solid subtype positive and negative LAC (S+ LAC, n=134 and S- LAC, n=109) according to whether the solid subtype was more than 5% of the tumor component or not.
Two pathologists (W. Wy and J. Ll) reviewed the hematoxylin and eosin (H&E)-stained pathologic slides for each case individually and were blinded to the patients' outcome and clinical stage. For controversial cases, the reviewers used a double-headed microscope to reach a consensus.
Tissues and cells
To get relatively pure sample of solid subtype, we collected a total of 279 primary LACs tissues that were stored in liquid nitrogen by Biobank of West China Hospital. All samples were used to produce formalin-fixed paran-embedded (FFPE) samples for further histological evaluation. 26 LAC FFPE samples with the pure solid subtype were isolated together with paired adjacent normal FFPE samples. Ethics approval was obtained from Ethics Committee at West China Hospital of Sichuan University for the use of these samples.
Human LAC cell lines A549 and SPC-A1 were both obtained from American type culture collection (ATCC, USA), grown in RPMI 1640 medium (Hyclone, USA) with 10% FBS (BI, Israel), and incubated at 5% CO2 at 37℃.
RNA extraction
Total RNA from FFPE samples was extracted using RecoverAll™ Total Nucleic Acid Isolation Kit (Ambion-1975, USA) according to the manufacturer's specifications. Trizol reagent (Invitrogen, Carlsbad, CA) was used to extract total RNA from cells. The yield of RNA was determined by a NanoDrop 2000 spectrophotometer (Thermo Scientific, USA), and the integrity of the extracted RNA was evaluated using agarose gel electrophoresis stained with ethidium bromide.
MicroRNA microarray
MiRNA microarray was performed following the manufacturer's protocol to screen the differential miRNA expression profile between solid subtype and adjacent normal tissue samples in 5 pairs of FFPE samples. Briefly, for all samples, total RNA was subjected to Poly A tailing and labeled with biotin by FlashTag Biotin HSR RNA Labeling Kit (Affymetrix, USA). Hybridization was then performed using the Affymetrix GeneChip miRNA 4.0, which contains 2578 human mature miRNAs probe sets (miRbase 20.0). Hybridization signals were produced using the Affymetrix Command Console Software (Affymetrix, USA). Raw data were normalized by the Robust Multichip Average (RMA) algorithm. Then the Linear Models for Microarray Data (LIMMA) algorithm was used to calculate the p value, FDR, and log2|Fold Change| between tumor and normal samples, and differentially expressed miRNAs were defined as those with a FDR<0.05 and log2|Fold Change|>3 or log2|Fold Change|<-3
Real-Time Quantitative Reverse-Transcription Polymerase Chain Reaction (QRT-PCR)
QRT-PCR was used to validate the differential expression between solid subtype and adjacent normal tissue for 4 miRNAs (miR-133b, miR-145-5p, miR-124-3p, miR-155-5p) in 26 pairs of samples, including the 5 pairs used in the microarray assay. Briefly, quantification was performed with a two-step reaction process: reverse transcription (RT) to generate the cDNA, and qPCR to quantify each miRNA. The latter was performed on a LightCycler® 480 II Real-time PCR Instrument (Roche, Swiss) using LightCycler® 480 SYBR Green I Master Mix (Roche, Swiss), with a universal primer (Qiagen, Germany) and microRNA-specific primers (Table 1). QPCR parameters were: 95℃ for 10 min, followed by 40 cycles of 95℃ for 10 s, 60℃ for 30 s. Each sample was run in triplicate. At the end of the PCR cycles, melting curve analysis was performed to validate the specific generation of the expected PCR product. The expression levels of microRNAs were normalized to U6 and were calculated using the 2-ΔΔCt method [30].
Primers of 4 miRNAs used for qPCR reaction
| Name | Primer Sequence |
|---|---|
| Universal Primer | Provided by Qiagen |
| U6 | CAAGGATGACACGCAAATTCG |
| hsa-miR-145-5p | GTCCAGTTTTCCCAGGAATCCCT |
| hsa-miR-133b | TTTGGTCCCCTTCAACCAGCTA |
| hsa-miR-124-3p | TAAGGCACGCGGTGAATGCC |
| hsa-miR-155-5p | TTAATGCTAATCGTGATAGGGGT |
Representative pathology images of subtypes of invasive LAC according to the reclassification system of 2011. (A) Lepidic (HE, ×60), (B) Acinar (HE, ×100), (C) Solid (HE, ×100), (D) papillary (HE, ×150), (E) Micropapillary (HE, ×200).
Cell transfection
The LAC cell lines A549 and SPC-A1 were seeded into six-well plates (2×105 or 5×105 separately) and grown to a suitable cell confluence. 24 hours later, miR-133b mimic (miR-133) and miRNA negative control (miR-NC) (Ribobio, China) were both transfected using the RiboFECTTM CP reagent (Ribobio, China) following the manufacturer's instructions. The final concentration of miR-133b or miR-NC was 100nM. Another miRNA mimic negative control labeled with FAM (miR-NC-FAM) (Ribobio, China) was used to assess the transfection efficiency.
