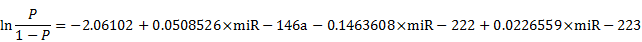Impact Factor
ISSN: 1837-9664
J Cancer 2017; 8(1):48-56. doi:10.7150/jca.16644 This issue Cite
Research Paper
Identification of A Panel of Serum microRNAs as Biomarkers for Early Detection of Lung Adenocarcinoma
Department of Laboratory Medicine, Shanghai Chest Hospital, Shanghai Jiao Tong University, Shanghai 200030, China.
* These authors contributed equally to this work.
Received 2016-6-29; Accepted 2016-9-18; Published 2017-1-1
Abstract
Introduction: Since currently no sensitive and specific biomarkers for early detection of lung adenocarcinoma (AD) exist and the majority of AD patients are diagnosed at late stages of disease, the development of effective screening tests for early-stage lung AD is urgently needed. Serum microRNAs (miRNAs) have been documented as novel noninvasive biomarkers in tumor diagnosis; thus, we studied the profile of serum miRNA in AD patients in order to identify the differentially expressed miRNAs as potential biomarkers for early detection of AD.
Patients and Methods: Serum samples were collected from 180 AD patients and 180 age- and sex-matched healthy controls. Serum miRNA profiling was performed by low-density array (LDA) using RNA extracted from blood samples of 20 patients and 20 controls. To validate the selected miRNAs, a stem-loop based RT-qPCR assay was used and serum samples from 160 patients and 160 controls were examined.
Results: Profiling data showed 11 differentially expressed miRNAs in the serum samples from AD patients compared with the controls. Among them, 6 selected miRNAs in AD patients, including miR-103, miR-146a, miR-151, miR-21, miR-221, miR-222, and miR-223, were validated by RT-qPCR. In particular, the top three, miR-146a, miR-222, and miR-223, were confirmed to be significantly expressed in stage I/II AD patients compared with healthy controls.
Conclusion: A panel of miRNAs with miR-146a, miR-222 and miR-223 could be used as potential noninvasive biomarkers for early detection of AD.
Keywords: Screening - Low-density array - Receiver operating characteristic (ROC) curve - Area under the ROC curve (AUC) - Logistic regression.
Introduction
Lung cancer is the most common malignancy in humans and the leading cause of cancer-related deaths worldwide [1-3]. Lung cancer is clinically classified into the following two types, small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). NSCLC accounts for 85-90% of all lung cancer cases [4], which is further histologically subdivided into four categories, including adenocarcinoma (AD), squamous cell carcinoma (SQ), large cell carcinoma (LC) and 'others'. Of those, lung AD is the most common histological type of NSCLC and is responsible for more than 500,000 deaths globally every year [5]. The incidence of AD is dramatically increasing in Chinese and East Asian populations, especially in females and never smokers [6, 7]. Moreover, the five-year survival rate of patients with AD is approximate 15%.
The survival and prognosis of patients with AD are closely associated with tumor stage at first diagnosis. Most of patients with AD are diagnosed at late stages of disease due to the lack of effective methods for early detection, leading to high mortality rates. Imaging and cytology-based screening strategies have been employed for early detection of tumors and are demonstrated to be sensitive tools for several types of tumors, but neither has shown to be sufficiently effective in either early detection for lung cancer or significant reduction of AD mortality [8, 9]. Therefore, development of effective screening tests for early-stage lung cancer, through the identification of sensitive biomarkers, is urgently needed.
miRNAs are small non-coding RNAs 19-24 nucleotides in length that negatively regulate gene expression at posttranscriptional levels. miRNAs can influence a wide variety of cellular functions, including differentiation and apoptosis, and have been implicated in tumor genesis according to various posttranscriptional regulation mechanisms [10-12]. Mounting evidence supports that miRNAs play very important roles during tumor initiation as well as tumor progression, migration, invasion, and metastasis [13-17]. Recent studies revealed that miRNAs are stably present in serum. The expression patterns of serum miRNAs are closely correlated with progression and prognosis in a variety of human tumors, including lung cancer, colorectal cancer, prostate cancer, gastric cancer, osteosarcoma, and so on [18-24]. Circulating miRNAs could be considered as a novel promising diagnostic and prognostic biomarker for cancer screening because: (ⅰ) serum miRNAs are readily detectable by RT-qPCR, a technique that widely used in clinical laboratories; (ⅱ) blood-based biomarkers are minimally invasive for the screening of high-risk subjects and early diagnosis of cancer.
