3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2018; 9(20):3776-3786. doi:10.7150/jca.26263 This issue Cite
Research Paper
miR-29b negatively regulates MMP2 to impact gastric cancer development by suppress gastric cancer cell migration and tumor growth
1. The First Affiliated Hospital of Xiamen University, Xiamen, Fujian, 361003, China
2. Department of Gastrointestinal Surgery, Zhongshan Hospital affiliated to Xiamen University, Xiamen, China 361004
3. Department of Basic Medical Sciences of Medical College, Xiamen University, Xiang'an, Xiamen, China 361102
4. State Key Laboratory Breeding Base of Marine Genetic Resources, Third Institute of Oceanography, State Oceanic Administration, Xiamen, China 361005
* These authors contributed equally to this work.
Received 2018-3-24; Accepted 2018-8-11; Published 2018-10-1
Abstract
MicroRNAs (miRNAs) are important regulators and associated with the development of many different types of cancer, including gastric cancer. However, their pathophysiologic role and their relevance to tumorigenesis, invasion and metastasis are still unknown. In our current study, we performed microRNA array and found that 28 of miRNAs were differentially expressed in INF type of gastric cancer. Among 28 miRNAs, miR-29b was one of the most significantly down-regulated miRNA. Further bioinformatics analysis showed that MMP2 was a potential target of miR-29b. Interestingly, luciferase analysis showed that miR-29b negatively regulates MMP2 by binding with the miRNA response element (MRE) on the 3'UTR of MMP2. In addition, overexpression of miR-29b significantly decreased the mRNA and protein level of MMP2 and the activity of MMP2 to suppress gastric cancer cell migration. Moreover, lentivirus mediated overexpression of miR-29b dramatically suppressed the ability of BGC823 cells to form colonies in vitro and their ability to develop tumor in vivo in nude mice. Finally, our qPCR and western blot analysis showed that miR-29b was significantly reduced in clinical gastric cancer tissue, whereas MMP2 protein was significantly up-regulated, suggesting that this aberrant down-regulation of miR-29b might be associated with the abnormal regulation of MMP2 and the development of gastric cancer. Significant apparent was also found between miR-29b expression and TNM staging, lymph node status, tumor differentiation and Ming classification. Together, our data suggest an important regulatory role of miR-29b in the development of gastric cancer. Thus, miR-29b and MMP2 might be important diagnostic or therapeutic targets for human tumor diseases.
Keywords: micorRNA-29b, MMP2, tumor migration, gastric cancer
Introduction
According to the data of the World Health Organization, gastric cancer (GC) is the fifth most common cancer and the third leading cause of cancer mortality in worldwide [1, 2]. Although a series of research progress has been made in diagnosis and treatment of GC, the 5-year survival rate of GC is no more than 25 percent [3]. As one of the most common malignant tumors, early diagnosis and treatment for GC has become an urgent issue. Studies have shown that a series of complicated process such as Helicobacter pylori infection [4], cell proliferation and cell cycle dysfunction [5], angiogenesis [6, 7], cell invasion and metastasis [8] cross-talked and promoted the development of GC. Multiple molecular alterations such as oncogenes and tumor suppressor genes were involved in the development of GC [9]. Tumor invasion and metastasis are the major cause of mortality in human GC patients [10, 11]. The mechanism of gastric tumor metastasis, however, is still not fully understood.
Matrix metalloproteinases (MMPs) are a kind of important endoproteinases including more than 25 members and divided into collagenases (MMP-1, -8, and -13), gelatinases (MMP-2 and -9), stromelysins (MMP-3, -10, and -11), membrane type (MMP-14, -15, -16, and -17) and other MMPs according to their substrate specificity, sequence similarity and domain organization [12-14]. Studies have demonstrated that MMPs serve as important proteolytic enzymes in degradation of extracellular matrix (ECM) are playing key roles in the process of tumor cell invasion and migration [15-17]. As a key member of the MMPs family, MMP-2 was highly expressed in many kinds of tumors such as breast cancer, cervical cancer, bladder cancer and gastric cancer and its expression was closely associated with tumor stage, lymphatic invasion, lymph node metastasis, and recurrence [18-20]. MMP2 expression can be induced by BCL-W and overexpression of MMP2 can promote gastric cancer cell invasion [21]. Moreover, Sewon Noh et al. analyzed the expression of MMP2 in the body fluid from gastric cancer patients and found that MMP2 can serve as a putative biomarker for carcinomatosis in gastric cancer [22]. However, the effect and mechanism of MMP2 gene activity on cell invasion and metastasis of gastric carcinoma remain enigmatic.
In our present study, we investigated the role of microRNA (miRNA)-mediated post-transcriptional regulation of MMP2 expression in BGC823 and MGC803 gastric cancer cells. We found that miR-29b was significantly down-regulated in INF type of gastric cancer and negatively regulates MMP2 by binding with the miRNA response element (MRE) on the 3'UTR of MMP2. In addition, miR-29b-induced repression of MMP2 expression is associated with suppressed gastric cancer cell migration, the ability of BGC823 cells to form colonies in vitro and their ability to develop tumor in vivo in nude mice. Moreover, the lower expression of miR-29b is correlated with substantial induction of MMP2 expression, TNM staging, lymph node status, tumor differentiation and Ming classification in human GC tissues. The results thus reveal an important gene silencing mechanism for the regulation of MMP2 and suggest that miR-29b can serve as a tumor suppressor.
Materials and Methods
Plasmid construction and lentivirus preparation
To construct lentiviral vector overexpressing miR-29b, human MIR-29b gene DNA fragment was PCR amplified from human BGC823 cell genomic DNA with the primers listed in Table S1. The PCR-amplified fragment was inserted into a lentiviral vector pLV-EF1α-MCS-IRES-Puro (pLV-MIR-ctrl) to generate pLV-MIR29b. Viral vector pLV-MIR29b or pLV-MIR-ctrl as well as three lentivirus packaging plasmids (pMDL, pVSVG and pREV) were co-transfected into HEK293T cells. pLV-EF1α-MCS-IRES-Puro, pMDL, pVSVG and pREV were kind gifts from Prof. Jiahuai Han from Xiamen University. Media containing lentiviruses (LV-MIR29b or LV-MIR-ctrl) were collected every 24 h for 3 times and the lentiviruses were purified by ultra-speed centrifugation.
