3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2019; 10(1):131-137. doi:10.7150/jca.28216 This issue Cite
Research Paper
Prognostic Value of PDZ-Binding Kinase/T-LAK Cell-Originated Protein Kinase (PBK/TOPK) in Patients with Cancer
Department of General Surgery, the First People's Hospital of Neijiang, Neijiang 641000, Sichuan Province, P. R. China.
Received 2018-6-30; Accepted 2018-10-24; Published 2019-1-1
Abstract
Background: PDZ-binding kinase/T-LAK cell-originated protein kinase (PBK/TOPK) plays a critical role in tumorigenesis and cancer progression. However, the prognostic roles in cancer patients are inconsistent or even controversial. Therefore, we performed a meta-analysis to investigate the prognostic value of PBK/TOPK in cancers.
Methods: Literature search was performed using several online databases (PubMed, Web of Science, Embase, Cochrane Library, and Google Scholar, National Knowledge Infrastructure and Wanfang) for eligible articles published up to May 1, 2018. The relationship between PBK/TOPK expression and prognosis in cancers was investigated by using pooled hazard ratios (HRs) with 95% confidence intervals (CIs) through STATA 12.0 software.
Results: Totally 20 eligible studies were included in this meta-analysis. The pooled results showed that carriers with high protein expression of PBK/TOPK were significantly associated with poor OS (HR: 1.69, 95% CI: 1.33-2.04) in various cancers, and patients with increased PBK/TOPK protein expression were significantly correlated with inferior RFS (HR: 1.63, 95% CI: 1.02-2.24) and short DFS (HR: 1.69, 95% CI: 1.16-2.23).
Conclusions: The findings suggest that PBK/TOPK protein expression might serve as a prognostic tumor marker in cancers.
Keywords: PBK/TOPK, solid tumor, prognosis, meta-analysis
Introduction
PBK (PDZ-binding kinase), also known as T-lymphokine-activated killer cell-originated protein kinase (TOPK), is a 322 amino-acid and mitogen- activated serine/threonine protein kinase [1-2]. PBK/TOPK protein is generally difficult to detect in normal tissues but frequently elevated in cancer tissues, such as breast cancer, lymphoma and bladder cancer [3-5]. Furthermore, PBK/TOPK has been reported to play vital roles in inflammation, cell apoptosis, and cell-cycle regulation and high expression of PBK/TOPK contributes to tumor growth, proliferation, and metastasis [6-9]. Accumulating evidence has reported that PBK/TOPK protein expression was correlated with clinical outcomes in various solid cancers, such as colorectal cancer, cholangiocarcinoma and lung cancer [10-12]. However, the potential prognostic value of PBK/TOPK in human solid tumors is inconsistent and even contradictory. For example, Hayashi et al [13] reported that over-expression of TOPK protein was significantly related with poor prognosis in glioma, which was consistent with the results conducted in lung adenocarcinoma by Wei et al [12], but there was no significant association between PBK/TOPK protein expression and OS in prostate, gastric and epithelial ovarian cancer [14-16]. Conversely, PBK/ TOPK up-regulation was reported to be correlated with longer overall survival in cholangiocarcinoma and oral squamous cell carcinoma [10,17].
To date, there is no study to comprehensively assess the correlation between PBK/TOPK protein expression and prognosis in cancer patients. And considering the limited number of individual study and varied results. Herein, we conducted the meta-analysis with published data to clarify the influence of high PBK/TOPK expression and its impact on the outcomes of different cancers as well as provide an overview of the current roles of PBK/TOPK in tumor prognosis and as a promising therapeutic target.
Materials and Methods
Search strategy and study selection
The literature search was performed by two authors (Zhang Y and Wang R) in the electronic platforms of PubMed, Web of Science, Embase, Cochrane Library, Google scholar, Wanfang and National Knowledge Infrastructure (CNKI). The last search date was May 1, 2018. The search strategy was used as: 'PDZ binding kinase', 'PBK', 'TOPK', 'T-LAK cell-originated protein kinase', 'lymphokine-activated killer T-cell-originated protein kinase', 'Nori-3 protein', 'TOPK' and 'cancer', 'tumour', 'carcinoma', 'neoplasm', 'tumor', 'neoplasia'. Article language was limited to English and Chinese. The references of all relevant articles were manually reviewed to find potentially relevant studies.
