Impact Factor ISSN: 1837-9664
J Cancer 2016; 7(1):107-114. doi:10.7150/jca.13503 This issue Cite
Review
HPV-Testing in Follow-up of Patients Treated for CIN2+ Lesions
1. HPV-UNIT, Regina Elena National Cancer Institute of Rome, Italy
2. Division of Laboratory Medicine, European Institute of Oncology, Milan, Italy
3. Department of Obstetrics and Gynecology - University of Turin, Italy
4. Obstetrics and Gynecology Unit, Vita-Salute San Raffaele University and IRCCS San Raffaele Hospital, Milan, Italy
5. MF Toniolo Hospital, Bologna, Italy
6. Villa Montallegro, Genoa, Italy
† Deceased
Received 2015-8-8; Accepted 2015-11-1; Published 2016-1-1
Abstract
Persistent positivity of HPV-DNA testing is considered a prognostic index of recurrent disease in patients treated for CIN2+. HPV detection, and particularly genotyping, has an adequate high rate of sensitivity and specificity (along with an optimal reproducibility), for accurately predicting treatment failure, allowing for an intensified monitoring activity. Conversely, women with a negative HPV-test 6 months after therapy have a very low risk for residual/recurrent disease, which leads to a more individualized follow-up schedule, allowing for a gradual return to the normal screening scheme. HPV testing should be routinely included (with or without cytology) in post-treatment follow-up of CIN2+ patients for early detection of recurrence and cancer progression. HPV genotyping methods, as a biological indicator of persistent disease, could be more suitable for a predictive role and risk stratification (particularly in the case of HPV 16/18 persistence) than pooled HPV-based testing. However, it is necessary to be aware of the performance of the system, adhering to strict standardization of the process and quality assurance criteria.
Keywords: CIN recurrence, HPV-testing, genotyping, CIN2+, follow-up
Introduction and HPV-Testing
The preventive strategy for cervical cancer is based on identification and treatment of high-grade cervical lesions (HG-CIN) in order to prevent cancer progression. Thus, although conservative, it represents a surgically-based approach (usually by means of laser, cold-knife cone biopsy, or loop electrosurgical excision procedure [LEEP]), which results in an overall high-rate of success. Nevertheless, treatment failures within 2 years (in terms of residual or recurrent high-grade cervical disease) may occur in approximately 5-15% of cases, or even more [[1],[2],[3],[4]], thus often requiring subsequent re-excisional therapy. Moreover, CIN2 and mainly CIN3 treated patients (recognized as the true pre-cancer cervical lesion) remain at high-risk for cervical or other lower-genital tract HPV-related diseases over time. Indeed, for many years the post-treatment risk of invasive cervical cancer is still five-fold greater in such women than that in the general population [[5]]. Furthermore, it has been estimated in the UK that 16% of diagnosed cervical cancers had previously been treated for intraepithelial neoplasia [[6]], confirming the need for a careful and close follow-up of such patients, at least for 10 years after therapy.
For these reasons, high-grade CIN-treated patients are usually kept under clinical post-treatment surveillance, including a combination of cytology, HPV-DNA testing and colposcopic examination. The objective of these monitoring activities is the early detection of patients at high-risk for residual/recurrent disease, with the aim of intensifying the diagnostic procedures or, conversely, returning to normal screening intervals.
Although many patient-related factors (age, smoking habits, number of sexual partners) and pathological characteristics (surgical margins, glandular involvement, lesion size, histological grade and antecedent cytological result) may affect the clinical outcome, the recurrence risk prediction is suboptimal. Women with clear margins following conservative treatment could be at risk for disease recurrence because a multifocal lesion may occur. Conversely, most women with positive resection margins (which suggest an incomplete excision of the lesion) do not develop recurrent disease over time, indicating a limited usefulness of this marker. Moreover, although usually conducted in the surveillance work-up, colposcopic examination has been shown to add little information to the detection rate of residual/recurrent HG-CIN [7, 8].Therefore an accurate test (or combination of tests) able to successfully predict clinical outlook, allowing reduction in the follow-up period and related negative issues (such as anxiety, psychosexual outcomes and overall health cost), would be particularly helpful.
Currently, despite the low-accuracy in detection of recurrent disease (corresponding to high sensitivity, but low specificity [9]), most women treated for high-grade CIN are still monitored by periodical cytological examination. Before returning to routine screening Luesley [10] proposed for CIN2+ patients a cytological follow-up at 6 and 12 months after therapy, and thereafter annually for the next nine years. Indeed, the pap-smear is probably still the most widely accepted follow-up procedure after conization, since the European guidelines for clinical management of abnormal cervical cytology [11] also recommend this test at 6, 12 and 24 months after cone biopsy.
However, a false-negative rate of pap-smears during follow up after treatment has been reported so far [1, 12, 13, 14]. Moreover, in the past decade it has emerged that HPV-DNA testing plays a relevant predictive role, over cytology, after ablative cervical excision. A positive post-treatment HPV-DNA test could predict HG-CIN recurrence more accurately than either cytology, or positive surgical margins, as described in the regression analysis by Paraskevaidis in 2001 [12] and in a subsequent systematic review [1]. To date, a wide range of literature from observational studies of prospective cohorts [7, 15, 13, 17, 19, 20], retrospective case-control study [21, 22], a large study conducted in an integrated health system [23] and meta-analyses or systematic reviews [1,18, 24, 25,27] attest to the usefulness of HPV-DNA test in clinical management after HG-CIN treatment.
