3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2017; 8(1):146-151. doi:10.7150/jca.16108 This issue Cite
Research Paper
Low Incidence along with Low mRNA Levels of EGFRvIII in Prostate and Colorectal Cancers Compared to Glioblastoma
1. Department of Research and Development, Celther Polska, Ltd., Milionowa 23, 93-193 Lodz, Poland;
2. Department of Tumor Biology, Medical University of Lodz, Zeligowskiego 7/9, 90-752 Lodz, Poland;
3. Clinical Department of Neurosurgery, The Voivodal Specialistic Hospital in Olsztyn, Zolnierska 18, 10-561 Olsztyn, Poland.
Received 2016-5-9; Accepted 2016-10-15; Published 2017-1-1
Abstract
Background: The presence as well as the potential role of EGFRvIII in tumors other than glioblastoma still remains a controversial subject with many contradictory data published. Previous analyses, however, did not consider the level of EGFRvIII mRNA expression in different tumor types. Methods: Appropriately designed protocol for Real-time quantitative reverse-transcription PCR (Real-time qRT-PCR) was applied to analyze EGFRvIII and EGFRWT mRNA expression in 155 tumor specimens. Additionally, Western Blot (WB) analysis was performed for selected samples. Stable cell lines showing EGFRvIII expression (CAS-1 and DK-MG) were analyzed by means of WB, immunocytochemistry (ICC) and fluorescence in situ hybridization (FISH). Results: Our analyses revealed EGFRvIII expression in 27.59% of glioblastomas (8/29), 8.11% of colorectal cancers (3/37), 6.52% of prostate cancers (3/46) and none of breast cancers (0/43). Despite the average relative expression of EGFRvIII varying greatly among tumors of different tissues (approximately 800-fold) or even within the same tissue group (up to 8000-fold for GB), even the marginal expression of EGFRvIII mRNA can be detrimental to cancer progression, as determined by the analysis of stable cell lines endogenously expressing the oncogene.
Conclusion: EGFRvIII plays an unquestionable role in glioblastomas with high expression of this oncogene. Our data suggests that EGFRvIII importance should not be underestimated even in tumors with relatively low expression of this oncogene.
Keywords: EGFRvIII, prostate cancer, colorectal cancer, breast cancer, glioblastoma, Real-time quantitative reverse-transcription PCR.
Introduction
Type III epidermal growth factor receptor (EGFRvIII) is a common mutation of EGFR [1]. According to the current state of knowledge, EGFRvIII is tumor-specific, ligand independent and constitutively active receptor. Moreover, it might contribute towards more cancerous phenotype, drug resistance and thus is considered an attractive anticancer therapy target [1, 2]. EGFRvIII was confirmed to be expressed in patients with glioblastoma (GB) [3, 4]. On the other hand, there have been many contradictory reports on its presence in other tumor types [2, 5]. Previous research focused only on EGFRvIII detection and did not include any quantitative analysis [6, 7]. Therefore, we decided to analyze not only the occurrence but also the level of EGFRvIII expression in glioblastomas, prostate, breast and colorectal tumors as well as unique EGFRvIII-positive glioblastoma cell lines - CAS-1, DK-MG, and DK-MG subline with low EGFRvIII expression (DK-MGlow). For the first time the relative and absolute EGFRvIII expression level was compared between different tumor types.
Materials and Methods
Tumor samples
Surgical specimens were obtained from 46 patients diagnosed with prostate cancer (Pabianice Medical Center; Mikolaj Pirogow Regional Specialist Hospital in Lodz), 43 with breast cancer (Polish Mother's Health Center Research Institute in Lodz), 37 with colorectal cancer (Antoni Troczewski Local Government Hospital in Kutno, Clinical Hospital Military Memorial Medical Academy - Central Veterans' Hospital in Lodz) and 29 with glioblastoma (The Voivodal Specialistic Hospital in Olsztyn). All samples were collected according to the protocol approved by the Bioethical Committee at the Regional Medical Chamber in Lodz (Approval No. K.B. - 3/12 of February 8, 2012) and by the Bioethical Committee of Medical University of Lodz (Approval No. RNN/27/11/KE). Written informed consent was obtained from all patients and their data were processed and stored according to the principles described in the Declaration of Helsinki. Patients were diagnosed according to the World Health Organization Criteria.