Inclusion criteria
(1) studies concerning the prognostic values of PBK/TOPK protein expression in tissues from primary solid cancer; (2) studies with complete information for assessment of hazard ratios (HRs) and their 95% confidence intervals (CIs) for overall survival (OS), progression-free survival (PFS), disease-free survival (DFS) or recurrence-free survival (RFS); (3) original articles in English or Chinese.
Exclusion criteria
(1) articles focused on hematologic neoplasms; (2) study only detected the mRNA expression of PBK/TOPK; (3) letters, editorials, case reports, reviews, comments or abstracts.
Data extraction and quality assessment
The following extracted information was recorded by two independent authors (Zhang Y and Wang R): first author's surname, publication year, number of sample size, number of patients with overexpression of PBK/TOPK protein, country or region, clinical stage or tumor grade, detection method, cut-off value, analytical method, HRs with their 95 % CIs for OS/RFS/DFS/PFS.
To evaluate the quality of studies, the Newcastle- Ottawa Scale (NOS) was applied. We assigned the studies of high quality a scored ≥6.
Statistical analysis
In the meta-analysis, the pooled HRs and their 95% CIs were calculated to investigate the impact of PBK/TOPK expression on the survival of patients using STATA (Version 11.0; StataCorp, College Station, TX). I2 test and Q test were used to assess heterogeneity among the studies, Phet<0.10 and/or I2 > 50% indicates obvious heterogeneity among studies, and then the random effect model was used for significant heterogeneity. Otherwise, a fixed-effect model was applied. Egger's test and Begg's test were conducted to assess potential publication bias. Sensitivity analysis was carried out to explore the stability of the combined results. A p-value < 0.05 was considered as statistically significant.
Results
The study selection procedure is outlined in Figure 1. According to the selection criteria, the potential relevant full-texts were reviewed and further examined. Finally, 20 [10-29] eligible studies which fulfilled all inclusion criteria were selected into this systematic review and meta-analysis.
Up to 3470 participants from 20 studies were enrolled in this meta-analysis. The included articles were published from 2010 to 2018. The sample size varied from 24 to 520. A total of 20 studies investigated the association of PBK/TOPK expression with overall survival (OS) [10-29], and for the correlations between PBK/TOPK expression and secondary outcomes, 2 studies reported progression-free survival (PFS) [13, 16], 3 studies focused on recurrence-free survival (RFS) [12, 20, 28] and 4 studies covered disease-free survival (DFS) [18, 22, 24, 29]. As for cancer type, 12 different kinds of human solid cancers were investigated, including colorectal cancer (CRC) [10, 22, 25], cholangiocarcinoma (CCA) [11], lung cancer (LC) [12, 18, 19, 21], breast cancer (BC) [20], prostate cancer (PCa) [14, 28], oral squamous cell carcinoma (OSCC) [17], epithelial ovarian cancer (EOC) [16], esophageal squamous cell carcinoma (ESCC) [23], nasopharyngeal carcinoma (NPC) [24], gastric cancer (GC) [15, 26, 27], hepatocellular carcinoma (HCC) [29] and glioma [13]. The principal characteristics of all included studies are summarized in Table 1.
PBK/TOPK protein and OS
A total of 20 studies with 3470 patients reported the relationship between PBK/TOPK protein and OS in various cancers. The random-effects model was applied for large heterogeneity existed (I2 = 71.7%; Phet =0.000). As shown in Figure 2, the overall results showed that high PBK/TOPK expression was significantly correlated with poorer OS in various cancers (HR: 1.69, 95% CI: 1.33-2.04).
Flow diagram of studies selection procedure.
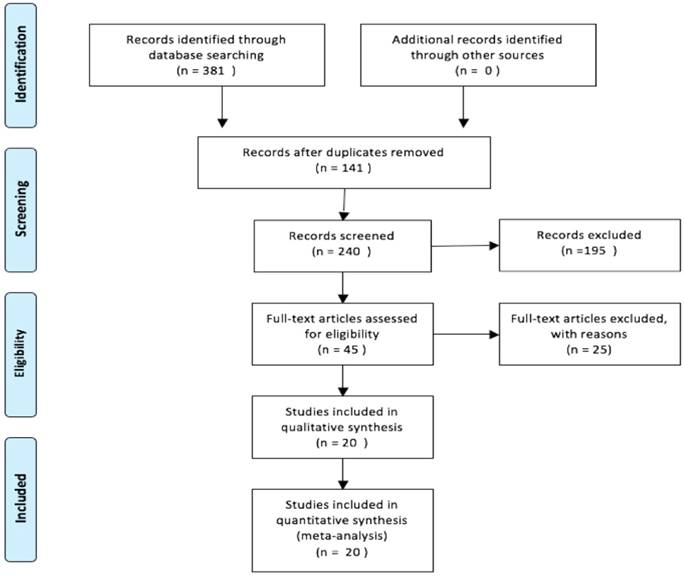
Forest plots of PBK/TOPK protein and OS in various cancers.