It has been documented [28] that the vast majority of CIN3 patients are clear of HPV infection after 6 months from the excisional therapy, dramatically reducing the risk of actual/subsequent recurrence and thus being widely protected against cancer progression. Conversely, HPV infection persistence (and in particular of high-risk genotypes, hr-HPV) during the follow-up period is strictly related to residual or relapsed disease [1,29]. This is because, as already stated [4], ablative treatment removes the dysplastic lesion but not necessarily all the infected tissue. Effective treatment thus results both from removal of the entire lesion and viral clearance as well. Therefore, hr-HPV-DNA persistence after therapy, which is a prerequisite of the original intraepithelial lesion and a critical key-point in cervical cancer oncogenesis [30, 69], may over time further promote the recurrence of disease [31,32]. In other words, the persistent positivity of HPV-DNA testing in the follow-up surveillance after CIN2+treatment (detected in up to one-third of women [1, 12, 19] and even more [29]) assumes a true predictive role.
According to this background, surveillance strategies including a pooled hr-HPV-DNA testing (also defined as a test-of-cure) are reported to be more effective than conventional cytology alone, predicting treatment failure with significantly higher sensitivity and similar (or not-significantly lower) specificity compared to repeat cytology (Table 1). Being a post-therapy monitor activity on high-risk women, not within a normal screening setting on general population, surveillance strategies should prefer sensitivity over the specificity rate, in order not to miss any recurrent disease. Indeed, a high sensitivity is repeatedly reported in the literature for HPV-DNA testing 6 months after conization, ranging from 90 to 100%, as well as a high negative predictive value, which is a relevant index to safely return to a normal screening schedule. Furthermore, hr-HPV-DNA test of cure would also be cost-saving compared with follow-up cytology, as reported by Legood et al [33] in a National Health Service Cervical Screening Programme setting.
Moreover, the combination of cytology and pooled hr-HPV-DNA testing [7, 34] in a “test-of-cure-setting” [35] has been reported to offer greater sensitivity (over 93%) than cytology alone and excellent post-treatment negative predictive value (close to 99% or over), which is highly relevant in this post-treatment group of patients. Therefore, despite the fact that follow-up after CIN2+ conservative treatment has not yet been standardized (varying in terms of timing, interval, and methods), it should include cytology and hr-HPV-DNA testing at 6 months, for early detection of any patients at increased risk of recurrence and cancer progression. The negative result of the two combined tests may further omit the scheduled 12 months check-up examination, to move directly to 24 months after treatment, thus being a less costly procedure. In this regard, Kocken et al [4, 27] stated, through a meta-analysis, that the 5-year risk of post-treatment CIN2+ after three consecutive negative cytological smears or negative co-testing (cytology and hr-HPV testing) at 6 and 24 months was similar to that of women with normal cytology in population-based screening.
However, although the combination strategy promises to better monitor CIN2+ treated patients compared with using cytology alone, it still does not represent the standard follow-up so far. In fact, Cochrane Review by van der Heijden et al [36] found no useful randomized clinical studies directly comparing the two modalities of follow-up strategy (pooled hr-HPV-DNA testing with cytology vs conventional post-treatment cytological at 6-12 months), thus advocating the need for appropriate perspective studies.
Sensitivity and specificity rate of 6 months post-treatment HPV-DNA testing for detecting residual/recurrent CIN 2+
| Author | Pts | % sensitivity | % specificity | % NPV | |
|---|---|---|---|---|---|
| Paraskevaidis E | 2001 | 123 | 93 | 84 | -- |
| Nobbenhius M | 2001 | 184 | 93 | 86 | 99 |
| HoufflinDebarge V | 2003 | 205 | 100 | 67 | 100 |
| Zielinski GD | 2004 | * | 91 | 79 | 98 |
| Sarian L | 2004 | 107 | 80 | 82.9 | |
| Alonso I | 2006 | 203 | 97.2 | 81.4 | 99.3 |
| Kreimer | 2006 | 485 | 90.6 | 63.8 | 99.7 |
| Verguts J | 2006 | 72 | 100 | 77.3 | 100 |
| Chan B | 2009 | ** | 90.7 | 74.6 | 100 |
| Kang WD | 2010 | 672 | 97.3 | 93.1 | 99.8 |
| Leguevaque P | 2010 | 352 | 90.7 | 79.9 | 98.4 |
| Kocken | 2011 | 435 | 92 | 76 | -- |
| Ryu | 2012 | 183 | 100 | 81.9 | 100 |
* pooledmeta-analysis
** systematic review
NPV: negative predictive value
HPV Genotyping
To date, in this surveillance setting, few clinical studies have been conducted to evaluate whether HPV-genotyping will even more accurately predict outcome of HG-CIN-treated patients than will the pooled-probe hr-HPV test.
The objective of the type-specific HPV-test is to distinguish, after conservative therapy, the persistence of the same oncogenic papillomavirus involved in the cervical lesion. Indeed, by definition, to be classified as “persistent” the cervical infection should be ascertained as being related to the same genotype found within the original intraepithelial lesion. Since up to one-third of treated women are still HPV-positive at follow-up, the clinical question is whether this persistence is related to the same original genotype, or belongs to a new infection. In this regard caution must be applied into the evaluation of younger women who may have, after the treatment, a high rate of transient infection.