Cell lines
DK-MG cell line (DSMZ, Germany) and its subline (DK-MGlow) were obtained and cultured as previously described by us [8]. CAS-1 cell line (ICLC, Italy) was cultured in DMEM (PAN-Biotech GmbH, Germany) supplemented with 10% FBS (Biowest, France), 1% Penicillin-Streptomycin (Gibco, France), 0.2% Gentamicin Sulfate (Biowest, France), maintained in 5% CO2 at 37°C and passaged with trypsin-EDTA (0.05% Trypsin, Gibco, France). Serial dilution in a 96-well plate format was employed to perform clonal selection of CAS-1 cell line.
Real-time qRT-PCR for EGFRvIII and EGFRWT
RNA was isolated using AllPrep RNA/DNA Mini Kit (Qiagen, Germany) according to the manufacturer's instructions. Isolation was performed on tissue specimens of 30-40 mg and 4-6 mm in diameter, with approximate RNA yield of 100 ng/µL. For each sample 250 ng of total RNA was reverse-transcribed into single-stranded cDNA using QuantiTect Reverse Transcription Kit (Qiagen, Germany) according to the manufacturer's protocol. To compare EGFRvIII expression level between different tissue samples, equal amounts of cDNA (20 ng) were analyzed in Real-time qRT-PCR reaction using StepOnePlus Real-Time PCR System (Applied Biosystems). PCR products were synthesized from cDNA samples using SYBR® Select Master Mix. TBP and HPRT1 genes were used as reference to normalize expression level of target genes. Primer sequences for TBP gene were 5'-GAGCTGTGATGTGAAGTTTCC-3', 5'-TCTGGGTTTGATCATTCTGTAG-3' while 5'-TGAGGATTTGGAAAGGGTGT-3', 5'-GAGCACACAGAGGGCTACAA-3' were used to amplify HPRT1 gene. The following specific primers were used for amplification of target genes: 5'-TAGCAGTCTTATCTAACTATGAT-3', 5'-CACTGCTGACTATGTCCCGC-3' for EGFRWT and 5' GGCTCTGGAGGAAAAGAAAGGTAATTATGT-3', 5' ACCAATACCTATTCCGTTACACACT-3' for EGFRvIII.
Normalized relative EGFRWT or EGFRvIII expression level in tested samples versus control sample was calculated utilizing the method described by Pfaffl et al. [9, 10], based on each sample's average Ct value and each gene's average PCR efficiency. cDNA from CAS-1 and DK-MG cell lines was used as a positive control of EGFRvIII and EGFRWT expression. BJ human neonatal foreskin fibroblasts (ATCC, USA) provided a negative control for EGFRvIII detection.
To generate cDNA pool from EGFRvIII-positive samples, mRNA isolated from 15 tumor samples expressing the mutated receptor was pooled and diluted 50 times. Specimens were classified as EGFRvIII-positive or -negative depending on generation of reaction product at cycle threshold (Ct) ≤ 37. The expression of EGFRvIII was normalized using pool cDNA, while EGFRWT was normalized using cDNA from fibroblasts. A fold change of more than 2 was considered as overexpression, while between 1 and 2 as expression within normal range [11].
The absolute quantification of EGFRvIII expression level within different tissues was performed according to standard curve method [12]. The number of EGFRvIII transcripts in tested samples was extrapolated from a standard curve based on Ct values obtained for dilution series of plasmid encoding EGFRvIII.
DK-MG and CAS-1 cell lines analyses
Immunocytochemistry (ICC) and Western Blotting (WB) for EGFR protein expression as well as FISH for EGFR copy number detection were performed as previously described by us [8, 13]. Additionally, WB analyses were performed for selected tumor specimens (with initial homogenization step - mechanical disruption of liquid nitrogen frozen sample). Phospho-WB for DK-MG cell line was performed as described previously with the use of rabbit anti-phospho-EGFR (Tyr1068) antibody (Cell Signaling Technology, Inc., Cat. No. 2234; 1 : 500) [8].