For the construction of WT of human MMP2-3'UTR, primers were designed based on the cDNA sequence from the NCBI, with the forward primer (Table S1) carrying a MluI site and the reverse primer (Table S1) carrying a HindIII site respectively. The 3'UTR fragment was PCR-amplified using cDNA prepared from human GC cell BGC823 with Pyrobest DNA Polymerase (Cat#R005A, TaKaRa, Dalian, China). The resultant DNA fragments were subcloned into pMIR-REPORTTM Luciferase plasmid (Cat#AM5795, Invitrogen, Carlsbad, CA, USA) to generate pMMP2-3'UTR-WT-luc, using the MluI and HindIII sites. The insertion sequences of the resultant plasmids were confirmed by sequencing. For the construction of MT of human MMP2-3'UTR, primers were designed (Table S1) based on the seeding sequences from both miRanda (http://www.microrna.org) and TargetScan (http://www.targetscan.org) algorithms. Site mutation MRE luciferase reporter plasmid was constructed by using Q5® Site-Directed Mutagenesis Kit (Cat# E0554S, New England BioLabs) and the resultant plasmid (pMMP2-3'UTR-MT-luc) was confirmed by sequencing. The resulting two plasmids are shown in Figure S1 in the Supplemental information section. prmt, promoter; Luc, luciferase; SV40-pA, SV40 poly-A.
Cell culture and transient transfection
Human BGC823 and MGC803 cells were obtained from the ATCC (Manassas, VA, USA), and the cells were maintained in DMEM medium (Cat#12491-015, Gibco, Grand island, NY, USA) supplemented with 10% fetal bovine serum (Cat#10099141, Gibco, Grand island, NY, USA) and incubated in a humidified incubator at 37 ºC in 5% CO2. Plasmid DNA or miR-29b precursor (Cat#AM17100, Ambion, Austin, TX, USA) transfections were performed with Lipofectamine 3000 reagent (Cat#L3000015, Invitrogen, Carlsbad, CA, USA) according to the manufacturer's protocol.
Real-time quantitative PCR (qPCR) analyses of mRNAs and miRNAs
For qPCR analyses of mRNA, reverse transcription was performed with TRIzol (Cat#15596-018, Invitrogen, Carlsbad, CA, USA)-extracted total RNAs using a ReverTra Ace-α® Kit as instructed (Cat#FSQ-101, Toyobo, Tokyo, Japan). qPCR was performed using the SYBR Green Real-Time PCR Master Mix (Cat#QPK-212, Toyobo, Tokyo, Japan) and the Step One Plus Real-Time PCR system (Applied Biosystems Inc., Foster City, CA, USA) using appropriate primer pairs as listed in Table S2, according to the manufacturers' protocols and with 18S rRNA as a control.
For miRNAs, qPCR was performed with the stem-loop primers as reported previously [23]. U6 RNA served as an internal control. The miRNA-specific stem-loop primers were listed in Table S2. qPCR was performed with total RNAs, using universal primer and miRNA-specific reverse LNA-primers as listed in Supplementary Table S2, U6 RNA served as an internal control.
Cell migration, invasion and colony formation assays
For wound-healing assays, about 5×105 BGC823 or MGC803 cells were seeded onto 6-well plates. After 24 hours, cells were transfected with 2.5 μg of pLV-MIR29b or control plasmid by using Lipofectamine 3000 reagent according to the manufacturer's protocol. Six hours after transfection, a wound was incised in the center of the confluent culture, followed by careful washing to remove detached cells and the addition of fresh medium. Phase contrast images of the wounded area were recorded using an inverted microscope at indicated time points.
Cell migration was analyzed by Transwell (Cat#3422, Corning, NY, USA) assays according to the manufacturer's instructions. BGC823 or MGC803 cells were seeded and transfected with 2.5 μg of pLV-MIR29b of control plasmid by using Lipofectamine 3000 reagent. 24 hours after transfection, 2.5×104 cells were seeded per upper chambers in serum-free DMEM whereas the lower chambers were loaded with DMEM containing 5% FBS. After 48 hours, the non-migrating cells on the upper chambers were removed by a cotton swab, and cells invaded through the membrane to the underside of the membrane were stained with a 0.1% crystal violet solution and counted manually in eight random microscopic fields.
For colony formation assays, LV-MIR29b or control lentivirus infected BGC823 cells were seeded on six-well plates and maintained in DMEM containing 10% FBS for 2 weeks. Cells were fixed with methanol and stained with 0.5% crystal violet in 50% methanol for 1 hour and colonies larger than 100 μm in diameter were counted.
Animal experiments
All experimental procedures involving animals were performed in accordance with animal protocols approved by the Institutional Animal Use and Care Committee of Xiamen University. For xenograft tumor growth, 6 mice were randomly divided into two groups. LV-MIR29b- or LV-MIR-ctrl-infected BGC823 cells (about 1 × 106) were suspended in 100 µL PBS and then injected s.c. into the right side of the posterior flank of male BALB/c athymic nude mouse at 5 to 6 week. Beginning from the 10th day after the injection, the size of the tumor was measured every 3 days by a Vernier caliper along two perpendicular axes. The volume of the tumors was calculated with the formula: volume (mm3) = length (mm) × width (mm)2 × 0.52. Twenty-five days after the injection, the mice were sacrificed and the tumors were dissected for further analyses.
Clinical samples
All clinical samples were collected with the informed consent of the patients and study protocols that were in accordance with the ethical guidelines of the Declaration of Helsinki (1975) and were approved by the Institutional Medical Ethics Committee of Xiamen University. GC pathological diagnosis was verified by at least two pathologists. 49 pairs of human GC specimens and paired adjacent epithelial tissues were obtained from the First Affiliated Hospital of Xiamen University from 25 January 2011 to October 2015.