Cell proliferation assay
The proliferation viability of A549 and SPC-A1 were evaluated by Cell Counting Kit-8 (CCK-8) (Dojindo, Japan). Cells were seeded at 1×103/well (A549) or 3×103/well (SPC-A1) into 96 plates, then transfected with miR-133b or miR-NC after incubating overnight. 48 hours later, 10μl of CCK8 was added into each well and incubated at 37°C for 2 hours. The plates were read on a robust microplate reader (Multiskan MK3, Thermo Scientific, USA) at a wavelength of 450nm. Absorbance values were normalized to media control.
Cell migration assay
24 hours after transfection, cells were trypsinized and counted to seed (2×104 for A549 and 4×104 for SPC-A1) into the upper chamber of the transwell (8μm pore size; Costar, USA), with 200μl of serum-free medium added into the top and 600μl of full medium added into the bottom of each well respectively. After incubation at 37°C in 5% CO2 for 24h, cells still on the upper surface of the chamber were removed, and the cells that migrated through the membrane were fixed by methanol and stained with crystal violet to assess the cell number.
Cell invasion assay
The procedure of the cell invasion assay was similar to that of the cell migration assay, with the addition that the filter was precoated for 4h with 50μl of Matrigel (BD, USA) diluted five-fold with serum-free medium.
Clinical characteristics of 255 patients with resected stage I-III invasive LACs according to the IASLC/ATS/ERS reclassification
| Acinar | Papillary | Solid | Lepidic | Micropapillary | Variant | Sum | |
|---|---|---|---|---|---|---|---|
| Sex | |||||||
| Men | 32 | 28 | 42 | 15 | 2 | 5 | 124(48.6%) |
| Women | 46 | 39 | 26 | 15 | 3 | 2 | 131 (51.4%) |
| Age | |||||||
| ≤60 | 46 | 35 | 43 | 20 | 3 | 6 | 153(60%) |
| >60 | 32 | 32 | 25 | 10 | 2 | 1 | 102(40%) |
| Smoking status | |||||||
| Smoking | 26 | 17 | 38 | 9 | 0 | 4 | 94 (36.9%) |
| Nonsmoking | 52 | 50 | 30 | 21 | 5 | 3 | 161(63.1%) |
| MTD | |||||||
| ≤3cm | 38 | 33 | 22 | 16 | 1 | 110(43.1%) | |
| >3cm,≤5cm | 28 | 24 | 33 | 9 | 2 | 4 | 100(39.2%) |
| >5cm | 12 | 10 | 13 | 5 | 2 | 3 | 45(17.6%) |
| Clinical stage | |||||||
| Stage I | 21 | 33 | 18 | 13 | 1 | 86(33.7%) | |
| Stage II | 22 | 15 | 23 | 8 | 1 | 3 | 72(28.2%) |
| stage III | 35 | 19 | 27 | 9 | 3 | 4 | 97(38.0%) |
| Bronchial stump | |||||||
| Positive | 3 | 2 | 4 | 2 | 1 | 12 (4.7%) | |
| Negative | 75 | 65 | 64 | 28 | 4 | 7 | 243(95.3%) (95.3%) |
LAC, lung adenocarcinoma; MTD, Maximum tumor diameter.
Statistical analysis
Correlation between two categorical parameters was analyzed by the χ2 test or the Fisher's exact test depending on the number of cells present. The Kaplan-Meier method was applied to estimate the differences in survival using the log-rank test. A Cox regression model was used for the multivariate analysis. For QRT-PCR, paired Wilcoxon's rank test was used to compare the differential expression of specific miRNAs and generate a p-value. For cellular experiments, data were expressed as means ± standard deviation (SD), and statistical significance was analyzed by one-way ANOVA or a two-tailed Student's t test. All the statistical analyses were completed by SPSS 21.0 (SPSS Inc., Chicago, IL, USA), and a p-value of <0.05 was considered significant.
Results
Patient Characteristics
The clinicopathologic characteristics of the 255 LAC cases used in our study, including age, sex, smoking status, the maximum tumor diameter (MTD), bronchial stump, and TNM stage are depicted in Table 2. In brief, there were 124 men (48.6%) and 131 women (51.4%), with a median age of 58 (ranging from 30 to 82). 94 patients (36.9%) had smoking history. 110 patients (43.1%) had an MTD≤3cm, 100 patients (39.2%) had an MTD between 3cm and 5cm, and 45 patients (17.6%) had an MTD>5cm. 86 patients (33.7%) were stage I, 72 patients (28.2%) were stage II, 97 patients (38.0%) were stage III. Bronchial stump invasion (bronchial stump positive) was seen in 12 patients (4.7%), and the remaining 243 patients (95.3%) had no bronchial stump invasion (bronchial stump negative).
According to the IASLC/ATS/ERS reclassification, all patients were invasive LACs, resulting in 6 predominant subtypes (table 2), including 78(30.6%) acinar predominant adenocarcinoma (APA), 68 (26.7%) solid predominant adenocarcinoma (SPA), 67 (26.3%) papillary predominant adenocarcinoma (PPA), 30 (11.8%) lepidic predominant adenocarcinoma (LPA), 5 (2.0%) micropapillary predominant adenocarcinoma (MPA), and 7 (2.7%) variant predominant LACs (VPA). The 5 MPAs and 7 VAPs were excluded for their small numbers, yielding 243 LACs for further analysis.