The possible use of blood-based biomarkers for NSCLC has been evaluated by several studies. Roth et al. reported serum miR-361-3p and miR-625* as blood-based markers for discriminating malignant lung tumors from benign ones [25]. Another study conducted by Shen et al. showed that four miRNAs, including miRNA-21, miRNA-126, miRNA-210 and miRNA-486-5p yielded 86.22% sensitivity and 96.55% specificity in distinguishing NSCLC patients from healthy controls [26]. Recently, Dou H et al. found that plasma let-7c and miR-152 of NSCLC patients are significantly reduced compared with healthy control, and the AUC of let-7c and miR-152 was 0.714 and 0.845, respectively, indicating that the two miRNAs could be promising diagnostic biomarkers for NSCLC screening [21].
However, the serum miRNA signature in patients with AD, a major type of NSCLC, has not been clearly identified. To identify miRNA-based noninvasive biomarkers that are capable of distinguishing AD patients from healthy individuals, we evaluated the expression of serum miRNAs in 180 Chinese patients with AD and 180 healthy controls by LDA (Exqion) and RT-qPCR. The diagnostic ability of differentiated expressed miRNAs was further evaluated by the AUC according to international association for the study of lung cancer (IASLC) stage system.
Patients and Methods
Patients and control individuals
All samples were collected from consenting individuals according to protocols approved by the ethics committee of Shanghai Chest Hospital. In this study, we recruited 180 primary AD patients who were treated in the Shanghai Chest Hospital and 180 control individuals. To identify serum miRNAs as promising markers for lung AD, a multiphase and case-control study was designed. First, to identify differentially expressed miRNAs between AD patients and matched controls, a miRNA-profiling array that includes a total of 766 miRNAs (miRBase 19.0) was initially performed by Exqion high-throughput PCR using pooled serum samples from 20 AD patients. Subsequently the validation of serum miRNA signature in AD patients was accomplished by a stem-loop RT-qPCR assay using the serum samples from 160 patients with AD and 160 controls. This part was subdivided into two phases: (a) a biomarker-selection phase, in which serum samples from 40 AD patients formed the training set and (b) a biomarker-validation phase, in which serum samples from the other 120 AD patients formed the validation set. All patients received a diagnosis of lung AD between 2014 and 2015 and their blood samples were collected before any therapeutic procedures, such as surgery, chemotherapy or radiotherapy. The final diagnosis of AD was made according to histopathological examination of surgical tumor resection and the tumor stage was defined by the operative findings. In the case where AD patients were unsuitable for surgical operation, their histopathology characteristics and tumor stage were confirmed by histobiopsy and imaging technology. Table 1 summarizes the demographic and clinical features of the patients. Cancer-free healthy controls were selected from a large pool of individuals seeking a routine health checkup at the Shanghai Chest Hospital. Those control individuals were matched to the patients by age, sex, and ethnicity.
Demographic and clinical features of AD patients and healthy subjects.
Variable | Training set | Validation set | p-value | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AD | Control | p-value | AD | Control | p-value | AD in | ||||||||
| (n=40) | (n=40) | (AD vs. | (n=120) | (n=120) | (AD vs. | training vs. | ||||||||
| No. | % | No. | % | control) | No. | % | No. | % | control) | validation) | ||||
| Average age (years) | 60.1±7.9 | 60.2±8.0 | 0.967 | 59.9±8.0 | 59.4±8.3 | 0.631 | 0.673 | |||||||
| Age (years) | ≤60 | 20 | 50% | 21 | 52.5% | 0.823 | 60 | 50% | 60 | 50.0% | 1.000 | 1.000 | ||
| >60 | 20 | 50% | 19 | 47.5% | 60 | 50% | 60 | 50.0% | ||||||
| Female | 20 | 50% | 20 | 47.5% | 58 | 48.3% | 59 | 49.2% | 0.855 | |||||
| Differentiation | Poor | 3 | 7.5% | 6 | 5% | 0.251 | ||||||||
| Moderate | 18 | 45% | 59 | 49.2% | ||||||||||
| High | 19 | 47.5% | 46 | 38.3% | ||||||||||
| Unknown | 0 | 0% | ` | 9 | 7.5% | |||||||||
| Stage | Ⅰ | 26 | 65% | 72 | 60% | 0.522 | ||||||||
| Ⅱ | 10 | 25% | 31 | 25.8% | ||||||||||
| Ⅲ | 4 | 10% | 10 | 8.3% | ||||||||||
| Ⅳ | 0 | 0% | 0 | 0% | ||||||||||
| Unknown | 0 | 0% | 7 | 5.8% | ||||||||||
Sample processing and RNA extraction
Sera from 20 patients with non-metastatic AD, or 20 controls with similar age and sex distributions, were used for the high-throughput PCR. Equal amounts (250 μL) of sera were combined to form two pools from cases and controls. miRNA was extracted from each serum pool using a mirVana PARIS Kit (Ambion, Grand Island, NY, USA), according to the manufacturer's instructions. The resulting RNA pellet was dissolved in 100 μL Elution Solution and stored at -80 °C until further analysis. For the RT-qPCR assay, miRNA was extracted from 200 μL serum of each sample with the mirVana PARIS Kit.