Luciferase reporter assays
Luciferase reporter activities were determined using a Luciferase Reporter Gene Assay System (Cat#E1601, Promega, Madison, WI, USA) as instructed. For all luciferase assays, β-galactosidase activities were determined to calibrate for the transfection efficiency. The calibrated value for a proper control was used to normalize all other values to obtain the normalized relative luciferase units (RLU) representing the activities of the MMP2-3' UTR.
Western blot analyses
Cell or tissue protein lysates were separated in 10% SDS-polyacrylamide gels and electrophoretically transferred to nitrocellulose membranes (Amersham). The detection was achieved using the Immobilon Western Chemiluminescent HRP Substrate Kit (Cat#WBKLS0500, Millipore, Darmstadt, Germany). The antibodies used were listed on Table S3.
Gelatin- and casein-zymography analysis
BGC823 cells were transfected with pLV-MIR29b or control plasmid. 24 hours after the transfection, the cell medium were collected and mixed with non-reducing SDS gel sample buffer and applied without boiling to a 10% polyacrylamide gel containing 0.1% SDS and 1 mg/ml gelatin or casein solution. After electrophoresis, the gels were washed in 50 mmol/l Tris-HCl (pH 7.5) containing 0.15 mol/l NaCl, 5 mmol/l CaC12, 5 μmol/l ZnCl, 0.02% NaN3, 0.25% Triton X-100 (three changes) at room temperature, and then incubated in the same buffer without Triton X-100 (two changes) at 37°C for 20 h. Proteins were stained by Coomassie Brilliant Blue R-250 solution.
Other data acquisition, image processing and statistical analyses
Western blot images were captured by Biosense SC8108 Gel Documentation System with GeneScope V1.73 software (Shanghai BioTech, Shanghai, China). Gel images were imported into Photoshop for orientation and cropping. Data are the means ± SEM. One-way ANOVA with Bonferonni's post-test was used for multiple comparisons and the Student's t test (two-tailed) for pair-wise comparisons.
Results
miRNAs are differentially expressed in human gastric cancer tissues
To explore the roles of miRNAs in the regulation of gastric cancer invasion and metastasis, we first compared the miRNA expression profiles of three pair of infiltrative (INF) type of gastric cancer and normal gastric samples. Among human miRNAs analyzed, 28 of them exhibited significantly differential expression in gastric tissues (P < 0.05, Table 1). Among the 21 down-regulated miRNAs, miR-29b (miR-29b-3p) was down-regulated by 3.51-fold (T/N). Interestingly, being one of the most significantly down-regulated miRNA from our array analysis, miR-29b turned out to be members of miR-29 family miRNAs that were found to play important roles in regulation of tumorigenicity of hepatocellular carcinoma (HCC) [24]. This lent credence to the validity of our dataset. However, the signal pathway(s) regulated by miR-29b and the role of miR-29b in tumorigenesis, tumor invasion and metastasis of gastric cancer are still largely unknown.
The down-regulation of miR-29b in INF type of gastric cancer promoted us to explore the possible biological significance of miR-29b in tumorigenesis. miR-29b is a member of miR-29 miRNA family and sequence alignment showed that the sequences of mature miR-29-3ps have high degrees similarity (Figure 1A). Actually, miR-29-3ps also share a common seed sequence of AGCACCA suggesting that these miR-29-3ps miRNAs might have common targets and similar functions (Figure 1A). It is generally accepted that miRNAs exert their function through regulating the expression of their downstream target genes. Thus, putative miR-29b targets were predicted using target prediction programs, TargetScan (http://www.targetscan.org) and miRanda (http://www.microrna.org). Our bioinformatics analysis revealed that MMP2, a member of the matrix metalloproteinase (MMP) gene family, was a potential target of miR-29b. The 3'UTR of human MMP2 mRNA is 1264 bp in length and contained a complementary site for the seed region of miR-29b (Figure 1B). In addition in a previous study, MMP2 had been shown to be up-regulated in gastric cancer [5, 25], suggesting that miR-29b may regulate gastric cancer development by targeting MMP2.
Summary of differentially expressed miRNAs in INF type of gastric cancer compared with normal gastric tissues. *, P values were calculated using Significance Analysis of Microarrays in the Bioconductor package. P value < 0.05 was considered statistically significant.