Survival analysis
As mentioned above, 243 patients covering the 4 most common LAC subtypes according to IASLC/ATS/ERS reclassification were analyzed for overall survival using a Kaplan-Meier survival analysis. Data showed that the predominant subtype of LAC were significantly associated with overall survival (p=0.009). While APA (n=74) and PPA (n=67) both have a relative moderate impact on patients' survival (with the 5-year OS being 50.8% and 45.2% respectively), LPA (n=30) represents the most favorable subtype and SPA (n=68) the least favorable subtype (with the 5-year OS being 77.5% and 33.5% respectively).
We then evaluated the extent of solid component on clinical outcome of the SPA group. Of all 68 SPA patients, there were 14 cases considered as with the pure solid subtype, with less than 5% of other subtypes. However, Kaplan-Meier analysis showed no significant difference between the pure solid subtype cases and other SPA cases (p=0.852).
To better understand the relationship between the presence of solid subtype and prognosis, as well as other clinical parameters, the 243 cases were further classified into two groups (S+ LAC, n=134; S- LAC, n=109) according to whether the solid subtype was more than 5% of the tumor component or not (Table 3). The presence of solid subtype was associated with poor outcome by univariate analysis. The solid subtype positive LACs (S+ LAC) had a 5-year OS of 40.3% compared with 5-year OS of 53.4% for the solid subtype negative LAC (S- LAC) (p = 0.010). In addition, MTD, clinical stage, and invasion of the bronchial stump was significantly associated with patient survival by univariate analysis, while sex, age, and smoking status was not significantly associated with survival (Table 4, Figure 2).
When considering the influence of MTD, clinical stage, bronchial stump, multivariate analysis demonstrated that the presence of solid subtype, as well as clinical stage, bronchial stump, could exert independent prognosis on the 5-year OS of LAC patients (Table 4).
Correlation between the presence of solid subtype and other clinicopathologic features
As demonstrated in Table 3, the presence of the solid LAC subtype was significantly positively correlated with the smoking status and clinical stage of LAC patients. Other clinical characteristics were not shown to be significantly correlated with the presence of the solid subtype.
Clinical characteristics associated with the presence of solid subtype (n=243)
| Clinical characteristics | S+ LAC | S- LAC | Sum | p value (Log rank test) |
|---|---|---|---|---|
| Sex | ||||
| Men | 68 | 49 | 117 | 0.369 |
| Women | 66 | 60 | 126 | |
| Age | 0.142 | |||
| ≤60 | 85 | 59 | 144 | |
| >60 | 49 | 50 | 99 | |
| Smoking status | 0.006 | |||
| Smoking | 60 | 30 | 90 | |
| Nonsmoking | 74 | 79 | 153 | |
| Clinical stage | 0.011 | |||
| Stage I | 36 | 49 | 85 | |
| Stage II | 44 | 24 | 68 | |
| Stage III | 54 | 36 | 90 | |
| Bronchial stump | 0.562 | |||
| Positive | 7 | 4 | 11 | |
| Negative | 127 | 105 | 232 | |
| MTD | 0.723 | |||
| ≤3cm | 55 | 52 | 109 | |
| >3cm, ≤5cm | 54 | 40 | 94 | |
| >5cm | 23 | 17 | 40 |
S+ LAC, solid subtype positive LAC; S- LAC, solid subtype negative LAC.
Univariate and multivariate analysis of the correlation of clinical characteristics with 5-year OS (n=243)