The literature on this matter, although based on few studies and a small number of patients, agrees to assign a prognostic role of type-specific HPV-testing, with a steady increase in sensitivity rate and a significantly higher specificity. Kreimer et al within the ALTS study [37], comparing three follow-up strategic methods, showed a better sensitivity for hr-HPV persistence with a better specificity compared with conventional follow-up. Analogous findings were reported by Heymans [21], where HPV genotyping showed both a significantly higher sensitivity and specificity to predict residual/recurrent CIN2+. Strand et al [38], recently reported that specific HPV genotype persistence predicts recurrent or residual disease more accurately than does simple pooled HPV-testing, improving the specificity (up to 100%) when using in post-treatment surveillance. Over three years of follow-up, no women with type-specific HPV-negative had recurrent disease and, vice versa, all of the recurrences were persistently positive with the same type. Similar data was also reported by Nobbenhuis [28], where 90% of recurrent CIN2+ were found to be infected by the same high-risk HPV type as before the initial treatment and, conversely, no recurrent disease has been reported in HPV-type specific negative cases. Similarly, outcomes have been reported by Kang et al [39] who stated that detection of the same hr-HPV genotype during follow-up was related to a sensitivity and negative predictive value of 100% for predicting residual/recurrent disease. Conversely, Brismar et al. [40] reported a reduction of sensitivity rate from 100% to 60%, as compared to oncogenic pooled test.
Similarly to sensitivity, specificity and positive predictive value are consistently described as increased due to HPV-genotyping [17,21,39,40]. As expected, among HPV-positive women during the follow-up period, different genotypes confer different risks for post-therapy recurrence and, within the same genotype, variants may also provide additional risk stratification [41]. Patients still infected at 6 months from non-carcinogenic types had a very low-risk of recurrence over 2 years (1.5%), thus very close to HPV-negative women [17]. In this regard, after ablative therapy, a repopulation of cervical and/or vaginal epithelium has been described with low-risk HPV types (from previously infected vaginal walls, or from subsequent contamination), which therefore do not show any significant cancer risk.
Conversely, among high-risk genotypes, patients with HPV16 infection have a greater risk of developing CIN3+ [42], and hence its persistence during the follow-up time may be associated with higher recurrence risk and warrant separate detection. In recurrent patients HPV-16 tends to clear slowly and lesser than the other oncogenic types [22], confirming the clinical evidence of the ALTS trial [17] and other studies [22,43,21,39], where persistent HPV-16 has been detected more frequently associated with recurrent disease. Women positive for HPV-16 at 6 months from cone biopsy [17] have higher risk (37%) of developing recurrent disease over 2 years, compared to those positive to HPV-18 (18.5%) or other carcinogenic genotypes (10.8%). Consistent with these data, Vintemyr et al. in a retrospective registry-based study in Norway [44], reported that 75% of recurrent CIN2+ cases were associated with types HPV 16/18. More recently, also Andrade et al. [22] confirmed a 3-fold greater risk of recurrence for HPV-16 persistence.
Although it has been recently reported [20] that a high percentage of HG-lesions related to HPV 16 or 18 may regress before treatment (thus suggesting that a pretreatment test would not have predictive information), hr-HPV genotyping promises to be useful into the clinical risk stratification. Women affected by HPV-16 high-grade disease need a more appropriate and intensive post-surgical follow-up, providing the opportune methodological test to identify its persistence over time. Conversely, although requiring more confirmation from the literature, a post-therapy detection of HPV genotype different from that present before cone-biopsy, may allow a reduction of the intensity of follow-up.
Finally, also viral load may be regarded as an HPV proxy measure of disease persistence. Many authors [7, 45, 46, 47] have reported that patients with low viral load were less likely to develop recurrent or persistent disease, while other do not confirm this [48, 49]. Although elevated hr-HPV loads have been associated with extensive lesions (more likely to recur), its predictive role is still debated. Indeed, many critical issues (regarding the value of single vs repeated load, pre- vs post-conization sample analysis, the impact of single vs multiple genotypes, the cut-off level expressed in RLU and the selected method for measurement as well) need to be further valued.
HPV Genotyping Methods
Currently, several systems are available for HPV genotyping (Table 2). However, they differ in their analytical performance, in terms of genotypes included, type-specific sensitivities and specificities. One of the main problems is the lack of standardization, and therefore the impossibility of comparing the results obtained by different laboratories. The results of an international proficiency study were recently published [50]: forty-three reference samples were prepared with complete genome of 17 HPV types (12 HR and 5 LR), cloned into plasmid vectors. Different amounts of plasmids were present in the samples and some contained multiple types. The more widely-used HPV genotyping systems were represented, as 96 laboratories participated, some of them providing data on different methods. The results of this survey indicate that, although there was an improvement over the results of the previous survey performed in 2008 [51], the systems still differ in their performance, and this could be due both to the test and to the overall performance of the laboratory.
Today two main groups of methods are commercially available. The first group is able to give a 'partial genotyping', as these tests can detect, through dedicated channels, the presence of HPV-16 and of HPV-18 individually, while the other hr-HPV are detected in pool. These systems have undergone analytical and clinical validation [52, 53, 54, 55, 56], and some of them have also been cleared by the US FDA for the triage of ASCUS cytology and recently the Cobas4800 HPV Test (Roche Diagnostics, Pleasanton, USA) was cleared as a primary test in screening.