| miRNAs | Fold change | P* |
|---|---|---|
| Decreased expression: | ||
| hsa-miR-29c | 0.162 | 0.0005 |
| hsa-miR-29b | 0.285 | 0.028 |
| hsa-miR-1184 | 0.398 | 0.006 |
| hsa-miRPlus-E1241 | 0.417 | 0.031 |
| hsa-miR-647 | 0.427 | 0.039 |
| hsa-miRPlus-E1018 | 0.528 | 0.002 |
| hsa-miRPlus-F1026 | 0.569 | 0.047 |
| hsa-miR-187* | 0.578 | 0.029 |
| hsa-let-7d | 0.579 | 0.038 |
| hsa-miR-124* | 0.614 | 0.009 |
| hsa-miR-185* | 0.615 | 0.006 |
| hsa-miR-150 | 0.629 | 0.035 |
| hsv1-miR-H7* | 0.641 | 0.041 |
| hsa-miR-650 | 0.693 | 0.033 |
| hsa-miR-602 | 0.739 | 0.035 |
| ebv-miR-BHRF1-1 | 0.757 | 0.024 |
| hsa-miRPlus-F1028 | 0.766 | 0.015 |
| hsa-miRPlus-E1093 | 0.797 | 0.016 |
| hsa-miR-589* | 0.814 | 0.041 |
| hsa-miRPlus-E1133 | 0.82 | 0.025 |
| hsa-miR-1204 | 0.851 | 0.048 |
| Increased expression: | ||
| hsa-miR-548d-5p | 3.214 | 0.013 |
| hsa-miR-130b* | 2.722 | 0.036 |
| hsa-miRPlus-E1245 | 2.656 | 0.045 |
| hsa-miR-21* | 2.137 | 0.017 |
| hsa-miR-671-5p | 2.034 | 0.005 |
| hsa-miR-516b | 1.328 | 0.025 |
| hsa-miR-320d | 1.09 | 0.006 |
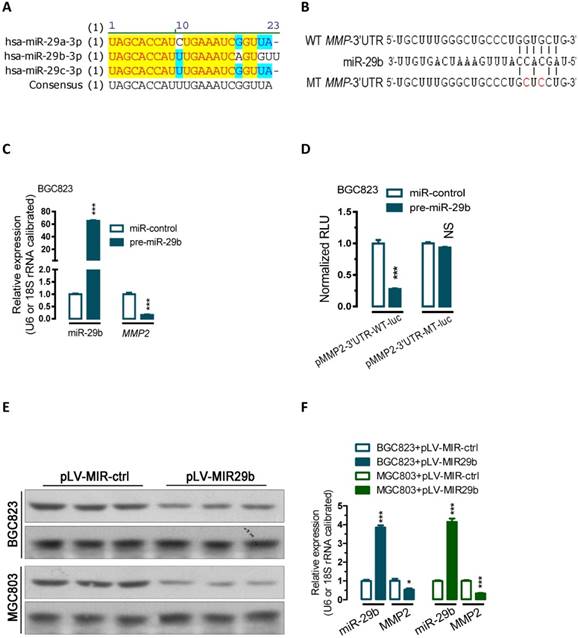
To examine the effect of miR-29b on the endogenous expression of MMP2, we performed transient transfection studies in cultured BGC823 cells. Our quantitative PCR (qPCR) analyses indicated that, compared to the control transfected group (miR-control-transfected group), transfection of BGC823 cells with miR-29b precursor significantly increased miR-29b expression (p < 0.001, Figure 1C), and this induction of the miRNA led to 84% of reduction in MMP2 mRNA (p < 0.001, Figure 1C).
To further determine whether miR-29b actually regulate MMP2 expression, a human MMP2-3'UTR fragment containing wild-type or mutant miR-29b-binding sequence (Figure 1B and Figure S1) was cloned downstream of the firefly luciferase reporter gene in pMIR-REPORTTM Luciferase plasmid. Interestingly, co-transfection with miR-29b precursor significantly decreased the luciferase reporter activity that contained wild type (WT) MMP2- 3'UTR (~73, Figure 1D, p < 0.001). In contrast, the luciferase activity of mutated type (MT) reporter was unaffected by simultaneous transfection of miR-29b precursor (Figure 1D), indicating that miR-29b may suppress MMP2 mRNA expression through binding with the miR-29b-MRE at the 3'UTR of MMP2.
Further protein analysis showed that overexpression of miR-29b by transfeced with a miR-29b overexpression plasmid (pLV-MIR29b) significantly suppressed MMP protein expression both in BGC823 and MGC803 cells (p < 0.05; p < 0.001, Figure 1E and 1F). In addition, Gelatin- and casein-zymography analysis showed that transfection with a plasmid carrying miR-29b precursor (pLV-MIR29b) significantly decreased the activity of MMP2 and MMP9 (Figure S2). These data suggested that miR-29b may inhibit the expression of MMP2 at posttranscriptional level by directly targeting the 3'UTR of MMP2 mRNA.
miR-29b regulate MMP2 post-transcriptionally. (A) Sequence alignment showing that miR-29a/b/c-3p are evolutionarily conserved in human. (B) Bioinformatic prediction of miR-29b recognition elements on the human MMP2-3' UTR. Analyses were performed with either the mirRanda or TargetScan algorithms. (C) Transfection of miR-29 precusor significantly increased miR-29b expression while reduced the MMP2 mRNA expression level in BGC823 cells. (D) miR-29b regulates MMP2 by interacting with the 3' UTR of MMP2. (E and F) Overexpression of miR-29b significantly repressed MMP2 protein expression in BGC823 and MGC803 gastric cancer cells. BGC823 or MGC803 cells were transfected with pLV-MIR29b or control plasmid as indicated. Data are represented as the mean ± SEM (n = 3). **, p < 0.01; ***, p < 0.001.
Effects of overexpression of miR-29b in BGC823 and MGC803 cells on cell migration. For wound-healing assay, BGC823 or MGC803 cells were seeded and transfected with pLV-MIR29b or the control plasmid. 24, 48 or 72 hours after the transfection, wound healing was recorded in photographs. For the transwell migration assay, approximately 2.5 × 104 transfected cells were plated in the upper chamber for transwell assay and the rest of the cells were collected for Western blot analysis. Data are expressed as the mean ± SEM (n = 3). ***, p < 0.001. (A and B) Overexpression of miR-29b significantly suppressed BGC823 and MGC803 cell migration as determined in a wound-healing assay. (C and D) Overexpression of miR-29b significantly suppressed cell migration as determined in a transwell assay.