High-throughput PCR array
Low-density array was performed by Exiqon high-throughput PCR array (Exiqon, Denmark) to examine the expression of 766 miRNAs in serum circulating RNA of 40 subjects, (20 patients with AD and 20 healthy subjects).
Quantification of miRNAs by RT-qPCR
Reverse transcription with 7 μL of RNA extracted from serum sample was performed using PrimeScript RT Enzyme System, following the manufacturer's protocol (TaKaRa, Japan). cDNA obtained from the RT reaction was diluted 1:10 with RNase-free water. The PCR reaction was performed with gene-specific primers whose sequences are described in (Supplementary Material) using SYBR Green I PCR Master Mix (TaKaRa, Japan) in the Roche 480 Real-Time PCR System (Roche, Switzerland). Each reaction was performed in a 20 μL volume containing 2 μL cDNA, 3 μL forward primer, 1.4 μL universal reverse primer and 10 μL Power SYBR Green I PCR Master Mix. The thermal cycling profile for PCR was set up as follows: pre-denaturation at 95°Cfor 5 min, followed by 40 cycles of denaturation for 15 s at 95°C, and annealing and extension for 1 min at 60 °C. A melt curve analysis was performed at the end of each PCR cycle to validate the PCR product specificity. The detection limits and dynamic range of the RT-qPCR assay were assessed with data analyzed. Meanwhile, the absolute concentration of target miRNA was calculated using a calibration curve established with the corresponding synthetic miRNA oligonucleotide (Supplementary Material). Each reaction was carried out in triplicate.
Statistical analysis
Statistical analysis was performed with Stata software (version12.0; StataCorp). Data are presented as median with inter-quartile range (IQR). The Mann-Whitney U, a non-parametric statistical test, was used to compare the differences between the serum miRNA level of cancer patients and that of healthy controls. A P-value < 0.05 was considered statistically significant. For each target miRNA, we established a Receiver operating characteristic (ROC) curve to calculate the area under the ROC curve (AUC) for evaluating both specificity and sensitivity of the prediction test of AD. The correlation between AD and serum miRNA level was determined by logistic regression analysis.
Results
Identification of serum miRNA signature in AD patients
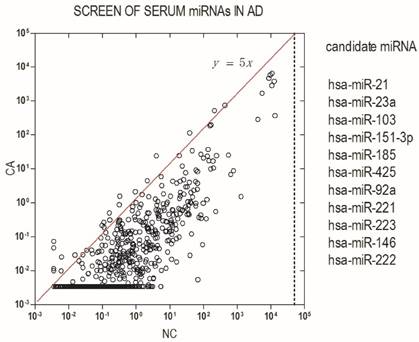
To identify differentially expressed miRNA in AD patients, we initially screened pooled serum from 20 patients and 20 healthy donors for the expression of 766 miRNAs using an Exqion LDA. The results showed that most miRNAs are down-regulated in AD patients as compared with health subjects. A differentially expressed miRNA was considered if their expression levels showed at least a 5-fold difference between AD patients and the control groups and had a Threshold cycle (Cp) value ≤ 35. Based on the above criteria, 11 differentially expressed miRNAs were found to be candidate miRNAs in our study (Fig.1).
Six miRNAs were differentially expressed in a large crowd of AD patients
We validated the expression level of 11 candidates of differentially expressed miRNAs that were selected from the initial screen. These differentially expressed miRNAs were confirmed by a stem-loop based RT-qPCR assay using serum samples from 160 patients with a clinical and pathologic diagnosis of AD and from 160 healthy control individuals. There were no significant differences between the cancer patients and control individuals in age distribution or sex (Table 1).