| Clinical characteristic | NO. of patients | 5-year OS | p value (Log rank test) | Multivariate Analysis | ||
|---|---|---|---|---|---|---|
| HR (95% CI) | p value | |||||
| Sex | 0.599 | Not included in the multivariate analysis | ||||
| Men | 115 | 45.7% | ||||
| Women | 124 | 47.2% | ||||
| Age | 0.843 | Not included in the multivariate analysis | ||||
| ≤60 | 143 | 69.9.0% | ||||
| >60 | 96 | 42.9% | ||||
| Smoking status | 0.382 | Not included in the multivariate analysis | ||||
| Smoking | 89 | 44.6% | ||||
| Nonsmoking | 150 | 47.1% | ||||
| Clinical stage | 0.000 | 1.530(1.176-1.989) | 0.002 | |||
| Stage I | 83 | 59.6% | ||||
| Stage II | 67 | 52.0% | ||||
| Stage III | 89 | 30.3% | ||||
| Bronchial stump | 0.000 | 3.627(1.811-7.267) | 0.000 | |||
| Positive | 11 | 0.0% | ||||
| Negative | 228 | 48.8% | ||||
| MTD | 0.027 | 1.218(0.936-1.584) | 0.142 | |||
| ≤3cm | 108 | 56.6% | ||||
| >3cm, ≤5cm | 91 | 49.2% | ||||
| >5cm | 40 | 27.1% | ||||
| subtype | 0.010 | 0.627(0.415-0.948) | 0.027 | |||
| S+ LAC | 134 | 40.3% | ||||
| S- LAC | 105 | 53.4% | ||||
The 5 year-OS of 4 cases were lost. 5-year OS, 5-year overall survival; HR, hazard ratio; 95% CI, 95% confidence. interval
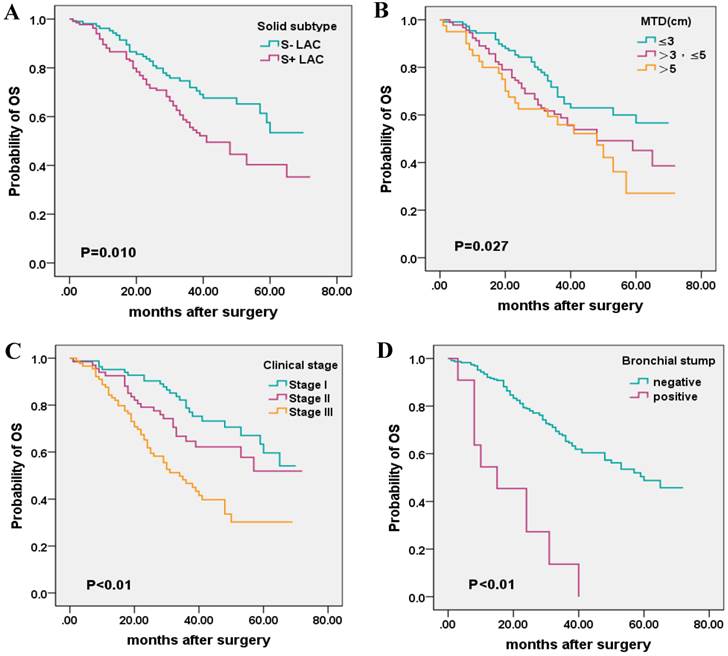
Kaplan-Meier survival plot for OS according to the presence of solid subtype, maximum tumor diameter (MTD), clinical stage, and invasion of the bronchial stump. (A), solid subtype; (B), MTD (C); clinical stage; (D), bronchial stump.
Differential miRNA expression profiles between solid subtype and paired normal tissues
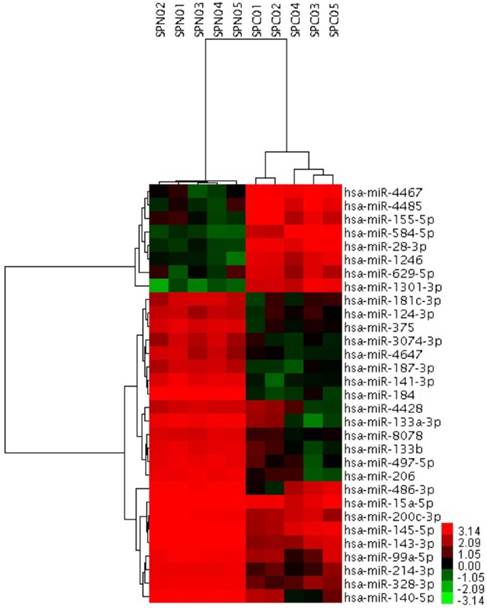
To further investigate the possible role of miRNAs in the solid subtype of LACs, we analyzed miRNAs expression profiles of 5 pairs of solid pure LAC (100% solid subtype) clinical samples and corresponding noncancerous tissues from Biobank of West China Hospital. miRNAs differentially expressed in the solid subtype of LACs were identified using the adjacent normal tissues as a comparison. Candidate miRNAs were those with a log2 |fold change| greater than 3 or smaller than -3, together with a FDR smaller than 0.05. Using this strict criteria, we identified 31 miRNAs that had significantly different expression levels in solid subtype of LAC compared with normal tissue, including 8 upregulated miRNAs and 23 downregulated miRNAs (Figure 3). Of these, several have previously been reported to be dysregulated in cancer. For example, miR-145-5p was identified as a tumor suppressor in many cancers, including our previous work to investigate its possible role in the metastasis of LAC [31]. Other miRNAs identified here that have been associated with cancer pathogenesis in other studies include miR-328-3p, miR-375, miR-99a-5p, miR-200c-3p, miR-15a-5p, miR-124-3p, miR-133b, miR-206, miR-214-3p, and miR-155-5p. Notably, miR-155-5p has previously been described as an onco-miRNA associated with LAC patient's survival [28]. However, there are also novel, differentially expressed miRNAs that had never before been identified as differentially expressed associated in malignancy, including miR-629.5p, miR-4428, miR-4647, and miR-8078.
31 miRNAs showing significantly differential expression between solid subtype and paired normal clinical samples. SPC01-SPC05, solid subtype tissue samples. SPN0-SPN1, paired normal clinical samples.