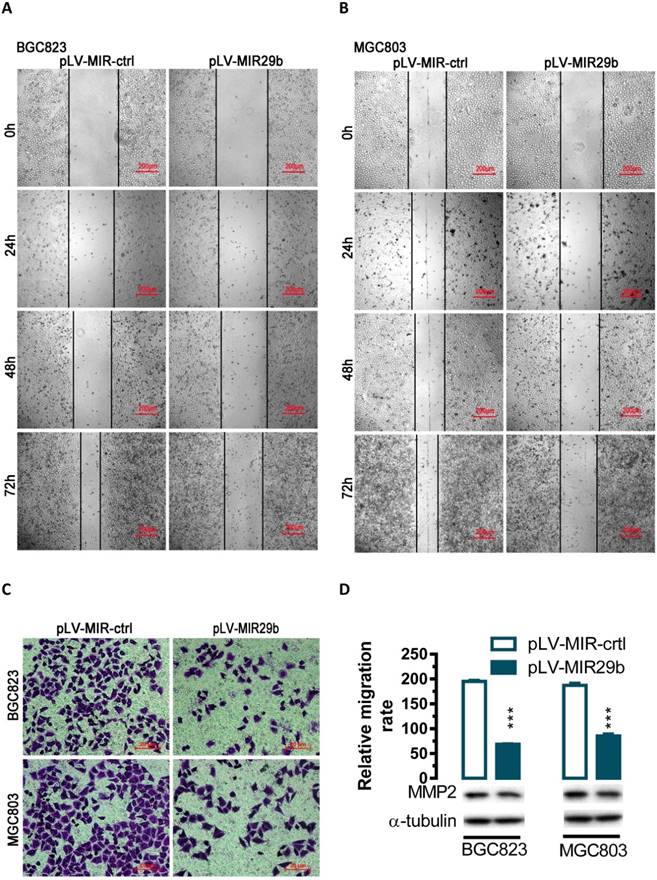
Overexpression of miR-29b suppresses gastric cancer cell migration
To evaluate the impact of miR-29b-mediated regulation of MMP2, we overexpressed miR-29b in BGC823 and MGC803 cells and performed wound healing and transwell experiments. In human BGC823 and MGC803 cells, overexpression of miR-29b greatly suppressed the ability of cell migration (Figure 2A and 2B). To further determine whether overexpression of miR-29b might affect cell migration, we performed transwell migration assays with BGC823 and MGC803 cells. In the transwell migration assay, we found that overexpression of miR-29b significantly suppressed MMP2 protein expression and decreased the likelihood of cell migration as determined by their ability to penetrate the Transwell membrane (P < 0.001, Figure 2C and 2D) in BGC823 and MGC803 cells.
Overexpression of miR-29b significantly inhibits GC cell colony formation in vitro and tumor growth in nude mice in vivo. (A and B) The effect of miR-29b on the colony formation of BGC823 cells. (C) Time-dependent growth of xenograft tumor tissues in nude mice. Lentivirus infected BGC823 cells were harvested and suspended in PBS and then inoculated subcutaneously (~1 × 106 cells/100 ml/mouse) into the right side of the posterior flank of 6-week-old nude mice. (D) Tumor growth in nude mice 25 days after injection with LV-MIR-ctrl- or LV-MIR29b -infected cells. (E) Comparison of tumor weights. (F) Altered expression of MMP2 and PCNA in dissected tumor tissues. Data are expressed as the mean ± SEM (n = 3). NS, not significant; *, p < 0.05; **, p < 0.01; ***, p < 0.001.
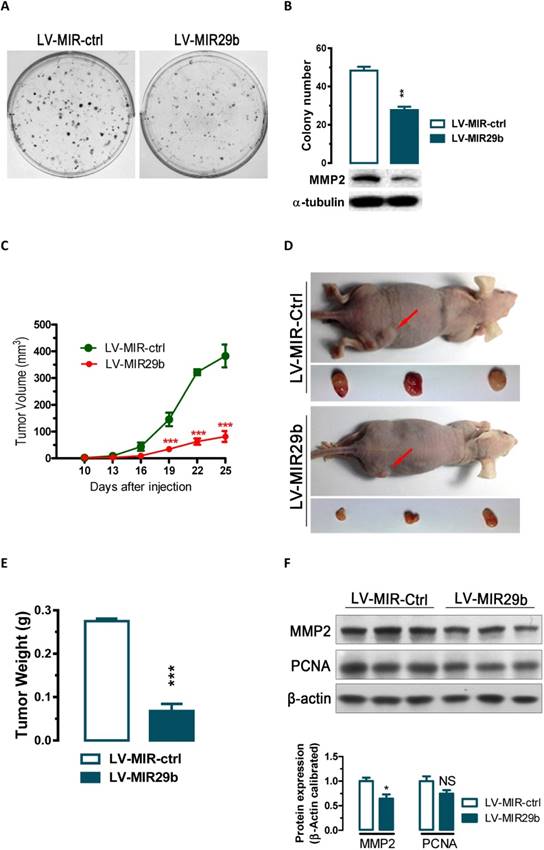
miR-29b suppressed colony formation in vitro and tumorigenicity in vivo in nude mice
The significant reduction of miR-29b expression in gastric cancer samples and suppression of migration and invasion by miR-29b overexpression prompted us to further explore the possible biological significance of miR-29b in tumorigenesis. As an initial step, the capacity of colony formation was evaluated on gastric cancer cell lines (BGC823) that were infected with miR-29b overexpression lentivirus LV-MIR29b or control virus. Interestingly, LV-MIR29b-infected cells displayed much fewer and smaller colonies compared with control virus infected cells (p < 0.05, Figure 3A and 3B). These data indicate a growth-inhibitory role of miR-29b. To further confirm the above findings, an in vivo model was used. Lentivirus infected BGC823 cells (LV-MIR29b- or LV-MIR-ctrl-infected BGC823 cells) were injected s.c. into the right posterior flank of nude mice. In nude mice, tumor growth of LV-MIR29b-infected cells was much slower than that with LV-MIR-ctrl-infected BGC823 cells (Figure 3C). The tumors became palpable from 10 days after inoculation and grew from average of 81 to 382 mm3 at the end of the observation (p < 0.001, Figure 3C). When dissected at the end of the study (day 25), the average tumor weight of the miR-29b overexpressed group were only 24.7% of that of the control group (p < 0.001, Figure 3D and 3E). In tumor tissues, miR-29b overexpression caused 36% (p < 0.05) and 25% (p = 0.0988) of reduction in MMP2 and PCNA expression, respectively (Figure 3F). These results indicate that the introduction of miR-29b significantly inhibits tumorigenicity of BGC823 cells in a nude mouse xenograft model by targeting MMP2.