Results
EGFRvIII presence and EGFRvIII and EGFRWT expression level in glioblastomas as well as prostate, breast and colorectal tumors
Real-time qRT-PCR analysis of 155 specimens revealed EGFRvIII expression in 14 tumor samples (28.39%), which constituted 9.03% of tested group. All specimens considered EGFRvIII-positive generated reaction product at Ct ≤ 37. Glioblastomas expressed EGFRvIII in 27.59% (8 out of 29 samples), while this variant was detected in three out of 37 colorectal tumors (8.11%) and three out of 46 prostate tumors (6.52%). All of 43 analyzed breast cancer samples were EGFRvIII-negative. A maximal thousandfold difference in relative EGFRvIII expression level was detected between samples (Table 1). The absolute number of EGFRvIII mRNA molecules varied remarkably between different specimens, still the general trend between relative and absolute values was followed (Table 1, Figure 1B).
Intriguingly, glioblastomas were divided into two groups with high and low EGFRvIII expression (100-fold difference on average) (Figure 1). These groups varied also in terms of EGFRWT expression, which was inversely proportional to the levels of EGFRvIII. Interestingly, expression of the oncogenic variant in non-CNS EGFRvIII-positive samples (prostate cancer and colon cancer) and glioblastoma specimens with low EGFRvIII expression was comparable (Figure 1A).
Comparison of relative EGFRvIII and EGFRWT mRNA expression levels and absolute copy number of EGFRvIII cDNA in EGFRvIII-positive samples by means of Real-time quantitative PCR.
| Sample name | Relative gene expression ± SD | Absolute number of EGFRvIII molecules | |
|---|---|---|---|
| EGFRWT | EGFRvIII | ||
| PC25 | 1.823 ± 0.062 | 0.053 ± 0.007 | 77.706 |
| PC33 | 2.283 ± 0.063 | 0.015 ± 0.014 | 20.382 |
| PC46 | 2.689 ± 0.079 | 0.844 ± 0.047 | 2935.646 |
| CC12 | 1.239 ± 0.255 | 0.019 ± 0.008 | 14.835 |
| CC13 | 1.257 ± 0.006 | 0.034 ± 0.015 | 8.708 |
| CC30 | 0.690 ± 0.078 | 0.024 ± 0.006 | 2.473 |
| GB16 | 11.435 ± 0.770 | 0.233 ± 0.033 | 28.609 |
| GB20 | 3.769 ± 0.062 *** | 21.441 ± 1.898 *** | 1730.176 |
| GB28 | 42.482 ± 7.466 | 38.299 ± 7.538 *** | 660833.331 |
| GB31 | 3.137 ± 0.436 *** | 67.149 ± 9.368 *** | 672855.128 |
| GB34 | 21.895 ± 0.666 *** | 0.020 ± 0.004 | 11.694 |
| GB45 | 105.487 ± 14.168 *** | 0.024 ± 0.019 | 18.747 |
| GB46 | 35.256 ± 0.886 | 1.820 ± 0.086 | 6887.642 |
| GB49 | 3.851 ± 0.625 | 0.008 ± 0.002 | 6.035 |
| DK-MG | 1.931 ± 0.272 | 16.288 ± 1.816 *** | 6392.209 |
| DK-MGlow | 2.877 ± 0.196 | 0.806 ± 0.030 | 265.193 |
| CAS-1 | 0.937 ± 0.011 | 0.786 ± 0.023 | 1467.265 |
| human fibroblasts | 1 ± 0.005 | 0 | 0 |
Note: Statistical significance calculated by one-way ANOVA with Tukey's post-comparison test; ***, p < 0.001; PC - prostate cancer; CC - colorectal cancer; GB - glioblastoma. Absolute number of molecules given per 20 ng cDNA.
Comparison of A. the average relative EGFRvIII and EGFRWT mRNA expression levels and B. the average number of EGFRvIII molecules per 20 ng of cDNA between two glioblastoma groups (with high and low EGFRvIII expression), prostate cancer and colon cancer. Error bars indicate SEM. Statistical significance calculated by one-way ANOVA with Tukey's post-comparison test; ***, p < 0.001, ns, not significant.