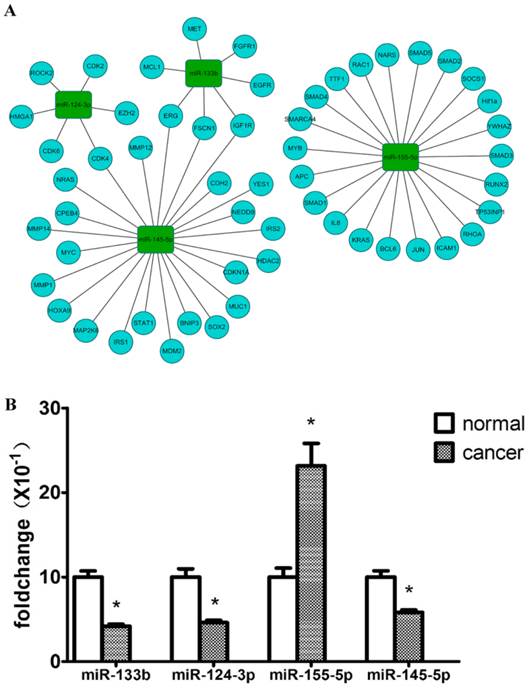
Validation of 4 key dysregulated miRNAs using QRT-PCR. (A) shows hotpot genes in lung cancer that were targeted by 4 key miRNAs. (B) Relative expression levels of 4 key miRNAs between solid LAC subtype and paired normal samples (n=26). Bar graph represents the fold change of log2 values between cancer and paired normal samples ((normalized with the inner control, U6). p<0.05 for all 4 miRNAs.
Bioinformatics analysis
To focus on key miRNAs most likely to play significant roles in the carcinogenesis of the solid subtype of LAC, miRTarbase was used to analyze the validated target genes of the 31 dysregulated miRNAs [32]. We found that these miRNAs target many cancer-associated genes including EGFR, KRAS, HRAS, ERBB2, CDKN2A, and PTEN. According to driver mutations previously reported in lung cancer, the miRNA-mRNA interaction for 4 key miRNAs was visualized by the open-source bioinformatics software platform Cytoscape (www.cytoscape.org) (Figure 4A).
MiRNA validation by QRT-PCR
To verify the accuracy and robustness of the differential expression of miRNAs identified by microarray, expression levels of 4 key miRNAs (miR-145-5p, miR-133b, miR-124-3p, miR-155-5p) were validated using QRT-PCR with an enlarged sample size (n=26), including the 5 pairs used for the microarrays. The significance of differential expression levels of these 4 miRNAs was confirmed with a paired Wilcoxon rank test. Generally, the differential expressions of the 4 miRNAs were consistent with the microarray results, except for variation in the fold difference in expression. As shown in Figure 4B, miR-133b and miR-124-3p were both downregulated in the solid subtype of LAC compared with adjacent normal tissue samples by more than 2-fold (p<0.001, p<0.05 respectively), and miR-145-5p expression was also proved to be significantly reduced in solid subtype of LAC (p<0.05), although with only a 1.5-fold decrease. The up-regulation of miR-155-5p was also verified in cancer tissue samples compared with paired normal tissue samples (p<0.05), with a fold change >2.
Overexpression of miR-133b suppresses the proliferation of LAC cell lines in vitro
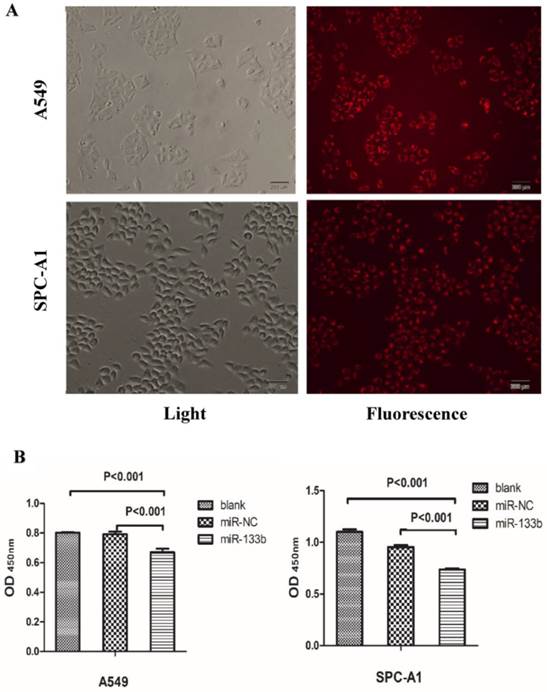
Given these QRT-PCR results, we hypothesized that these 4 key miRNAs likely act as onco-miRNAs or tumor suppressor miRNAs in the carcinogenesis of the solid subtype of LAC. We thus investigated the impact of these miRNAs on the malignant phenotype of LAC. In order to determine the effect of miR-133b on the proliferation viability of LAC cell lines, we utilized CCK8 assays with the LAC cell lines A549 and SPC-A1. miR-133b was overexpressed via transfection of miR-133b mimic. Transfection efficiency was evaluated by miR-NC-FAM (Figure 5A) and QRT-PCR (data not shown). After 48h, CCK8 assays demonstrated decreased proliferation of A549 and SPC-A1 cells overexpressing miR-133b compared with cells transfected with the negative control (miR-NC) or blanks (Figure 5B). Compared with the miR-NC group, the proliferation rate in the miR-133b group of A549 and SPC-A1 was decreased by 15.4% and 22.4%, respectively (p<0.05 for each).