Inverse correlation between miR-29b expression and MMP2 protein expression in GC clinical samples
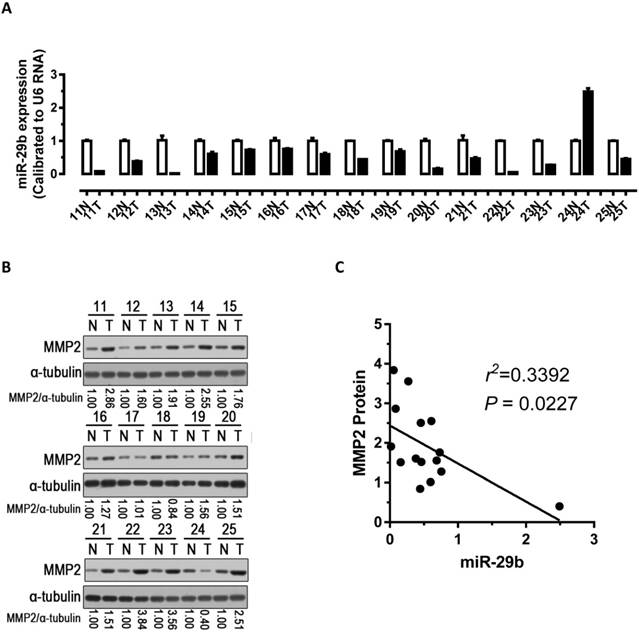
To explore the relationship between the expression of miR-29b and the expression of MMP2 protein, we performed qPCR and western blots with clinical samples. We analyzed miR-29b expression in 15 pairs of gastric cancer samples and found that miR-29b were significantly down-regulated in 93% (14 out of 15, Figure 4A) of the human GC tissues. Further, we analyzed MMP2 protein expression and found it was significantly up-regulated in 87% (13 out of 15, Figure 4B) of gastric cancers. Correlation analyses demonstrated significant inverse correlations between the expression of miR-29b and the expression of MMP2 protein (p < 0.05, Figure 4C) in the 15 pairs of GC samples tested. In addition, we analyzed the correlation of miR-29b expression with clinicopathologic features in 49 pair of gastric cancer tissues (Table 2). The expression of miR-29b was lower expressed in INF type of gastric cancer (p = 0.028) and advanced TNM stage (p = 0.034), and correlated with degree of tumor differentiation (p = 0.036) and lymph node status (p = 0.027). However, no apparent association was found between miR-29b expression with patient gender (p = 0.929), patient age (p = 0.574) and tumor diameter (p = 0.22). These results suggested that the down-regulation of miR-29b may affect the microenvironment of GC by targeting MMP2 to impact the invasion and metastasis of gastric cancer.
Analysis of miR-29b and MMP2 expression in GC samples. (A) miR-29b expression was significantly down-regulated in human GC tissues. (B) Western blot demonstrating the protein expression level of MMP2 in GC tissues. (C) Inverse correlation between the expression of miR-29b and the expression of MMP2 protein in 15 pairs of GC clinical samples. Abbreviations: T, GC tissue; N, adjacent noncancerous gastric tissue; U6 mRNA were calibrated for qPCR analysis; α-tubulin was used as a control for protein loading.
Correlation between the expression of miR-29b and the clinicopathological features of 49 GC patients. P value < 0.05 was considered statistically significant.
| Clinicopathological features | n | % | miR-29b | p Value | |
|---|---|---|---|---|---|
| Gender | M | 35 | 71.4 | 0.512±0.241 | 0.929 |
| F | 14 | 28.6 | 0.441±0.102 | ||
| Age | <60 | 25 | 51 | 0.434±0.167 | 0.574 |
| ≥60 | 24 | 49 | 0.527±0.179 | ||
| Maximum diameter of mass | <5cm | 22 | 44.9 | 0.405±0.19 | 0.22 |
| ≥5cm | 27 | 55.1 | 0.52±0.18 | ||
| TNM stage | Ⅰ, Ⅱ | 14 | 28.6 | 0.645±0.295 | 0.034 |
| Ⅲ, Ⅳ | 35 | 71.4 | 0.43±0.14 | ||
| Degree of differentiation | Low | 23 | 46.9 | 0.43±0.16 | 0.036 |
| Mid | 17 | 34.7 | 0.45±0.23 | ||
| High | 9 | 18.4 | 0.6±0.32 | ||
| Lymph node status | ≥1 | 40 | 71.6 | 0.44±0.17 | 0.027 |
| 0 | 9 | 18.4 | 0.6±0.465 | ||
| Ming classification | EXP | 34 | 69.4 | 0.522±0.142 | 0.028 |
| IFN | 15 | 30.6 | 0.383±0.224 | ||
Discussion
MicroRNAs (miRNAs) represent as oncogenes or tumor suppressors, are a class of key regulators to regulate target messenger RNA (mRNA) expression post-transcriptonally by targeting the 3' untranslated region (UTR) of mRNA [26, 27]. Recent studies demonstrated that aberrant up- or down-regulation of miRNAs were closely related to the occurrence and development of GC [28-30]. Stefano Volinia et al. analyzed miRNA expression profiles in six human solid tumors and found that 22 miRNAs were up-regulated and 6 miRNAs were down-regulated in GC [28]. E2F1 and its host gene Mcm7 regulated miR-106-25 cluster miRNAs were upregulated in a subset of human gastric tumors and play key roles in the development of TGFβ resistance in GC [29]. In our previous study, we found that down-regulated miR-15a-3p and miR-16-1-3p repress Twist1 expression post-transcriptionally and impact GC cell EMT progress to contribute to tumor cell invasion and metastasis [30]. Together these studies demonstrated that abnormal expressed miRNAs are associated with the development and metastasis of tumors.