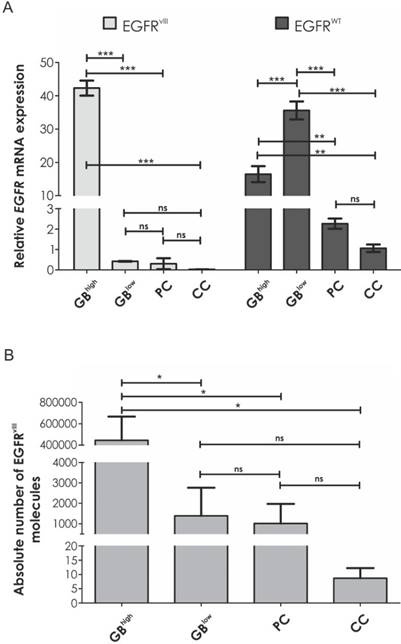
Considering EGFRvIII-positive samples, Real-time qRT-PCR analysis demonstrated on average 20 times higher EGFRWT expression level in comparison to tumors other than GB (p < 0.001) and at least three times higher in GB specimens when compared to fibroblasts (8/8, ranging from 3.137 to 105.487).
It is worth mentioning that EGFRvIII was undetectable at protein level in two out of three tumors with comparably low level of this oncogene mRNA expression. In our analysis, the threshold of EGFRvIII detection was 0.8 (Figure 2A, Table 1).
Correlation between EGFRvIII mRNA level and protein expression in CAS-1, DK-MG and DK-MGlow cells
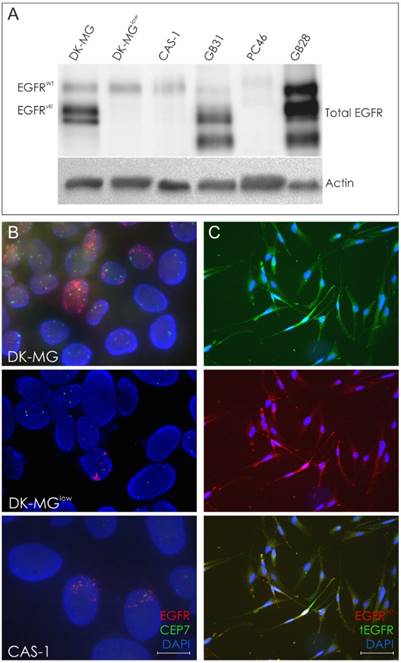
We established DK-MG subline with a minimum content of EGFRvIII-positive cells at 5% (DK-MGlow) [8]. FISH analysis of CAS-1 cell line revealed numerous extrachromosomal amplicons, in a similar staining pattern to the one observed for DK-MG line, in approximately 1% of cells (Figure 2B). Furthermore, double ICC staining with antibodies against total as well as wild-type EGFR confirmed presence of the truncated receptor and indicated the fraction of EGFRvIII-positive cells to be around 1% (Figure 2C). In case of CAS-1, an attempt to derive a population with smaller fraction of EGFRvIII-expressing cells by means of clonal selection was unsuccessful.
Molecular analyses in glioblastoma cell lines and selected tumor specimens. A. Western Blot analysis of DK-MG, DK-MGlow, CAS-1 cell lines and selected EGFRvIII-positive tumor specimens. B. FISH analysis of DK-MG, DK-MGlow and CAS-1 cell lines. The number of cells positive for EGFRvIII amplicons varies among these lines; magnification 600x. C. Representative images of EGF-mediated degradation of EGFRWT following 1h EGF treatment in CAS-1 cell line, demonstrated as intracellular dots. Cells presenting high cell membrane expression of total EGFR under these conditions are likely to be EGFRvIII-positive.
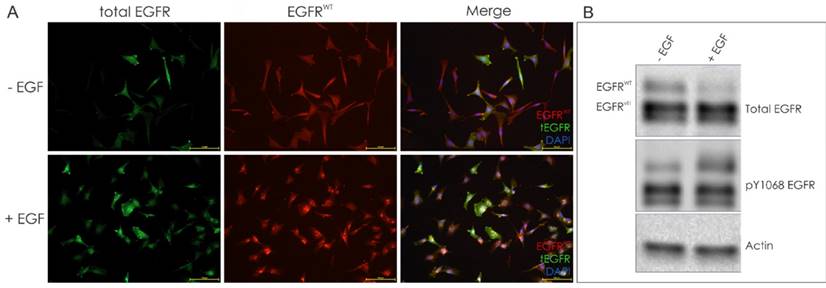
The fraction of EGFRvIII-positive cells in the DK-MG population was further confirmed when population of cells was stimulated with EGF, inducing degradation of the EGFRWT, observed as intracellular dots representing endocytosed protein (Figure 3A). The ligand-mediated degradation of the wild-type receptor, but not EGFRvIII, was confirmed by Western Blotting (Figure 3B).