Overexpression of miR-133b did not repress the migration and invasion of LAC cell lines
Migration and invasion are key steps in metastasis that represent an advanced clinical stage of many malignant cancers. To shed light on the possible role of miR-133b in the migration and invasion of LAC cell lines, transwell assays were performed. However, we did not observe any significant effect on the migration and invasion of A549 and SPC-A1 by overexpression of miR-133b (data not shown).
Overexpression of miR-133b can significantly suppress the proliferation of A549 and SPC-A1. (A) The transfection rate of miR-133b was evaluated by a FAM reporter assay using the miR-NC-FAM; (B)CCK-8 assay showed decreased proliferation viability of A549 and SPC-A1 in the miR-133b transfected group compared with miR-NC or blank groups.The OD values were checked 48h after transfection and were normalized to media control. Data are shown as mean OD ± SD from 3 independent experiments. p<0.05 significance was analyzed by one-way ANOVA.
Discussion
As a hallmark of cancer, genome instability [33] generates a variety of mutations to produce subpopulations of cells for Darwinian selection in a given microenvironment, resulting in both inter- and intra-tumor heterogeneity [34, 35]. Despite intense research by high-throughput genetic and genomic technologies in the past several years, tumor heterogeneity still represents a major challenge for personalized-medicine and biomarker development [36].
An important aspect of LAC prognostication specifically remains the histological heterogeneity and resultant subtypes of LAC. In 2011, the new IASLC/ATS/ERS classification system recommended that pathologists assess the most predominant subtype by a 5% increment in mixed LAC. The prognostic value of the predominant subtype of LAC has been well documented [5, 9, 37]. In the present study, univariate analysis validated the prognostic value of LAC predominant subtype in 243 Han Chinese patients with LAC. In accordance with most of the previous studies, we found that SPA had the lowest 5-year OS (33.5%) among the 4 most common predominant subtypes, followed by PPA (45.2%) and APA (50.8%). LPA was the most favorable predominant subtype with a 5-year OS of 77.5%, suggesting clinical utility of this classification system for Chinese patients.
We further explored the prognostic impact of solid subtype by regrouping the patients into S+ LAC group and the S- LAC group. Kaplan-Meier analysis showed that the presence of the solid subtype was a statistically significant risk factor for poor OS, i.e., the S+ LAC group and the S- LAC group each had a 5-year OS of 40.3% and 53.4% separately. This finding differed from a report by Cha et al. [10], who enrolled a cohort of Korea patients. However, another study of Chinese LAC patients came to the similar conclusion to our study [38], suggesting the possibility of ethnic or treatment differences. Besides subtype, Kaplan-Meier survival analysis also showed a significant association of 5-year OS with clinical stage, bronchial stump, and MTD in our study.
A Cox regression model incorporating solid subtype, clinical stage, bronchial stump, and MTD revealed an independent negative prognostic value of the presence of the solid LAC subtype. In addition, we found a significant correlation between the presence of solid subtype, smoking status and clinical stage.
MiRNAs have been one hotspot of investigation for various cancers in recent years, which prompted us to elucidate the possible role of important miRNAs in the pathogenesis of the solid subtype of LAC. We hoped this would help shed light on the relatively poor survival of patients with this subtype. To understand the molecular mechanisms, we compared miRNA expression profiles between 5 pairs of pure solid subtype clinical samples and paired adjacent normal samples to identify 31 dysregulated miRNAs, including 8 upregulated and 23 downregulated miRNAs. Expression levels of 4 of these miRNAs, identified by an informatics analysis as likely to play a pivotal role in carcinogenesis, were confirmed via QRT-PCR, showing miR-133b, miR-124-3p and miR-145-5p had significantly lower expression while miR-155-5p demonstrated increased expression in the solid LAC subtype.
Interestingly, the dysregulation of each of these miRNAs has been previously documented in human cancers. MiR-145 has been shown to be downregulated in prostate [39], colon, and ovarian cancer [40], while miR-155-5p has been shown to be upregulated in LAC [28] and B cell lymphomas [41]. However, to our knowledge, our study is the first to examine the differential expression of miRNAs at the subtype level of LAC.
That miRNAs may affect tumor phenotype via cell functions including proliferation, invasion, migration, cell apoptosis, and cell autophagy is well known. Thus we further performed cellular assays to investigate the possible role of one down regulated miRNA, miR-133b. In previous studies, miR-133b was reported to be a tumor suppressor in bladder cancer [42], esophageal squamous cell carcinoma [43], colorectal cancer [44], and gastric cancer [45]. In contrast, in cervical carcinoma miR-133b has been reported to act as an onco-miRNA to promote tumorigenesis and metastasis [46]. In the present study, we found miR-133b could significantly inhibit the proliferation of LAC cell lines A549 and SPC-A1 in vitro, but appeared to have no effect on the migration and invasion of these cell lines.
There are several limitations to our study. First, data on disease free survival are lacking. Second, cases of MPA and VPA were excluded due to their small numbers, then a larger cohort will be required to more thoroughly investigate the prognostic role of those subtypes. Finally, a larger cohort size is needed to verify the differential expression of these miRNAs and to elucidate their clinical relevance for S+ LAC patients.