Characteristics of eligible studies in this meta-analysis.
| First author | Year | Cancer type | Country | Sample size | Number of OE | Clinical stage/ tumor grade | Criterion of OE | Test methods | Outcome measures | MVA | NOS score | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Zlobec et al [10] | 2010 | CRC | Switzerland | 87 | 36 | NA | diffuse cytoplasmic TOPK staining in >90% of colorectal tumor cells | IHC | OS | Yes | 6 | |
| He et al [11] | 2010 | CCA | China | 24 | 11 | tumor grade 1-2-3 | > the percentages of TOPK-positive cells (10 %) | IHC | OS | Yes | 6 | |
| Wei et al [12] | 2012 | LAC | China | 203 | 136 | I | IHC score >3 | IHC | OS, RFS | Yes | 7 | |
| Shih et al [18] | 2012 | LC | China | 119 | 72 | I-II-III-IV | IHC score (2, 3) | IHC | OS, DFS | Yes | 6 | |
| Lei et al [19] | 2013 | NSCLC | China | 279 | 125 | I-II-III-IV | > the median percentage of positively stained cells ( 11.5%) | IHC | OS | Yes | 8 | |
| O Leary et al [20] | 2013 | BC | Ireland | 290 | 104 | tumor grade 1-2-3 | percentage of cells stained multiplied by intensity score >48 | IHC | OS, RFS | Yes | 8 | |
| Lei et al [21] | 2015 | LAC | China | 127 | 67 | I-II-III-IV | > the average percentages of TOPK-positive cells (13.3%) | IHC | OS | Yes | 7 | |
| Chen et al [14] | 2015 | PCa | China | 98 | NA | NA | immunoreactivity scores (IRS) | IHC | OS | No | 6 | |
| Xiao et al [22] | 2015 | CRC | China | 186 | 135 | tumor grade 1-2-3 | IHC score >3 | IHC | OS, DFS | Yes | 7 | |
| Chang et al [17] | 2016 | OSCC | China | 287 | 75 | I-II-III-IV | mean PBK/TOPK expression score | IHC | OS | Yes | 7 | |
| Ikeda et al [16] | 2016 | EOC | Japan | 163 | 84 | I-II-III-IV | 2+, 3++ | IHC | OS, PFS | No | 7 | |
| Ohashi et al [23] | 2016 | ESCC | Japan | 54 | 10 | 0-I-II-III-IV | intensity plus proportion scores ≥5 | IHC | OS | Yes | 6 | |
| Wang et al [24] | 2016 | NPC | China | 185 | 92 | NA | NA | IHC | OS, DFS | Yes | 7 | |
| Zou et al [25] | 2017 | CRC | China | 80 | 43 | NA | NA | IHC | OS | No | 6 | |
| Kwon et al [15] | 2017 | GC | Korea | 385 | 79 | I-II-III | +,++ | IHC | OS | Yes | 8 | |
| Su et al [26] | 2017 | GC | China | 79 | 73 | I-II-III-IV | +,++,+++,++++ | IHC | OS | No | 6 | |
| Ohashi et al [27] | 2017 | GC | Japan | 144 | 24 | I-II-III | PBK/TOPK presenting scores ⩾5 tumor cells | IHC | OS | Yes | 7 | |
| Pirovano et al [28] | 2017 | PCa | UK | 128 | NA | NA | > the median score of PBK IHC score | IHC | OS, RFS | Yes | 7 | |
| Yang et al [29] | 2017 | HCC | China | 520 | 292 | I-II-III-IV | > the median score of PBK IHC | IHC | OS, DFS | Yes | 8 | |
| Hayashi et al [13] | 2018 | Glioma | Japan | 32 | 23 | II--III-IV | > median percentage of TOPK-positive cells (12.7%) | IHC | OS, PFS | Yes | 6 |
CRC: colorectal cancer; CCA: cholangiocarcinoma; LAC: lung adenocarcinoma; LC: lung cancer; NSCLC: non-small-cell lung cancer; BC: breast cancer; PCa: prostate cancer; OSCC: oral squamous cell carcinoma; EOC: epithelial ovarian cancer; ESCC: esophageal squamous cell carcinoma; NPC: nasopharyngeal carcinoma; GC: gastric cancer; HCC: hepatocellular carcinoma; IHC: immunohistochemistry; OS: overall survival; RFS: recurrence-free survival; PFS: progression-free survival; DFS: disease-free survival; OE: over-expression; MVA: multivariate analysis NA: not available.