It is important to note that results of ICC and FISH were consistent for CAS-1, DK-MG and DK-MGlow cells, in each case demonstrating similar percentage of cells with EGFRvIII expression and amplicons, respectively (Figure 2B, 2C and 3A).
Our data suggests, however, that in case of EGFRvIII there is no simple correlation between mRNA and protein level. In spite of the fact that both CAS-1 and DK-MGlow showed similar EGFRvIII mRNA expression, the percentage of EGFRvIII-positive cells detected by means of Immunocytochemistry differed between the lines and was undetectable with Western Blotting (Figure 2C), even on overexposed blots (data not shown). Hence, we suggest that Western Blotting may be insufficiently sensitive to detect the low protein level. Moreover, WB analysis of EGFRvIII in frozen tumor sections may give misleading results, due to possible detection of EGFRWT degradation products, which might appear similar in molecular weight to EGFRvIII (Figure 2C).
Discussion
The presence of EGFRvIII in glioblastoma is unquestionable [3, 4], while its occurrence in other tumor types still remains very controversial [6, 7, 14, 15, 16, 17]. There are no reports analyzing EGFRvIII mRNA level in a wider range of tumors, since previously it was considered satisfactory to classify tumor samples as EGFRvIII-positive or -negative.
Our work showed for the first time the relative levels of EGFRvIII mRNA in specimens of glioblastomas, as well as prostate, colorectal and breast cancer. Interestingly, those levels varied substantially between analyzed tumor types - several thousandfold differences were detected between specimens with the highest and lowest expression, even within glioblastoma cases. It has to be noted that in contrary to other tumor types, glioblastoma is resected with a minimal non-malignant tissue margin. Consequently, the substantial diversity in EGFRvIII expression levels and absolute number of molecules observed by us within the group of glioblastoma specimens (Table 1) cannot be explained solely by the variation in extension of surgical resection (resulting from differences in content of normal cells in analyzed specimens). Interestingly, all of the glioblastoma samples investigated by us can be subdivided into two subgroups, depending on the level of EGFRvIII mRNA expression (Figure 1A). On the other hand, prostate and colorectal cancers showed relatively low expression of EGFRvIII mRNA and low frequency of this alteration when compared to glioblastoma.
Considering other analyzed tumor types, no breast cancer specimen expressing EGFRvIII was identified, which is in accordance with Rae et al. [16] and contradicts the data published by Del Vecchio et al. and Silva et al. [18, 19]. Therefore, it may be suggested that previous analyses overestimated the number of cases showing EGFRvIII expression in tumors other than glioblastoma [6, 15, 16].
Our choice of Real-time qRT-PCR as a method to investigate EGFRvIII expression in tumors of different origin has been dictated predominantly by reports on relatively low specificity of anti-EGFRvIII antibodies, low sensitivity and semi-quantitative nature of Western blot or limited use of DNA-based methods in assessment of high copy number of genes encoded by amplicons [15, 20-22]. Still, we are aware that percentage of normal cells in tumor specimens, percentage of tumor cells showing EGFRvIII expression, number of EGFRvIII amplicons per cell, as well as number of EGFRvIII mRNA molecules per cell remain variables that may hinder the interpretation of Real-time qRT-PCR results.
Despite the variables described above, the strong discrepancies in mRNA expression levels, even within the same tumor type, suggest different roles or modes of action the oncogene can employ to induce tumor progression. EGFRvIII-positive cells have been described to constitute only a small fraction of the overall cancer cell population, however, they remain dispersed throughout the tumor tissue rather than forming cohesive lumps. Taking into account the reports on the EGFRvIII being implicated in inducing secretion of cytokines and growth factors associated with tumor progression [23-25], or the oncogene being secreted in the form of extracellular vesicles that merge with surrounding cells, it ensures that even very limited expression of EGFRvIII can contribute towards cancer progression [26]. It remains to be seen, whether different levels of mRNA correlate with any particular oncogenic action.