Like the Cobas 4800 HPV Test, the Abbott Real Time High Risk HPV test (Abbott Molecular, Des Plaines, IL, USA), belongs to the group of next-generation Real-Time-PCR based HPV DNA assays in which the detection of 14 HPV genotypes is combined with concurrent genotyping for HPV-16 and HPV-18: this test was launched on the European market in January 2009 and is currently used in many laboratories worldwide for routine detection of high risk HPV [57].
Another HPV test clinically validated for HPV genotyping is the Cervista HPV 16/18 test (Hologic, Inc., Madison, WI). This was the first HPV genotyping test approved by the FDA, and permits the differentiation only of HPV-16 and HPV-18 genotypes [58]. Among the different systems available, Qiagen HR-HPV 16/18/45 Probe Set Test is a signal amplification assay based on the most widespread hybrid capture technology (the first FDA validated method) using a probe mix comprising short sequence-specific synthetic probes designed for HPV-16, -18 and -45 detection.
Available systems for HPV genotyping (the grey area is a pooled analysis of HPV genotypes)
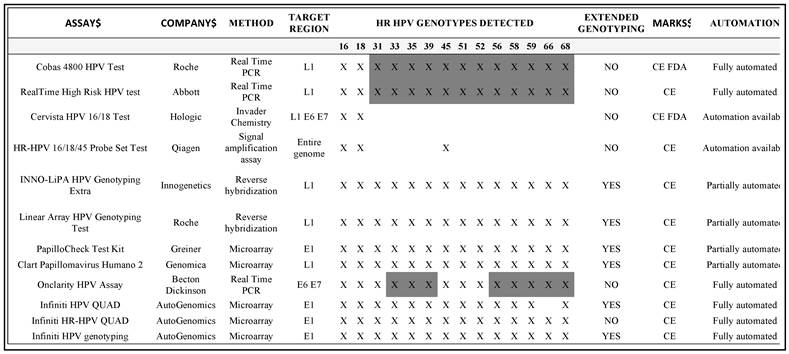
Although these systems may provide sufficient information in more than 50% of patients, who are HPV-16 or HPV-18 positive with CIN2+ lesions, they are not suitable for the follow up of women bearing another hr-HPV. In these cases a second group of HPV genotyping methods, which provide an 'extended' genotyping, should be used. Few systems have been validated for this purpose. They can give a specific response on the presence of single or multiple infections, and are based on different approaches. For example the Linear ArrayTM HPV genotyping test (Roche Diagnostics) is a partially manual method, which is based on reverse line blot hybridization for the detection of 37 genotypes [59]. The same approach is also used for the INNOLIPA HPV Genotyping Extra (Innogenetics, Belgium), although the L1 region of the HPV genome used for amplification is much smaller compared to the Linear Array and the genotypes detected number twenty-eight [56]. Other methods are based on microarray technology. Examples include the Greiner Bio-One PapilloCheck assay (Greiner Bio-One GmbH, Germany), based on the amplification of a 350 bp fragment within the E1-region, which allows the simultaneous identification of 24 genotypes [60, 61], or the Clart® Papillomavirus Humano 2 (Genomica, Spain), which can detect 35 genotypes [62], or the fully automated InfinitiTM Analyzer, based on the detection of different groups of HPV genotypes by using three different reagent kits [63].
Other systems will appear in the near future, and many are now under clinical validation. For example the BD Onclarity HPV assay(BD Diagnostics, Sparks, MD, USA): the Onclarity is a fully-automated E6-E7 DNA-based Real Time PCR assay that detects 16, 18, 31, 45, 51, 52 as single genotypes and the remaining eight genotypes in three groups (33/58, 56/59/66, 35/39/68)[64].
The large number of methods available means that it is important to be aware of the performance characteristics of the particular system in use, and to observe strict standardization of the process and observation of quality assurance criteria.
Conclusion
Since HG-CIN recurrence after surgical therapy is a major marker for progression to invasive cancer, the strategy and procedures of follow-up represent a critical point for risk stratification. Monitoring activity with pooled hr-HPV-DNA testing during the surveillance period has been identified as an independent risk factor associated with recurrent disease, and thus may select those women at high-risk for recurrence [21, 17], reducing unnecessary and expensive procedures. The combination of pooled hr-HPV-DNA testing with cytology represents an accurate index of disease clearance and has been widely accepted (though not formally included into nationwide guidelines) in the routine post-treatment work-out of HG-CIN-treated patients. Furthermore, positive HPV-testing has been also identified, at any time during follow-up, as the most significant independent predictor of recurrent disease in the management of adenocarcinoma in situ [66]. Moreover, based on scientific knowledge and recent emerging data, type-specific HPV after HG-CIN therapy may be an even more significant marker (compared to pooled-based testing) for the development of disease recurrence. Detection at 6 months after conization of the same high-risk HPV genotype is reported to be a better predictor of recurrence in respect to pooled hr-HPV-DNA testing, with a comparable sensitivity and accompanied by a higher specificity. In particular, persistence of HPV 16/18 genotypes are reported to significantly impact on the recurrence rate after conization, thus requiring a close follow-up with immediate reference to colposcopy. Conversely, women negative for HPV type-specific persistence are considered at very low risk for recurrence, leading to a more individualized follow-up schedule. In Table 3, a flow-chart for surveillance of HG-CIN, with an integration of HPV genotyping and cytology 6 months after conservative treatment, is suggested.