We also investigated the prognostic values of PBK/TOPK in several subgroups (Table 2). We found that PBK/TOPK expression level could serve as a prognostic biomarker in lung cancer (HR=2.30, 95% CI: 1.79-2.81) and CRC (HR=2.39, 95% CI: 1.81-2.97). However, there was no significant difference but a strong trend between PBK/TOPK expression level and OS in prostate cancer (HR=1.61, 95% CI: 0.12-3.10) and gastric carcinoma (HR=1.13, 95% CI: 0.86-1.41). Furthermore, increased expression of PBK/TOPK was related to poor OS in Asia cohorts (HR=1.65, 95% CI: 1.28-2.03) but no significant association was found for European cohorts (HR=2.08, 95% CI: 0.94-3.23). Interestingly, PBK/TOPK expression could be an independent predictor for OS in multiple cancers (HR=1.88, 95% CI:1.42-2.34).
PBK/TOPK protein and DFS/PFS/RFS
The pooled HR for secondary outcomes in cancer patients are presented in Table 3.
Patients with elevated expression of PBK/TOPK protein were significantly associated with inferior RFS (HR: 1.63, 95% CI: 1.02-2.24) and shorter DFS (HR: 1.69, 95% CI: 1.16-2.23), and there was a strong trend but not statistical difference between PBK/TOPK expression and PFS (HR: 2.56, 95% CI: 0.80-4.32).
Publication bias
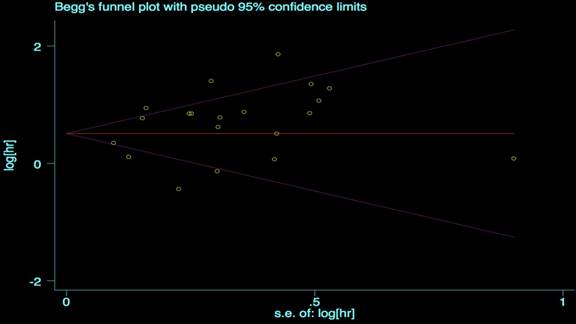
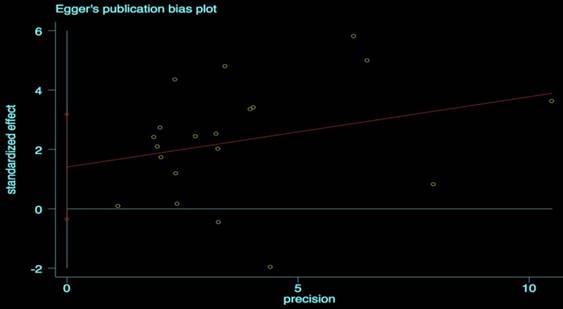
We performed the Begg's and Egger's test to identify potential publication bias. The visible plots were shown in Figure 3 and Figure 4, the test results all indicated that there was no significant publication bias in this meta-analysis (Pr Begg's test > |z| = 0.538 (continuity corrected); Pr Egger's test > |z|=0.111).
Sensitivity analysis
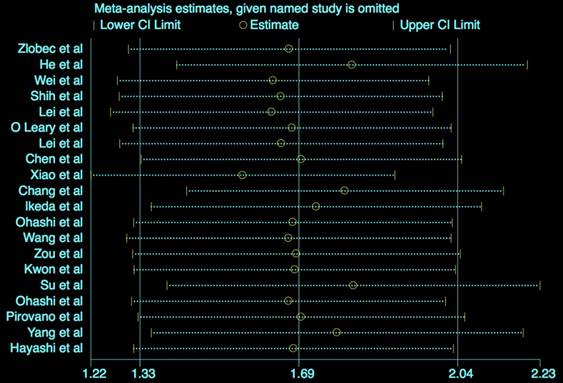
The results (Figure 5) showed that the exclusion of any individual study did not change the statistical significance, suggesting the robustness of our results.