The important role of EGFRvIII in cancer cell progression is further supported by the study on stable cancer cell lines that endogenously express the oncogenic receptor. In case of DK-MG line we were able to reduce the number of EGFRvIII-positive cells below the 5% threshold of the population, but never to zero. Additionally, we were unable to decrease the number of EGFRvIII-positive cells in CAS-1 cell line. Taken together, our results suggest that the aforementioned models are fairly accurate representation of the tumor tissue and correspond with Nishikawa et al. who demonstrated that low percentage of EGFRvIII-positive cells is sufficient to maintain proper environment for glioblastoma progression [27]. Additionally, models characterized by low incidence of a particular mutation may be suitable especially for analysis of cancer stem cells hypothesis. In fact, EGFRvIII-positive cells have already been reported to demonstrate characteristic features of cancer stem cells [28].
With the use of Real-time qRT-PCR we demonstrated that prostate and colorectal cancer resections were positive for EGFRvIII expression at 6.52% and 8.11% incidence rate, respectively, albeit the relative expression level was low. In contrast, glioblastoma specimens (with incidence rate at 27.59%) varied greatly in respect to the relative and absolute number of mutant receptor's mRNA.
The correlation between varying levels of EGFRvIII mRNA and the reported elsewhere pro-oncogenic function of EGFRvIII remain to be elucidated. However, it is clear that even the lowest levels of the oncogenic receptor's expression might be sufficient for cancer progression.
EGF-mediated degradation of the wild-type EGFR allows for identification of EGFRvIII-positive cells. A. Representative images of DK-MG cells treated for 1h with EGF, as indicated. Intracellular dots represent EGFRWT undergoing endocytosis. Cells strongly positive for total EGFR signal are likely to be EGFRvIII-positive. B. Western blot analysis confirms degradation of the EGFRWT following ligand stimulation.
Abbreviations
CNS: Central Nervous System; EGFRvIII: Epidermal Growth Factor Receptor variant III; EGFRWT: Epidermal Growth Factor Receptor wild-type; GB: Glioblastoma; Real-time qRT-PCR: real-time quantitative reverse-transcription PCR
Acknowledgements
This work was supported by the Polish Agency for Enterprise Development (grant number POIG.01.04.00-10-037/11-00). Studies on glioblastoma samples were possible thanks to collaboration with the Department of Tumor Biology, Medical University of Lodz, Poland and supported by National Science Center (grant number 2012/05/B/NZ4/02623).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Pedersen MW, Meltorn M, Damstrup L. et al. The type III epidermal growth factor receptor mutation. Biological significance and potential target for anti-cancer therapy. Ann Oncol. 2001;12:745-60
2. Sok JC, Coppelli FM, Thomas SM. et al. Mutant epidermal growth factor receptor (EGFRvIII) contributes to head and neck cancer growth and resistance to EGFR targeting. Clin Cancer Res. 2006;12:5064-73
3. Ohgaki H, Kleihues P. Genetic pathways to primary and secondary glioblastoma. Am J Pathol. 2007;170:1445-53
4. Saikali S, Avril T, Collet B. et al. Expression of nine tumour antigens in a series of human glioblastoma multiforme: interest of EGFRvIII, IL-13Ralpha2, gp100 and TRP-2 for immunotherapy. J Neurooncol. 2007;81:139-48
5. Chau NG, Perez-Ordonez B, Zhang K. et al. The association between EGFR variant III, HPV, p16, c-MET, EGFR gene copy number and response to EGFR inhibitors in patients with recurrent or metastatic squamous cell carcinoma of the head and neck. Head Neck Oncol. 2011;3:11
6. Moscatello DK, Holgado-Madruga M, Godwin AK. et al. Frequent expression of a mutant epidermal growth factor receptor in multiple human tumors. Cancer Res. 1995;55:5536-9