As a member of matrix metalloproteinase (MMP) family, Metalloproteinase-2 was encoded by MMP2 gene also known as 72 kDa type Ⅳ collagenase or gelatinase A [31]. It has been reported that MMP2 was a direct target of numerous miRNAs associated with the development of many kinds of human tumors such as hepatocellular carcinoma [32, 33], renal cell carcinoma [34], no-small cell lung cancer (NSCLC) [35], breast cancer [36] and pancreatic cancer [37, 38]. Alio Ghosh et al. performed in vitro and in vivo analysis and found that miR-199a-3p can regulate extracellular matrix remodeling by targeting MMP2 to participate in the regulation of the tumor microenvironment [33]. Downregulation of miR-106b was also found in breast cancer and miR-106b can directly target MMP2 to regulate breast cancer cell invasion and motility [36]. Studies have also shown that MMP2 was a direct target of miR-29b in hepatocellular carcinoma [32] and no-small cell lung cancer (NSCLC) [35] as well as miR-29c in pancreatic cancer [37]. The regulatory mechanism and function of MMP2 on cell invasion and metastasis of GC, however, are largely unknown. Here we show that miR-29b represses the mRNA and protein expression of MMP2 by binding with the miRNA response element (MRE) on the 3'UTR of MMP2. Furthermore, lentivirus mediated overexpression of miR-29b significantly suppressed GC cell migration thereby suppressed the ability of GC cells to form colonies in vitro and their ability to develop tumors in vivo in nude mice.
Based on miRBase 21 (http://www.mirbase.org), in human, the miR-29 miRNA family contains MiR-29a, MiR-29b-1, MiR-29b-2 and MiR-29c four precusors and miR-29a/b-1/b-2/c-5p and miR-29a/b/c-3p seven mature miRNAs (Figure 1A and Figure S3). Two miRNA precusors (MiR-29a and MiR-29b-1) were located at chromosome 7q32 and other two (MiR-29b-1 and MiR-29c) were located at chromosome 1q32. Interestingly, our microarray data show that miR-29b and miR-29c were significantly down regulated in human GC tissues from patients (Table 1), this may because MiR-29b-1 and MiR-29c located at the same chromosome position and share one promoter. Since the mature sequence of miR-29a/b/c are conserved evolutionarily in mammals and share the seam seed sequence which determines the target genes (Figure 1A), MMP2 may also be a potential target for miR-29a/c, but this need to be further confirmed.
An important finding in our present study is that miR-29b was significantly down-regulated while MMP2 protein was up-regulated in GC tissues taken from patients. Meanwhile, the expression of miR-29b and MMP2 were inverse correlated in clinical tumor tissues. These results demonstrated that the abnormal expression of miR-29b contributed to the MMP2 related tumor invasion and metastasis. Interestingly, apparent association were also found between miR-29b expression with tumor TNM stage, the degree of tumor differentiation, Lymph node status and Ming classification. This suggest us that miR-29b can serve as a clinical diagnostic marker for the development of GC. Consistent with our results, Sewon Noh et al. found that the expression of MMP2 might be a prognostic marker in ascites of advanced gastric patients with disseminated metastasis [22]. Together these results indicate that miR-29b and MMP2 can server as diagnostic biomarkers and used for the diagnosis of clinical gastric cancer.
In summary, we investigated the roles of miR-29b in GC development and found that miR-29b can severe as a tumor suppressor gene, repressing GC cell migration by targeting the MMP2 gene. The novel regulatory mechanism of MMP2 may play important role in GC growth and recurrence. Theoretically, the identification of miR-29b-MMP2 axis sheds light on the GC gene therapy.
Abbreviations
GC: gastric cancer; miRNA: microRNA; MRE: miRNA recognition elements; 3' UTR: 3' untranslated region; LV: Lentivirus; mRNA: messenger RNA; MMP: metalloproteinase; ECM: extracellular matrix; EMT: epithelial-mesenchymal transition; INF: infiltrative; EXP: expanding; WT: wild type; MT: mutated type; SEM: standard error of the mean.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
We would like to thank Prof. Jiahuai Han from Xiamen University for providing the lentivirus packaging plasmids (pLV-EF1α-MCS-IRES-Puro, pMDL, pVSVG and pREV).
Funding
This work was supported in part by grants from the National Science Foundation for Young Scientists of China (#81502035 and #31501114), China Postdoctoral Science Foundation (#2013M531549) and Fujian Provincial Department of Science and Technology (#2016D009).
Competing Interests
The authors have declared that no competing interest exists.
References
1. McGuire S. World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Advances in nutrition. 2016;7:418-9
2. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2012;136:E359-86
3. Nagini S. Carcinoma of the stomach: A review of epidemiology, pathogenesis, molecular genetics and chemoprevention. World J Gastrointest Oncol. 2012;4:156-69
4. Leung WK, Wong IO, Cheung KS, Yeung KF, Chan EW, Wong AY. et al. Effects of Helicobacter PYLORI Treatment on Incidence of Gastric Cancer in Older Individuals. Gastroenterology. 2018 [Epub ahead of print]
5. Yang GL, Tao HR, Wang HW, Sun Y, Zhang LD, Zhang C. et al. Ara-C increases gastric cancer cell invasion by upregulating CD-147-MMP-2/MMP9 via the ERK signaling pathway. Oncol Rep. 2015;33:2045-51
6. Townley-Tilson WH, Callis TE, Wang D. MicroRNAs 1, 133, and 206: critical factors of skeletal and cardiac muscle development, function, and disease. Int J Biochem Cell Biol. 2010;42:1252-5
7. Javle M, Smyth EC, Chau I. Ramucirumab: successfully targeting angiogenesis in gastric cancer. Clin Cancer Res. 2014;20:5875-81