In summary, the presence of solid subtype may act as an independent negative prognostic factor incorporating with MTD, bronchial stump and clinical stage. 4 miRNAs including miR-133b, miR-124-3p, miR-145-5p and miR-155-5p are dysregulated in solid subtype of LAC, and thus may be associated with the poor outcome of S+ LAC. One dysregulated miRNA, MiR-133b, can suppress the cell proliferation of LAC cell lines, but had no significant effect on LAC migration and invasion in vitro.
Acknowledgements
This work was supported by grants from the National Basic Research Program of China Project (grant NO. 2013CB967504). We acknowledged Shanghai Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China) and Shanghai OE Biotech. Co. Ltd. (Shanghai, China) for their assistance in miRNA microarray or QRT-PCR assays.
Conflict of Interest
The authors declare that they have no conflicts of interest with the contents of this article.
References
1. Ferlay J, Soerjomataram I, Dikshit R. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. International Journal of Cancer. 2015;136:E359-386
2. Goldstraw P, Ball D, Jett JR. et al. Non-small-cell lung cancer. Lancet. 2011;378:1727-1740
3. Travis WD, Brambilla E, Noguchi M. et al. International association for the study of lung cancer/american thoracic society/european respiratory society international multidisciplinary classification of lung adenocarcinoma. Journal of Thoracic Oncology. 2011;6:244-285
4. Yoshizawa A, Motoi N, Riely GJ. et al. Impact of proposed IASLC/ATS/ERS classification of lung adenocarcinoma: prognostic subgroups and implications for further revision of staging based on analysis of 514 stage I cases. Modern Pathology. 2011;24:653-664
5. Yoshizawa A, Sumiyoshi S, Sonobe M. et al. Validation of the IASLC/ATS/ERS Lung Adenocarcinoma Classification for Prognosis and Association with EGFR and KRAS Gene Mutations Analysis of 440 Japanese Patients. Journal of Thoracic Oncology. 2013;8:52-61
6. Warth A, Muley T, Meister M. et al. The Novel Histologic International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society Classification System of Lung Adenocarcinoma Is a Stage-Independent Predictor of Survival. Journal of Clinical Oncology. 2012;30:1438-1446
7. Solis LM, Behrens C, Raso MG. et al. Histologic patterns and molecular characteristics of lung adenocarcinoma associated with clinical outcome. Cancer. 2012;118:2889-2899
8. Yanagawa N, Shiono S, Abiko M. et al. New IASLC/ATS/ERS Classification and Invasive Tumor Size are Predictive of Disease Recurrence in Stage I Lung Adenocarcinoma. Journal of Thoracic Oncology. 2013;8:612-618
9. Hung JJ, Yeh YC, Jeng WJ. et al. Predictive value of the international association for the study of lung cancer/American Thoracic Society/European Respiratory Society classification of lung adenocarcinoma in tumor recurrence and patient survival. Journal of Clinical Oncology. 2014;32:2357-2364
10. Clay TD, Do H, Sundararajan V. et al. The clinical relevance of pathologic subtypes in metastatic lung adenocarcinoma. Journal of Thoracic Oncology. 2014;9:654-663
11. Cha MJ, Lee HY, Lee KS. et al. Micropapillary and solid subtypes of invasive lung adenocarcinoma: clinical predictors of histopathology and outcome. Journal of Thoracic and Cardiovascular Surgery. 2014;147:921-928 e922
12. Urer HN, Kocaturk CI, Gunluoglu MZ. et al. Relationship between lung adenocarcinoma histological subtype and patient prognosis. Annals of Thoracic and Cardiovascular Surgery. 2014;20:12-18
13. von der Thusen JH, Tham YS, Pattenden H. et al. Prognostic Significance of Predominant Histologic Pattern and Nuclear Grade in Resected Adenocarcinoma of the Lung Potential Parameters for a Grading System. Journal of Thoracic Oncology. 2013;8:37-44
14. Ohtaki Y, Yoshida J, Ishii G. et al. Prognostic significance of a solid component in pulmonary adenocarcinoma. Annals of Thoracic Surgery. 2011;91:1051-1057
15. Zhang Y, Sun Y, Pan Y. et al. Frequency of Driver Mutations in Lung Adenocarcinoma from Female Never-Smokers Varies with Histologic Subtypes and Age at Diagnosis. Clinical Cancer Research. 2012;18:1947-1953
16. de Melo AC, Karen de Sa V, Sternberg C. et al. Mutational Profile and New IASLC/ATS/ERS Classification Provide Additional Prognostic Information about Lung Adenocarcinoma: A Study of 125 Patients from Brazil. Oncology. 2015;89(3):175-186
17. Hu H, Pan Y, Li Y. et al. Oncogenic mutations are associated with histological subtypes but do not have an independent prognostic value in lung adenocarcinoma. Onco Targets and Therapy. 2014;7:1423-1437
18. Pan Y, Zhang Y, Li Y. et al. ALK, ROS1 and RET fusions in 1139 lung adenocarcinomas: a comprehensive study of common and fusion pattern-specific clinicopathologic, histologic and cytologic features. Lung Cancer. 2014;84:121-126
19. Tomonaga N, Nakamura Y, Yamaguchi H. et al. Analysis of Intratumor Heterogeneity of EGFR Mutations in Mixed Type Lung Adenocarcinoma. Clinical Lung Cancer. 2013;14:521-526