Begg's funnel plot for OS.
Egger's publication bias plot for OS.
Sensitivity analysis for OS.
Meta-analysis results of the relationship between PBK/TOPK protein and OS.
| Group/ subgroup | No. of data set | No. of cases | Pooled HR (95%CI) | p-value | Model | Phet | I² (%) |
|---|---|---|---|---|---|---|---|
| Overall [10-29] | 20 | 3470 | 1.69 (1.33-2.04) | <0.001 | Random | 71.7 | 0.0 |
| Lung cancer [12, 18, 19, 21] | 4 | 728 | 2.30 (1.79-2.81) | <0.001 | Fixed | 0.524 | 0.0 |
| Prostate cancer [14, 28] | 2 | 226 | 1.61 (0.12-3.10) | NS | Fixed | 0.853 | 0.0 |
| Gastric carcinoma [15, 26, 27] | 3 | 608 | 1.13 (0.86-1.41) | NS | Fixed | 0.141 | 49 |
| Colorectal cancer [10, 22, 25] | 3 | 353 | 2.39 (1.81-2.97) | <0.001 | Fixed | 0.643 | 0.0 |
| European cohorts [10, 20, 28] | 3 | 505 | 2.08 (0.94-3.23) | NS | Fixed | 0.593 | 0.0 |
| Asia cohorts [11-19, 21-27, 29] | 17 | 2965 | 1.65 (1.28-2.03) | <0.001 | Random | 0.000 | 74.9 |
| Multivariate analysis [10-13, 15, 17-24, 27-29] | 16 | 3050 | 1.88 (1.42-2.34) | <0.001 | Random | 0.000 | 77.0 |
Analyses of secondary outcomes for PBK/TOPK protein in tumors.
| Secondary outcomes | No. of data set | No. of cases | Pooled HR (95% CI) | p-value | Model | Phet | I2 (%) |
|---|---|---|---|---|---|---|---|
| RFS [12, 20 ,28] | 3 | 621 | 1.63 (1.02-2.24) | <0.001 | Fixed | 0.490 | 0.0 |
| DFS [18, 22, 24, 29] | 4 | 1010 | 1.69 (1.16-2.23) | <0.001 | Fixed | 0.677 | 0.0 |
| PFS [13, 16] | 2 | 195 | 2.56 (0.80-4.32) | NS | Fixed | 0.594 | 0.0 |
Discussion
Since the prognosis potential of PBK/TOPK protein in human solid cancers is inconsistent and debatable, a quantitative meta-analysis is employed in our study. To our knowledge, this is the first meta-analysis to comprehensively evaluate the effect of PBK/TOPK protein level on prognosis in various tumors. By searching against several online databases, a total of 20 observational studies were ultimately included in our work altogether 3470 subjects, containing 12 different cancer types. After analyzing the data extracted from those eligible articles, we revealed that PBK/TOPK protein expression level was associated with clinical outcomes in solid tumors.
The pooled HR results suggested that a high expression level of PBK/TOPK protein was significantly associated with poor OS (HR: 1.69, 95% CI: 1.33-2.04) in human cancers. According to different cancer types, we found the prognostic value of PBK/TOPK in LC (HR=2.30, 95% CI: 1.79-2.81) and CRC (HR=2.39, 95% CI: 1.81-2.97). But no significant differences were found between PBK/TOPK expression and OS in prostate and gastric cancer, this might be for the relatively small number of studies and needed to be explored by future larger studies. As for different ethnicities, the current findings showed that PBK/TOPK expression might be a significant prognostic predictor in Asian patients but not in Caucasian populations, the findings of different ethnic background should be confirmed by future multi-center researches. In addition, we also found that PBK/TOPK protein might be an independent predictor for OS in human cancers (HR=1.88, 95% CI:1.42-2.34). Meanwhile, the relationships between PBK/TOPK protein level and secondary outcomes were also investigated, and PBK/TOPK was found to be a useful prognostic factor for RFS (HR: 1.63, 95% CI: 1.02-2.24) and DFS (HR: 1.69, 95% CI: 1.16-2.23), but not significant for PFS (HR: 2.56, 95% CI: 0.80-4.32). Generally, PBK/TOPK exhibited the potential clinical utility as a candidate molecular marker in human malignancies.