Flow-chart with integration of HPV-DNA test and pap-test 6 months after conservative treatment for CIN2+
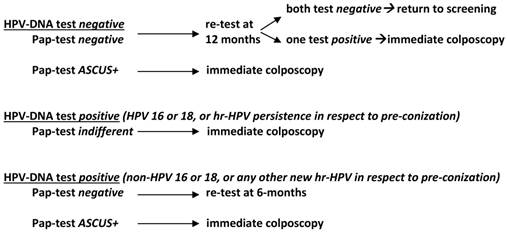
In conclusion, the excellent clinical performance of the combination strategy with HPV genotyping and cytology is considered a powerful tool for managing the post-treatment follow-up of high-grade cervical intraepithelial lesions. In addition, the choice of HPV-test remains of relevant importance given the increasing number of available assays on the market. All of the Authors of the present review use, in their own Institution, validated HPV-test (HC2, Cobas 4800 or Abbott Real Time) in a strategy as shown in Table 3 to follow CIN2+ treated patients. Although most of the commonly used assays were sensitive for detection of CIN2+ disease after therapy [67, 68], it is necessary to be aware of the performance of the system, observing strict standardization of the process and adhering to quality assurance criteria.
Acknowledgements
We thank Samuel William for the excellent review of the English language.
Authors' Contributions
LM conceived and wrote the study.
MTS participated in the international data collection.
MP reviewed the data on HPV pooled system.
MO reviewed the data on cytology.
SC participated in the design of the study.
PC coordinated and helped to draft the manuscript.
FB updated the international data.
MS was the original designer of the review (deceased).
All authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Paraskevaidis E, Arbyn M, Sotiriadis A, Diakomanolis E, Martin-Hirsch P, Koliopoulos G, Makrydimas G, Tofoski J, Roukos DH. The role of HPV DNA testing in the follow-up period after treatment for CIN: a systematic review of the literature. Cancer Treat Rev. 2001;30:205-11
2. Arbyn M, Sasieni P, Meijer CJ, Clavel C, Koliopoulos G, Dillner J. Clinical applications of HPV testing: A summary of meta-analyses. Vaccine. 2006;24:78-89
3. Zielinski GD, Bais AG, Helmerhorst TJ, Verheijen RH, de Schipper FA, Snijders PJ, Voorhorst FJ, van Kemenade FJ, Rozendaal L, Meijer CJ. HPV testing and monitoring of women after treatment of CIN 3: review of the literature and meta-analysis. ObstetGynecol Surv. 2004;59:543-53
4. Kocken M, Uijterwaal MH, de Vries AL, Berkhof J, Ket JC, Helmerhorst TJ, Meijer CJ. Risk of recurrent high-grade cervical intraepithelial neoplasia after successful treatment: a long-term multi-cohort study. Lancet Oncol. 2011;12:441-50
5. Soutter WP, de Barros Lopes A, Fletcher A, Monaghan JM, Duncan ID, Paraskevaidis E, Kitchener HC. Invasive cervical cancer after conservative therapy for cervical intraepithelial neoplasia. Lancet. 1997;349:978-80
6. Soutter WP, Sasieni P, Panoskaltsis T. Long-term risk of invasive cervical cancer after treatment of squamous cervical intraepithelial neoplasia. Int J Cancer. 2006;118:2048-55
7. Alonso I, Torné A, Puig-Tintoré LM, Esteve R, Quinto L, Campo E, Pahisa J, Ordi J. Pre- and post-conization high-risk HPV testing predicts residual/recurrent disease in patients treated for CIN 2-3. Gynecol Oncol. 2006;103:631-6
8. Bigrigg A, Haffenden DK, Sheehan AL, Codling BW, Read MD. Efficacy and safety of large loop excision of the transformation zone. Lancet. 1994;343:32-4
9. Bollen LJ, Tjong-A-Hung SP, van der Velden J, Mol BW, ten Kate FW, Schegget J, Bleker OP. Prediction of recurrent cervical dysplasia by human papillomavirus detection among patients with abnormal cytology. Gynecol Oncol. 1999;72:199-201
10. Luesley D, Leeson S. Colposcopy and Programme Management: Guidelines for the NHS Cervical Screening Programme. NHSCSP Publication No. 20. 2nd Edition. Sheffield: NHSCSP. 2010
11. Jordan J, Martin-Hirsch P, Arbyn M, Schenck U, Baldauf JJ, Da Silva D, Anttila A, Nieminen P, Prendiville W. European guidelines for clinical management of abnormal cervical cytology, Part 2. Cytopathology. 2009;20:5-16
12. Paraskevaidis E, Koliopoulos G, Alamanos Y, Malamou-Mitsi V, Lolis ED, Kitchener HC. Human papillomavirus testing and the outcome of treatment for cervical intraepithelial neoplasia. Obstet Gynecol. 2001;98:833-6
13. Chan J, Bronselaer B, Donders G, Arbyn M, Van Eldere J, Drijkoningen M, Poppe W. Prediction of recurrence after treatment for high-grade cervical intraepithelial neoplasia: the role of human papillomavirus testing and age at conisation. BJOG. 2006. 2006;113:1303-7
14. Jain S, Tseng CJ, Horng SG, Soong YK, Pao CC. Negative predictive value of human papillomavirus test following conization of the cervix uteri. Gynecol Oncol. 2001;82:177-80