The measurement of the level of serum miRNAs using RT-qPCR proved to be reliable and reproducible, because semi logarithmic plots of the calibration curves for various levels of the synthetic single-strand miRNA calibrators were linear from 1 fmol/L to 100 pmol/L (Fig. S1, A-J). Cp values of replicate assays were very similar (r2 = 0.990), indicating that the RNA extraction method was reproducible. We eliminated a candidate miRNA, hsa-miR-92a, because it exhibited non-liner amplification.
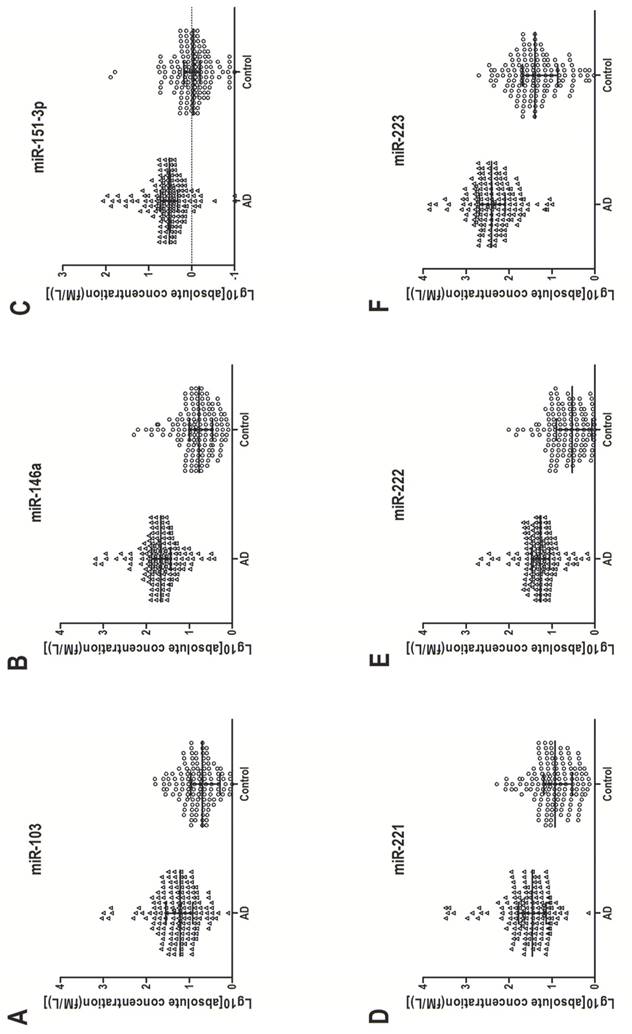
In the training set, individual serum samples from 40 AD patients and 40 healthy controls were used for miRNA screening, in which only miRNAs with a mean change ≥ 1.5-fold and a P-value ≤ 0.005 were selected for further analysis. According to these criteria, six differentially expressed miRNAs, including miR-103, miR-146a, miR-151-3p, miR-221, miR-222 and miR-223, were discovered (Table 2). These six miRNAs were further examined by RT-qPCR in a larger cohort consisting of 120 AD patients and 120 matched controls. Consistent with the results from the training set, the serum levels of the above six miRNAs in AD patients were significantly higher than those in healthy controls. The increased range in those serum miRNAs was from 3.46-fold to 11.59-fold (Table 2). Figure 2 shows the differences in serum levels of the six differentially expressed miRNAs between 160 AD patients and 160 control individuals enrolled in both training and validation sets.
miRNA concentrations in AD serum samples and control samples in the training and validation sets.
| Training set | Validation set | |||||||
|---|---|---|---|---|---|---|---|---|
| AD | Control | Fold | AD | Control | Fold | |||
| (n=40) | (n=40) | change | p-value | (n=120) | (n=120) | change | p-value | |
| miR-103 | 17.20(27.70) | 9.61(13.99) | 1.79 | <0.0005 | 17.15(27.46) | 4.96(5.47) | 3.46 | <0.0001 |
| miR-146a | 35.20(46.20) | 5.20(6.81) | 6.77 | <0.0001 | 53.00(47.30) | 6.98(6.85) | 7.60 | <0.0001 |
| miR-151-3p | 3.86(3.22) | 1.19(1.37) | 3.25 | <0.0001 | 3.29(3.14) | 0.91(0.86) | 3.62 | <0.0001 |
| miR-221 | 31.50(51.47) | 12.56(12.56) | 2.51 | <0.0005 | 33.10(54.00) | 6.76(11.28) | 4.90 | <0.0001 |
| miR-222 | 14.30(14.29) | 7.07(8.07) | 2.02 | <0.0003 | 21.70(19.80) | 3.76(6.19) | 5.78 | <0.0001 |
| miR-223 | 171.00(195.10) | 31.80(53.60) | 5.38 | <0.0001 | 295.00(391.00) | 25.45(38.20) | 11.59 | <0.0001 |
Annotations: The unit for miRNA concentrations is fM/L. The values before parentheses represent average of the miRNA concentrations, and the values in parentheses represent the inter-quartile range.