7. Olapade-Olaopa EO, Moscatello DK, MacKay EH. et al. Evidence for the differential expression of a variant EGF receptor protein in human prostate cancer. Br J Cancer. 2000;82:186-94
8. Stec WJ, Rosiak K, Siejka P. et al. Cell line with endogenous EGFRvIII expression is a suitable model for research and drug development purposes. Oncotarget. 2016 http://dx.doi.org/10.18632/oncotarget.8201
9. Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29:e45
10. Pfaffl MW, Horgan GW, Dempfle L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 2002;30:e36
11. Lam AK, Lau KK, Gopalan V. et al. Quantitative analysis of the expression of TGF-alpha and EGFR in papillary thyroid carcinoma: clinicopathological relevance. Pathology. 2011;43:40-7
12. Larionov A, Krause A, Miller W. A standard curve based method for relative real time PCR data processing. BMC Bioinformatics. 2005;6:62
13. Stoczynska-Fidelus E, Piaskowski S, Bienkowski M. et al. The failure in the stabilization of glioblastoma-derived cell lines: spontaneous in vitro senescence as the main culprit. PLoS One. 2014;9:e87136
14. Wikstrand CJ, Hale LP, Batra SK. et al. Monoclonal antibodies against EGFRvIII are tumor specific and react with breast and lung carcinomas and malignant gliomas. Cancer Res. 1995;55:3140-8
15. Jungbluth AA, Jungbluth AA, Stockert E. et al. A monoclonal antibody recognizing human cancers with amplification/overexpression of the human epidermal growth factor receptor. Proc Natl Acad Sci. 2003;100:639-44
16. Rae JM, Scheys JO, Clark KM. et al. EGFR and EGFRvIII expression in primary breast cancer and cell lines. Breast Cancer Res Treat. 2004;87:87-95
17. Azuma M, Danenberg KD, Iqbal S. et al. Epidermal growth factor receptor and epidermal growth factor receptor variant III gene expression in metastatic colorectal cancer. Clin Colorectal Cancer. 2006;6:214-8
18. Silva HA, Abraúl E, Raimundo D. et al. Molecular detection of EGFRvIII-positive cells in the peripheral blood of breast cancer patients. Eur J Cancer. 2006;42:2617-22
19. Del Vecchio CA, Jensen KC, Nitta RT. et al. Epidermal growth factor receptor variant III contributes to cancer stem cell phenotypes in invasive breast carcinoma. Cancer Res. 2012;72:2657-71
20. Gupta P, Han SY, Holgado-Madruga M. et al. Development of an EGFRvIII specific recombinant antibody. BMC Biotechnol. 2010;10:72
21. Gan HK, Cvrljevic AN, Johns TG. The epidermal growth factor receptor variant III (EGFRvIII): where wild things are altered. FEBS J. 2013;280:5350-70
22. Weller M, Kaulich K, Hentschel B. et al. Assessment and prognostic significance of the epidermal growth factor receptor vIII mutation in glioblastoma patients treated with concurrent and adjuvant temozolomide radiochemotherapy. Int J Cancer. 2013;134:2437-47
23. Bonavia R, Inda MM, Vandenberg S. et al. EGFRvIII promotes glioma angiogenesis and growth through the NF-κB, interleukin-8 pathway. Oncogene. 2012;31:4054-66
24. Ramnarain DB, Park S, Lee DY. et al. Differential gene expression analysis reveals generation of an autocrine loop by a mutant epidermal growth factor receptor in glioma cells. Cancer Res. 2006;66:867-74
25. Inda MD, Bonavia R, Mukasa A. et al. Tumor heterogeneity is an active process maintained by a mutant EGFR-induced cytokine circuit in glioblastoma. Genes Dev. 2010;24:1731-45
26. Al-Nedawi K, Meehan B, Micallef J. et al. Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nat Cell Biol. 2008;10:619-24
27. Nishikawa R, Sugiyama T, Narita Y. et al. Immunohistochemical analysis of the mutant epidermal growth factor, deltaEGFR, in glioblastoma. Brain Tumor Pathol. 2004;21:53-6
28. Emlet DR, Gupta P, Holgado-Madruga M. et al. Targeting a glioblastoma cancer stem-cell population defined by EGF receptor variant III. Cancer Res. 2014;74:1238-49
Author contact
![]() Corresponding author: Joanna Peciak, e-mail: joanna.peciakcom.
Corresponding author: Joanna Peciak, e-mail: joanna.peciakcom.

 Global reach, higher impact
Global reach, higher impact