8. Li X, Zhang Y, Zhang H, Liu X, Gong T, Li M. et al. miRNA-223 promotes gastric cancer invasion and metastasis by targeting tumor suppressor EPB41L3. Mol Cancer Res. 2011;9:824-33
9. Kanda M, Kodera Y. Recent advances in the molecular diagnostics of gastric cancer. World J Gastroenterol. 2015;21:9838-52
10. Yang D, Hendifar A, Lenz C, Togawa K, Lenz F, Lurje G. et al. Survival of metastatic gastric cancer: Significance of age, sex and race/ethnicity. J Gastrointest Oncol. 2011;2:77-84
11. Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57-70
12. Stamenkovic I. Matrix metalloproteinases in tumor invasion and metastasis. Semin Cancer Biol. 2000;10:415-33
13. Kasper G, Glaeser JD, Geissler S, Ode A, Tuischer J, Matziolis G. et al. Matrix metalloprotease activity is an essential link between mechanical stimulus and mesenchymal stem cell behavior. Stem Cells. 2007;25:1985-94
14. Vu TH, Werb Z. Matrix metalloproteinases: effectors of development and normal physiology. Genes Dev. 2000;14:2123-33
15. Kalluri R. Basement membranes: structure, assembly and role in tumour angiogenesis. Nat Rev Cancer. 2003;3:422-33
16. Liotta LA, Kohn EC. The microenvironment of the tumour-host interface. Nature. 2001;411:375-9
17. Kessenbrock K, Plaks V, Werb Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell. 2010;141:52-67
18. Sheu BC, Lien HC, Ho HN, Lin HH, Chow SN, Huang SC. et al. Increased expression and activation of gelatinolytic matrix metalloproteinases is associated with the progression and recurrence of human cervical cancer. Cancer Res. 2003;63:6537-42
19. Chan OTM, Furuya H, Pagano I, Shimizu Y, Hokutan K, Dyrskjot L. et al. Association of MMP-2, RB and PAI-1 with decreased recurrence-free survival and overall survival in bladder cancer patients. Oncotarget. 2017;8:99707-21
20. Shi Y, An D, Liu Y, Feng Q, Fang X, Pan G. et al. Propoxur enhances MMP-2 expression and the corresponding invasion of human breast cancer cells via the ERK/Nrf2 signaling pathway. Oncotarget. 2017;8:87107-23
21. Bae IH, Park MJ, Yoon SH, Kang SW, Lee SS, Choi KM. et al. Bcl-w promotes gastric cancer cell invasion by inducing matrix metalloproteinase-2 expression via phosphoinositide 3-kinase, Akt, and Sp1. Cancer Res. 2006;66:4991-5
22. Noh S, Jung JJ, Jung M, Kim TS, Park CH, Lim SJ. et al. MMP-2 as a putative biomarker for carcinomatosis in gastric cancer. Hepato-gastroenterology. 2011;58:2015-9
23. Raymond CK, Roberts BS, Garrett-Engele P, Lim LP, Johnson JM. Simple, quantitative primer-extension PCR assay for direct monitoring of microRNAs and short-interfering RNAs. Rna. 2005;11:1737-44
24. Xiong Y, Fang JH, Yun JP, Yang J, Zhang Y, Jia WH. et al. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology. 2010;51:836-45
25. Zhao ZS, Wang YY, Chu YQ, Ye ZY, Tao HQ. SPARC is associated with gastric cancer progression and poor survival of patients. Clin Cancer Res. 2009;16:260-8
26. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281-97
27. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215-33
28. Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A. 2006;103:2257-61
29. Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I. et al. E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell. 2008;13:272-86
30. Wang T, Hou J, Li Z, Zheng Z, Wei J, Song D. et al. miR-15a-3p and miR-16-1-3p Negatively Regulate Twist1 to Repress Gastric Cancer Cell Invasion and Metastasis. Int J Biol Sci. 2017;13:122-34
31. Devarajan P, Johnston JJ, Ginsberg SS, Van Wart HE, Berliner N. Structure and expression of neutrophil gelatinase cDNA. Identity with type IV collagenase from HT1080 cells. J Biol Chem. 1992;267:25228-32
32. Fang JH, Zhou HC, Zeng C, Yang J, Liu Y, Huang X. et al. MicroRNA-29b suppresses tumor angiogenesis, invasion, and metastasis by regulating matrix metalloproteinase 2 expression. Hepatology. 2011;54:1729-40
33. Ghosh A, Dasgupta D, Ghosh A, Roychoudhury S, Kumar D, Gorain M. et al. MiRNA199a-3p suppresses tumor growth, migration, invasion and angiogenesis in hepatocellular carcinoma by targeting VEGFA, VEGFR1, VEGFR2, HGF and MMP2. Cell death & disease. 2017;8:e2706
34. Chen X, Ruan A, Wang X, Han W, Wang R, Lou N. et al. miR-129-3p, as a diagnostic and prognostic biomarker for renal cell carcinoma, attenuates cell migration and invasion via downregulating multiple metastasis-related genes. J Cancer Res Clin Oncol. 2014;140:1295-304
35. Wang HY, Tu YS, Long J, Zhang HQ, Qi CL, Xie XB. et al. SRF-miR29b-MMP2 axis inhibits NSCLC invasion and metastasis. Int J Oncol. 2015;47:641-9
36. Ni X, Xia T, Zhao Y, Zhou W, Wu N, Liu X. et al. Downregulation of miR-106b induced breast cancer cell invasion and motility in association with overexpression of matrix metalloproteinase 2. Cancer science. 2014;105:18-25
37. Zou Y, Li J, Chen Z, Li X, Zheng S, Yi D. et al. miR-29c suppresses pancreatic cancer liver metastasis in an orthotopic implantation model in nude mice and affects survival in pancreatic cancer patients. Carcinogenesis. 2015;36:676-84
38. Xu Q, Li P, Chen X, Zong L, Jiang Z, Nan L. et al. miR-221/222 induces pancreatic cancer progression through the regulation of matrix metalloproteinases. Oncotarget. 2015;6:14153-64
Author contact
![]() Corresponding author: Prof. Jinchun Xing, PhD, The First Affiliated Hospital of Xiamen University, Xiamen, Fujian, 361003, Tel/Fax: (86-592) 213-9314, Email: xmcua2007com or Prof. Jianchun Cai, PhD, Zhongshan Hospital affiliated to Xiamen University, Xiamen, China 361004, Tel/Fax: (86-592) 229-2799, Email: jianchunfh2com
Corresponding author: Prof. Jinchun Xing, PhD, The First Affiliated Hospital of Xiamen University, Xiamen, Fujian, 361003, Tel/Fax: (86-592) 213-9314, Email: xmcua2007com or Prof. Jianchun Cai, PhD, Zhongshan Hospital affiliated to Xiamen University, Xiamen, China 361004, Tel/Fax: (86-592) 229-2799, Email: jianchunfh2com

 Global reach, higher impact
Global reach, higher impact