This present study showed that PBK/TOPK expression was associated with patients' clinical outcomes. A number of researches have studied the roles of PBK/TOPK in cancer development, PBK/TOPK acts as an oncogene in multiple cancers and contributes to tumor progression [8, 21, 23, 30-31]. PBK/TOPK is reported to be highly expressed in cancer tissues and cell lines, and high PBK/TOPK expression tends to indicate higher biological malignant aggressiveness [19, 32]. TOPK knockdown could significantly inhibit tumor growth, invasion and metastasis in vitro and in vivo, conversely, PBK up-regulation displays multiple tumorigenic properties, it could promote cell growth, metastasis and enhance tumor transformation [23-26, 30-31]. PBK/TOPK also correlates with drug response and tumor resistance and showed to be the valid target for antineoplastic kinase inhibitors and serves as a potential therapeutic target for various cancers, such as CRC [33-36], lung cancer [37-38], breast cancer [39] and prostate cancer [40-41].
Nevertheless, there are several limitations in our study. First, we only included the studies published in English or Chinese, and most of the studies included came from the Asian countries. Second, the total sample size was not sufficient enough and all enrolled were retrospective studies. Third, the prognostic potentials of PBK/TOPK protein in other special types of cancers were needed for further investingation. Fourth, there was obvious heterogeneity for OS. Finally, the definition of PBK/TOPK protein overexpression varied in different studies.
In summary, the current meta-analysis indicates that elevated PBK/TOPK protein expression is correlated with poor cancer prognosis, and PBK/TOPK protein might be a promising candidate to predicate the cancer survival. If replicated in future larger-scale, multicenter prospective studies, this finding may support the clinical value of PBK/TOPK as a biomarker in tumors.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Gaudet S, Branton D, Lue RA. Characterization of PDZ-binding kinase, a mitotic kinase. Proc Natl Acad Sci U S A. 2000;97:5167-72
2. Hu F, Gartenhaus RB, Eichberg D, Liu Z, Fang HB, Rapoport AP. PBK/TOPK interacts with the DBD domain of tumor suppressor p53 and modulates expression of transcriptional targets including p21. Oncogene. 2010;29:5464-74
3. Park JH, Lin ML, Nishidate T, Nakamura Y, Katagiri T. PDZ-binding kinase/T-LAK cell-originated protein kinase, a putative cancer/testis antigen with an oncogenic activity in breast cancer. Cancer Res. 2006;66:9186-95
4. Simons-Evelyn M, Bailey-Dell K, Toretsky JA, Ross DD, Fenton R, Kalvakolanu D. et al. PBK/TOPK is a novel mitotic kinase which is upregulated in Burkitt's lymphoma and other highly proliferative malignant cells. Blood Cells Mol Dis. 2001;27:825-9
5. Singh PK, Srivastava AK, Dalela D, Rath SK, Goel MM, Bhatt ML. Expression of PDZ-binding kinase/T-LAK cell-originated protein kinase (PBK/TOPK) in human urinary bladder transitional cell carcinoma. Immunobiology. 2014;219:469-74
6. Valiente M, Andrés-Pons A, Gomar B, Torres J, Gil A, Tapparel C. et al. PBK/TOPK promotes tumour cell proliferation through p38MAPK activity and regulation of the DNA damage response. J Biol Chem. 2005;280:28936-43
7. Xiao Y, Feng M, Ran H, Han X, Li X. Identification of key differentially expressed genes associated with non-small cell lung cancer by bioinformatics analyses. Mol Med Rep. 2018;17:6379-6386
8. Seol MA, Park JH, Jeong JH, Lyu J, Han SY, Oh SM. Role of TOPK in lipopolysaccharide-induced breast cancer cell migration and invasion. Oncotarget. 2017;8:40190-40203