15. Cecchini S, Carozzi F, Confortini M, Zappa M, Ciatto S. Persistent human papilloma virus infection as an indicator of risk of recurrence of high-grade cervical intraepithelial neoplasia treated by the loop electrosurgical excision procedure. Tumori. 2004;90:225-8
16. Sarian LO, Derchain SF, Andrade LA, Tambascia J, Morais SS, Syrjanen KJ. HPV DNA test and Pap smear in detection of residual and recurrent disease following loop electrosurgical excision procedure of high-grade cervical intraepithelial neoplasia. Gynecol Oncol. 2004;94:181-6
17. Kreimer AR, Guido RS, Solomon D, Schiffman M, Wacholder S, Jeronimo J, Wheeler CM, Castle PE. Human papillomavirus testing following loop electrosurgical excision procedure identifies women at risk for post-treatment cervical intraepithelial neoplasia grade 2 or 3 disease. CEBP. 2006;15:908-14
18. Chan BKS, Melnikow J, Slee CA. et al. Posttreatment human papillomavirus testing for recurrent cervical intraepithelial neoplasia: a systematic review. Am J Obstet Gynecol. 2009;200:422
19. Costa S, De Simone P, Venturoli S, Cricca M, Zerbini ML, Musiani M, Terzano P, Santini D, Cristiani P, Syrjänen S, Syrjänen K. Factors predicting human papillomavirus clearance in cervical intraepithelial neoplasia lesions treated by conisation. GynecolOncol. 2003;90:358-65
20. Gosvig CF, Huusom LD, Deltour I, Andersen KK, Duun-Henriksen AK, Madsen EM, Petersen LK, Elving L, Schouenbourg L, Iftner A, Svare E, Iftner T, Kjaer SK. Role of human papillomavirus testing and cytology in follow-up after conization. Acta Obstet Gynecol Scand. 2015;94:405-11
21. Heymans J, Benoy IH, Poppe W, Depuydt CE. Type-specific HPV genotyping improves detection of recurrent high-grade cervical neoplasia after conisation. Int J Cancer. 2011;129:903-9
22. Andrade CE1, Scapulatempo-Neto C, Longatto-Filho A, Vieira MA, Tsunoda AT, Da Silva ID, Fregnani JH. Prognostic scores after surgical treatment for cervical intraepithelial neoplasia: a proposed model and possible implications for post-operative follow-up. ActaObstet Gynecol Scand. 2014;93:941-48
23. Katki HA, Schiffman M, Castle PE, Fetterman B, Poitras NE, Lorey T, Cheung LC, Raine-Bennett T, Gage JC, Kinney WK.Five-year risk of recurrence after treatment of CIN 2, CIN 3, or AIS. performance of HPV and Pap cotesting in post-treatment management. J Low Genit Tract Dis. 2013;17:S78-84
24. Zielinski GD, Bais AG, Helmerhorst TJ, Verheijen RH, de Schipper FA, Snijders PJ, Voorhorst FJ, van Kemenade FJ, Rozendaal L, Meijer CJ. HPV testing and monitoring of women after treatment of CIN 3: review of the literature and meta-analyis. ObstetGynecolSurv. 2004;59:543-53
25. Arbyn M, Paraskevaidis E, Martin-Hirsch P, Prendiville W, Dillner J. Clinical utility of HPV DNA detection: triage of minor cervical lesions, follow-up of women treated for high-grade CIN. An update of pooled evidence. Gynecol Oncol. 2005;99(Suppl 3):7-11
26. Leguevaque P, Motton S, Decharme A, Soulé-Tholy M, Escourrou G, Hoff J. Predictors of recurrence in high-grade cervical lesions and a plan of management. Eur J Surg Oncol. 2010;36(11):1073-9
27. Kocken M, Uijterwaal MH, de Vries AL, Berkhof J, Ket JC, Helmerhorst TJ, Meijer CJ. High-risk human papillomavirus testing versus cytology in predicting post-treatment disease in women treated for high-grade cervical disease: A systematic review and meta-analysis. Gynecol Oncol. 2012;125:500-507
28. Nobbenhuis MA, Meijer CJ, van den Brule AJ, Rozendaal L, Voorhorst FJ, Risse EK, Verheijen RH, Helmerhorst TJ. Addition of high-risk HPV testing improves the current guidelines on follow up after treatment for cervical intraepithelial neoplasia. BJC. 2001;84:796-801
29. Elfgren K, Jacobs M, Walboomers JM, Meijer CJ, Dillner J. Rate of human papillomavirus clearance after treatment of cervical intraepithelial neoplasia. Obstet Gynecol. 2002;100:965-71
30. Schlecht NF, Kulaga S, Robitaille J, Ferreira S, Santos M, Miyamura RA, Duarte-Franco E, Rohan TE, Ferenczy A, Villa LL, Franco EL. Persistent human papillomavirus infection as a predictor of cervical intraepithelial neoplasia. JAMA. 2001;286:3106-14
31. Kim YT, Lee JM, Hur SY, Cho CH, Kim SC, Kang SB. Clearance of human papillomavirus infection after successful conization in patients with cervical intraepithelial neoplasia. Int J Cancer. 2010;126:1903-1909
32. Duesing N, Schwarz J, Choschzick M, Jaenicke F, Gieseking F, Mahner RS, Woelber L. Assessment of cervical intraepithelial neoplasia (CIN) with colposcopic biopsy and efficacy of loop electrosurgical excision procedure (LEEP). Arch Gynecol Obstet. 2012;286:1549-1554