20. Carthew RW, Sontheimer EJ. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009;136:642-655
21. Lytle JR, Yario TA, Steitz JA. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5 ' UTR as in the 3 ' UTR. Proceedings of the National Academy of Sciences of the United States of America. 2007;104:9667-9672
22. Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nature Reviews Genetics. 2009;10:704-714
23. Farazi TA, Spitzer JI, Morozov P. et al. miRNAs in human cancer. Journal of Pathology. 2011;223:102-115
24. Volinia S, Calin GA, Liu CG. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proceedings of the National Academy of Sciences of the United States of America. 2006;103:2257-2261
25. Dacic S, Kelly L, Shuai YL. et al. miRNA expression profiling of lung adenocarcinomas: correlation with mutational status. Modern Pathology. 2010;23:1577-1582
26. Arima C, Kajino T, Tamada Y. et al. Lung adenocarcinoma subtypes definable by lung development-related miRNA expression profiles in association with clinicopathologic features. Carcinogenesis. 2014;35:2224-2231
27. Tan X, Qin W, Zhang L. et al. A 5-microRNA signature for lung squamous cell carcinoma diagnosis and hsa-miR-31 for prognosis. Clinical Cancer Research. 2011;17:6802-6811
28. Yanaihara N, Caplen N, Bowman E. et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006;9:189-198
29. Yu SL, Chen HY, Chang GC. et al. MicroRNA signature predicts survival and relapse in lung cancer. Cancer Cell. 2008;13:48-57
30. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402-408
31. Zhao CY, Xu Y, Zhang YQ. et al. Downregulation of miR-145 contributes to lung adenocarcinoma cell growth to form brain metastases. Oncology Report. 2013;30:2027-2034
32. Hsu SD, Tseng YT, Shrestha S. et al. miRTarBase update 2014: an information resource for experimentally validated miRNA-target interactions. Nucleic Acids Research. 2014;42:D78-85
33. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646-674
34. Burrell RA, McGranahan N, Bartek J. et al. The causes and consequences of genetic heterogeneity in cancer evolution. Nature. 2013;501:338-345
35. McGranahan N, Swanton C. Biological and therapeutic impact of intratumor heterogeneity in cancer evolution. Cancer Cell. 2015;27:15-26
36. Gerlinger M, Rowan AJ, Horswell S. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. New England Journal of Medicine. 2012;366:883-892
37. Russell PA, Wainer Z, Wright GM. et al. Does Lung Adenocarcinoma Subtype Predict Patient Survival? A Clinicopathologic Study Based on the New International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society International Multidisciplinary Lung Adenocarcinoma Classification. Journal of Thoracic Oncology. 2011;6:1496-1504
38. Zhang Y, Li J, Wang R. et al. The prognostic and predictive value of solid subtype in invasive lung adenocarcinoma. Scientific Reports. 2014;4:7163
39. Chen X, Gong J, Zeng H. et al. MicroRNA145 targets BNIP3 and suppresses prostate cancer progression. Cancer Research. 2010;70:2728-2738
40. Xu Q, Liu LZ, Qian X. et al. MiR-145 directly targets p70S6K1 in cancer cells to inhibit tumor growth and angiogenesis. Nucleic Acids Research. 2012;40:761-774
41. Eis PS, Tam W, Sun L. et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proceedings of the National Academy of Sciences of the United States of America. 2005;102:3627-3632
42. Chen XN, Wang KF, Xu ZQ. et al. MiR-133b regulates bladder cancer cell proliferation and apoptosis by targeting Bcl-w and Akt1. Cancer Cell International. 2014;14:70
43. Kano M, Seki N, Kikkawa N. et al. MiR-145, miR-133a and miR-133b: Tumor-suppressive miRNAs target FSCN1 in esophageal squamous cell carcinoma. International Journal of Cancer. 2010;127:2804-2814
44. Hu G, Chen D, Li X. et al. miR-133b regulates the MET proto-oncogene and inhibits the growth of colorectal cancer cells in vitro and in vivo. Cancer Biology & Therapy. 2010;10:190-197
45. Zhao Y, Huang J, Zhang L. et al. MiR-133b is frequently decreased in gastric cancer and its overexpression reduces the metastatic potential of gastric cancer cells. Bmc Cancer. 2014:14
46. Qin W, Dong P, Ma C. et al. MicroRNA-133b is a key promoter of cervical carcinoma development through the activation of the ERK and AKT1 pathways. Oncogene. 2012;31:4067-4075
Author contact
Corresponding author: Dr. Xun Hu, Biorepository, State Key Laboratory of Biotherapy, West China Hospital, West China Medical School, Sichuan University, 1 Keyuan 4 Lu, Hi-Tech District, Chengdu, Sichuan 610041, P.R. China. Telephone: +86 028 85164121. Fax: +86 028 85164121. E-mail: hxxhu99com or Dr. Kang Zhang, Institute for Genomic Medicine, University of California, San Diego, La Jolla, CA 92093-0838, USA. Telephone: 858 246 0823; Fax: 858 246 0873. E-mail: kang.zhangcom.

 Global reach, higher impact
Global reach, higher impact