9. Fan X, Duan Q, Ke C, Zhang G, Xiao J, Wu D. et al. Cefradine blocks solar-ultraviolet induced skin inflammation through direct inhibition of T-LAK cell-originated protein kinase. Oncotarget. 2016;7:24633-45
10. Zlobec I, Molinari F, Kovac M, Bihl MP, Altermatt HJ, Diebold J. et al. Prognostic and predictive value of TOPK stratified by KRAS and BRAF gene alterations in sporadic, hereditary and metastatic colorectal cancer patients. Br J Cancer. 2010;102:151-61
11. He F, Yan Q, Fan L, Liu Y, Cui J, Wang J. et al. PBK/TOPK in the differential diagnosis of cholangiocarcinoma from hepatocellular carcinoma and its involvement in prognosis of human cholangiocarcinoma. Hum Pathol. 2010;41:415-24
12. Wei DC, Yeh YC, Hung JJ, Chou TY, Wu YC, Lu PJ. et al. Overexpression of T-LAK cell-originated protein kinase predicts poor prognosis in patients with stage I lung adenocarcinoma. Cancer Sci. 2012;103:731-8
13. Hayashi T, Hayakawa Y, Koh M, Tomita T, Nagai S, Kashiwazaki D. et al. Impact of a novel biomarker, T-LAK cell-originating protein kinase (TOPK) expression. Neuropathology. 2018;38:144-153
14. Chen JH, Liang YX, He HC, Chen JY, Lu JM, Chen G. et al. Overexpression of PDZ-binding kinase confers malignant phenotype in prostate cancer via the regulation of E2F1. Int J Biol Macromol. 2015;81:615-23
15. Kwon CH, Park HJ, Choi YR, Kim A, Kim HW, Choi JH. et al. PSMB8 and PBK as potential gastric cancer subtype-specific biomarkers associated with prognosis. Oncotarget. 2016;7:21454-68
16. Ikeda Y, Park JH, Miyamoto T, Takamatsu N, Kato T, Iwasa A. et al. T-LAK cell-originated protein kinase (TOPK) as a prognostic factor and a potential therapeutic target in ovarian cancer. Clin Cancer Res. 2016;22:6110-6117
17. Chang CF, Chen SL, Sung WW, Hsieh MJ, Hsu HT, Chen LH. et al. PBK/TOPK expression predicts prognosis in oral cancer. Int J Mol Sci. 2016:17 pii: E1007
18. Shih MC, Chen JY, Wu YC, Jan YH, Yang BM, Lu PJ. et al. TOPK/PBK promotes cell migration via modulation of the PI3K/PTEN/AKT pathway and is associated with poor prognosis in lung cancer. Oncogene. 2012;31:2389-400
19. Lei B, Liu S, Qi W, Zhao Y, Li Y, Lin N. et al. PBK/TOPK expression in non-small-cell lung cancer: its correlation and prognostic significance with Ki67 and p53 expression. Histopathology. 2013;63:696-703
20. O Leary PC, Penny SA, Dolan RT, Kelly CM, Madden SF, Rexhepaj E. et al. Systematic antibody generation and validation via tissue microarray technology leading to identification of a novel protein prognostic panel in breast cancer. BMC Cancer. 2013;13:175
21. Lei B, Qi W, Zhao Y, Li Y, Liu S, Xu X. et al. PBK/TOPK expression correlates with mutant p53 and affects patients' prognosis and cell proliferation and viability in lung adenocarcinoma. Hum Pathol. 2015;46:217-24
22. Xiao YC, Yang ZB, Cheng XS, Fang XB, Shen T, Xia CF. et al. CXCL8, overexpressed in colorectal cancer, enhances the resistance of colorectal cancer cells to anoikis. Cancer Lett. 2015;361:22-32
23. Ohashi T, Komatsu S, Ichikawa D, Miyamae M, Okajima W, Imamura T. et al. Overexpression of PBK/TOPK contributes to tumor development and poor outcome of esophageal squamous cell carcinoma. Anticancer Res. 2016;36:6457-6466
24. Wang MY, Lin ZR, Cao Y, Zheng LS, Peng LX, Sun R. et al. PDZ binding kinase (PBK) is a theranostic target for nasopharyngeal carcinoma: driving tumor growth via ROS signaling and correlating with patient survival. Oncotarget. 2016;7:26604-16