33. Legood R, Smith M, Lew JB, Walker R, Moss S, Kitchener H, Patnick J, Canfell K. Cost effectiveness of human papillomavirus test of cure after treatment for cervical intraepithelial neoplasia in England: economic analysis from NHS Sentinel Sites Study. BMJ. 2012;345:e7086
34. Prato B, Ghelardi A, Gadducci A, Marchetti I, Di Cristofano C, Di Coscio G, Bevilacqua G, Genazzani AR. Correlation of recurrence rates and times with posttreatment human papillomavirus status in patients treated with loop electrosurgical excision procedure conization for cervical squamous intraepithelial lesions. Int J Gynecol Cancer. 2008;18:90-94
35. Jones J, Saleem A, Rai N, Shylasree T.S, Ashman S, Gregory K, Powell N, Tristram A, Fiander A, Hibbitts S. Human Papillomavirus genotype testing combined with cytology as a 'test of cure' post treatment: The importance of a persistent viral infection. Journal of Clinical Virology. 2011;52:88-92
36. van der Heijden E, Lopes AD, Bryant A, Bekkers R, Galaal K. Follow-up strategies after treatment (large loop excision of the transformation zone (LLETZ)) for cervical intraepithelial neoplasia (CIN): Impact of human papillomavirus (HPV) test (Review). Cochrane Database of Systematic Reviews. 2015;1:CD010757
37. Houfflin Debarge V, Collinet P, Vinatier D, Ego A, Dewilde A, Boman F, Leroy JL. Value of human papillomavirus testing after conization by loop electrosurgical excision for high-grade squamous intraepithelial lesions. Gynecol Oncol. 2003;90(3):587-92
38. Söderlund-Strand A, Kjellberg L, Dillner J. Human Papillomavirus Type-Specific Persistence and Recurrence After Treatment for Cervical Dysplasia. Journal of Medical Virology. 2014;86:634-641
39. Kang WD, Oh MJ, Kim SM, Nam JH, Park CS, Choi HS. Significance of human papillomavirus genotypingwith high-grade cervical intraepithelial neoplasia treated by a loop electrosurgical excision procedure. Am J Obstet Gynecol. 2010;203:72
40. Brismar S, Johansson B, Borjesson M, Arbyn M, Andersson S. Follow-up after treatment of cervical intraepithelial neoplasia by human papillomavirus genotyping. Am J Obstet Gynecol. 2009;20:17
41. Xi LF, Kiviat NB, Wheeler CM, Kreimer A, Ho J, Koutsky LA. Risk of Cervical Intraepithelial Neoplasia Grade 2 or 3 after Loop Electrosurgical Excision Procedure Associated with Human Papillomavirus Type 16 Variants. J Infect Dis. 2007;195:1340-4
42. Khan MJ, Castle PE, Lorincz AT, Wacholder S, Sherman M, Scott DR, Rush BB, Glass AG, Schiffman M. The elevated 10-year risk of cervical precancer and cancer in women with human papillomavirus (HPV) type 16 or 18 and the possible utility of type-specific HPV testing in clinical practice. J Natl Cancer Inst. 2005;20:1072-9
43. Gök M, Coupé VM, Berkhof J, Verheijen RH, Helmerhorst TJ, Hogewoning CJ, Snijders PJ, Meijer CJ. HPV16 and increased risk of recurrence after treatment for CIN. Gynecol Oncol. 2007;104:273-5
44. Vintemyr OK, Iversen O, Thorensen S. et al. Recurrent high-grade cervical lesion after primary conization is associated with persistent human papillomavirus infection in Norway. Gyencol Oncol. 2014;133:159-166
45. Rodriguez-Manfredi A, Alonso I, del Pino M, Fusté P, Torné A, Ordi J. Predictors of absence of cervical intraepithelial neoplasia in the conization specimen. Gynecol Oncol. 2013;128:271-276
46. Ryu A, Nam K, Chung S, Kim J, Lee H, Koh E, Bae D. Absence of dysplasia in the excisedcervix by a loop electrosurgical excision procedure in the treatment of cervical intraepithelial neoplasia. J Gynecol Oncol. 2010;21:87-92
47. Park JY, Lee KH, Dong SM, Kang S, Park SY, Seo SS. The association of pre-conization high-risk HPV load and the persistence of HPV infection and persistence/recurrence of cervical intraepithelial neoplasia after conization. GynecolOncol. 2008;108:549-54
48. Bae JH, Kim CJ, Park TC, Namkoong SE, Park JS. Persistence of human papillomavirus as a predictor for treatment failure after loop electrosurgical excision procedure. Int J Gynecol Cancer. 2007;17:1271-7
49. Kulmala SM, Shabalova IP, Petrovitchev N, Syrjänen KJ, Gyllensten UB, Johansson BC, Syrjänen SM. New Independent States of the former Soviet Union Cohort Study Group.Type-specific persistence of high-risk human papillomavirus infections in the New Independent States of the former Soviet Union Cohort Study. CEBP. 2007;16:17-22
50. Eklund C, Forslund O, Wallin KL, Dillner J.Global improvement in genotyping of human papillomavirus DNA. the 2011 HPV LabNet International Proficiency Study. JClinMicrobiol. 2014;52:449-59