25. Zou J, Kuang W, Hu J, Rao H. miR-216b promotes cell growth and enhances chemosensitivity of colorectal cancer by suppressing PDZ-binding kinase. Biochem Biophys Res Commun. 2017;488:247-252
26. Su CL, Zhang B, Huang K, Zheng H, Yang D, Sun L. et al. TOPK promotes cell migration and is associated with poor prognosis in gastric cancer. Int J Clin Exp Pathol. 2017;10:3876-3884
27. Ohashi T, Komatsu S, Ichikawa D, Miyamae M, Okajima W, Imamura T. et al. Overexpression of PBK/TOPK relates to tumour malignant potential and poor outcome of gastric carcinoma. Br J Cancer. 2017;116:218-226
28. Pirovano G, Ashton TM, Herbert KJ, Bryant RJ, Verrill CL, Cerundolo L. et al. TOPK modulates tumour-specific radiosensitivity and correlates with recurrence after prostate radiotherapy. Br J Cancer. 2017;117:503-512
29. Yang YF, Pan YH, Cao Y, Fu J, Yang X, Zhang MF. et al. PDZ binding kinase, regulated by FoxM1, enhances malignant phenotype via activation of β-Catenin signaling in hepatocellular carcinoma. Oncotarget. 2017;8:47195-47205
30. Zykova TA, Zhu F, Wang L, Li H, Bai R, Lim DY. et al. The T-LAK cell-originated protein kinase signal pathway promotes colorectal cancer metastasis. EBioMedicine. 2017;18:73-82
31. Ayllón V, O'connor R. PBK/TOPK promotes tumour cell proliferation through p38 MAPK activity and regulation of the DNA damage response. Oncogene. 2007;26:3451-61
32. Luo Q, Lei B, Liu S, Chen Y, Sheng W, Lin P. et al. Expression of PBK/TOPK in cervical cancer and cervical intraepithelial neoplasia. Int J Clin Exp Pathol. 2014;7:8059-64
33. Yang J, Yuan D, Xing T, Su H, Zhang S, Wen J. et al. Ginsenoside Rh2 inhibiting HCT116 colon cancer cell proliferation through blocking PDZ-binding kinase/T-LAK cell-originated protein kinase. J Ginseng Res. 2016;40:400-408
34. Zeng X, Liu L, Zheng M, Sun H, Xiao J, Lu T. et al. Pantoprazole, an FDA-approved proton-pump inhibitor, suppresses colorectal cancer growth by targeting T-cell-originated protein kinase. Oncotarget. 2016;7:22460-73
35. Zhu F, Zykova TA, Kang BS, Wang Z, Ebeling MC, Abe Y. et al. Bidirectional signals transduced by TOPK-ERK interaction increase tumorigenesis of HCT116 colorectal cancer cells. Gastroenterology. 2007;133:219-31
36. Kim DJ, Li Y, Reddy K, Lee MH, Kim MO, Cho YY. et al. Novel TOPK inhibitor HI-TOPK-032 effectively suppresses colon cancer growth. Cancer Res. 2012;72:3060-8
37. Li Y, Yang Z, Li W, Xu S, Wang T, Wang T. et al. TOPK promotes lung cancer resistance to EGFR tyrosine kinase inhibitors by phosphorylating and activating c-Jun. Oncotarget. 2016;7:6748-64
38. Park JH, Inoue H, Kato T, Zewde M, Miyamoto T, Matsuo Y. et al. TOPK (T-LAK cell-originated protein kinase) inhibitor exhibits growth suppressive effect on small cell lung cancer. Cancer Sci. 2017;108:488-496
39. Dou X, Wei J, Sun A, Shao G, Childress C, Yang W, Lin Q. PBK/TOPK mediates geranylgeranylation signaling for breast cancer cell proliferation. Cancer Cell Int. 2015;15:27
40. Sun H, Zhang L, Shi C, Hu P, Yan W, Wang Z. et al. TOPK is highly expressed in circulating tumor cells, enabling metastasis of prostate cancer. Oncotarget. 2015;6:12392-404
41. Brown-Clay JD, Shenoy DN, Timofeeva O, Kallakury BV, Nandi AK, Banerjee PP. PBK/TOPK enhances aggressive phenotype in prostate cancer via β-catenin-TCF/LEF-mediated matrix metalloproteinases production and invasion. Oncotarget. 2015;6:15594-609
Author contact
![]() Corresponding authors: Rong Wang, E-mail: 460532929com and Xu Zhang, E-mail: 1990486976com
Corresponding authors: Rong Wang, E-mail: 460532929com and Xu Zhang, E-mail: 1990486976com

 Global reach, higher impact
Global reach, higher impact