51. Eklund C, Zhou T, Dillner J. WHO Human Papillomavirus Laboratory Network.Global proficiency study of human papillomavirus genotyping. J Clin Microbiol. 2010;48:4147-55
52. Einstein MH, Martens MG, Garcia FA, Ferris DG, Mitchell AL, Day SP, Olson MC. Clinical validation of the Cervista HPV HR and 16/18 genotyping tests for use in women with ASC-US cytology. GynecolOncol. 2010;118:116-22
53. Carozzi FM, Burroni E, Bisanzi S, Puliti D, Confortini M, Giorgi Rossi P, Sani C, Scalisi A, Chini F. Comparison of clinical performance of Abbott RealTime High Risk HPV test with thatofhybridcapture 2 assay in a screening setting. J Clin Microbiol. 2011;49:1446-51
54. Poljak M, Ostrbenk A, Seme K, Ucakar V, Hillemanns P, Bokal EV, Jancar N, Klavs I. Comparison of clinical and analytical performance of the Abbott Realtime High Risk HPV test to the performance of hybrid capture 2 in population-based cervical cancer screening. J ClinMicrobiol. 2011;49:1721-9
55. Lloveras B, Gomez S, Alameda F, Bellosillo B, Mojal S, Muset M, Parra M, Palomares JC, Serrano S. HPV testing by cobas HPV test in a population from Catalonia. PLoS One. 2013;8:e58153
56. Heideman DA, Hesselink AT, Berkhof J, van Kemenade F, Melchers WJ, Daalmeijer NF, Verkuijten M, Meijer CJ, Snijders PJ. Clinical validation of the cobas 4800 HPV test for cervical screening purposes. J ClinMicrobiol. 2011;49:3983-5
57. Poljak M, Cuzick J, Kocjan BJ, Iftner T, Dillner J, Arbyn M. Nucleic acid tests for the detection of alpha human papillomaviruses. Vaccine. 2012;30(Suppl 5):F100-6
58. Boers A, Wang R, Slagter-Menkema L, van Hemel BM, Ghyssaert H, van der Zee AG, Wisman GB, Schuuring E. Clinical validation of the Cervista HPV HR test according to the international guidelines for human papillomavirus test requirements for cervical cancer screening. J Clin Microbiol. 2014;52:4391-3
59. Gravitt PE, Peyton CL, Apple RJ, Wheeler CM. Genotyping of 27 human papillomavirus types by using L1 consensus PCR products by a single-hybridization, reverse line blot detection method. JClin Microbiol. 1998;36:3020-3027
60. Dalstein V, Merlin S, Bali C, Saunier M, Dachez R, Ronsin C. Analytical evaluation of the PapilloCheck test, a new commercial DNA chip for detection and genotyping of human papillomavirus. J Virol Methods. 2009;156:77-83
61. Jones J, Powell NG, Tristram A, Fiander AN, Hibbitts S. Comparison of the PapilloCheck DNA micro-array Human Papillomavirus detection assay with Hybrid Capture II and PCR-enzyme immunoassay using the GP5/6+ primer set. J Clin Virol. 2009;45:100-4
62. Pista A, Verdasca N, Oliveira A. Clinical performance of the CLART human papillomavirus 2 assay compared with the hybrid capture 2 test. J Med Virol. 2011;83(2):272-6
63. Erali M, Pattison DC, Wittwer CT, Petti CA. Human papillomavirus genotyping using an automated film-based chip array. J Mol Diagn. 2009;11:439-45
64. Bottari F, Sideri M, Gulmini C, Igidbashian S, Tricca A, Casadio C, Carinelli S, Boveri S, Ejegod D, Bonde J, Sandri MT. Comparison of the performance of the BD Onclarity HPV Assay with Hybrid Capture II in detection of CIN2+ lesions. J Clin Microbiol. 2015;53:2109-14
65. Wright TC Jr, Massad LS, Dunton CJ, Spitzer M, Wilkinson EJ, Solomon D. et al. 2006 consensus guidelines for the management of women with cervical intraepithelial neoplasia or adenocarcinoma in situ. J Low Genit Tract Dis. 2007;11:223-39
66. Costa S, Venturoli S, Origoni M, Preti M, Mariani L, Cristoforoni P, Sandri MT. Performance of HPV DNA testing in the follow-up after treatment of high-grade cervical lesions, adenocarcinoma in situ (AIS) and microinvasive carcinoma. ecancer. 2015;9:528
67. Cubie A, Canham M, Moore C, Pedraza J, Graham C, Cuschieri K. Valuation of commercial HPV assays in the context of post-treatment follow-up: Scottish Test of Cure Study (STOCS-H). J Clin Pathol. 2014;67:458-463
68. Phillips S, Garland SM, Tan JH, Quinn MA, Tabrizi SN. Comparison of the Roche Cobas4800 HPV assay to Digene Hybrid Capture 2, Roche Linear Array and Roche Amplicor for detection of High-Risk Human Papillomavirus Genotypes in Women undergoing treatment for cervical dysplasia. Journal of Clinical Virology. 2015;62:63-65
69. Vici P, Mariani L, Pizzuti L, Sergi D, Di Lauro L, Vizza E, Tomao F, Tomao S, Mancini E, Vincenzoni C, Barba M, Maugeri-Saccà M, Giovinazzo G, Venuti A. Emerging biological treatments for uterine cervical carcinoma. J Cancer. 2014Jan5;5(2):86-97
Author contact
Corresponding author: Luciano Mariani, MD, HPV-UNIT, Regina Elena National Cancer Institute, Via Elio Chianesi 53, Rome (Italy). Email: marianiit

 Global reach, higher impact
Global reach, higher impact