Identification of serum miRNA signature in AD. Serum miRNA signature was identified by a screening test using high-throughput PCR assay. Differentially expressed miRNAs were considered according to at least a 5-fold difference between AD patients and control groups and a Cp value ≤ 35.
Up-regulation of six serum miRNAs in AD. Serum concentration of the six miRNAs was measured in 160 AD patients and 160 healthy control individuals (in both the training and validation sets) with a stem-loop-based RT-qPCR assay. Cp values were converted to absolute values from the calibration curves. Each point represents the median of results for triplicate samples.
Three of the six miRNAs were optimized as a serum biomarker for AD screening
To further evaluate the diagnostic value of the screening test for AD, we examined the sensitivity and specificity of the test using the above mentioned six differentially expressed miRNAs (Figure S2). On the basis of the AUC values, a logistic regression model with the best prediction was defined as following:
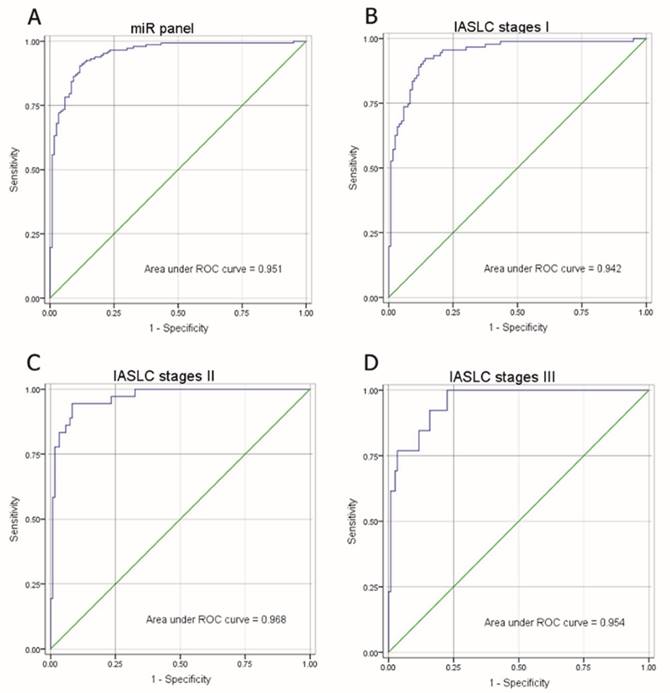
As shown in Figure 3A, the composite panel of the above three miRNAs (miR-146a, miR-222 and miR-223) produced 0.951 AUC (95% CI, 0.92605-0.97644). Using an optimal cutoff value (p = 0.5015), the sum of the sensitivity (84.35%) and specificity (90.83%) was maximal for this prediction model. The analysis confirmed the effectiveness of this miRNA panel with high accuracy (87.27%) in discriminating AD from healthy subjects.
The three miRNAs was associated with the stages of AD patients
The correlation of the miRNA panel and AD stages according to IASLC Lung Cancer stages (Seventh Edition), a proposal for the TNM classification for lung cancer, was also evaluated (Fig. 3B-D). The corresponding AUCs for patients with IASLC Lung Cancer stages I, II and III were 0.942, 0.968, and 0.954, respectively, as compared to healthy subjects. These results suggest that even at an early stage of AD (stage I), this miRNA panel is still able to discriminate AD patients from healthy subjects.
The three miRNAs was specifically correlated with AD
Finally, we evaluate the specificity of the miRNA panel for AD: squamous cell carcinoma (SQ), lung inflammation (INF) and small cell lung cancer (SCLC) are used as other lung disease control (Fig. S3). As the result, the serum miRNA panel level of the patient with AD was significantly higher than other lung disease.
Discussion
Previous studies revealed that miRNAs are stably present in plasma and serum, exhibiting distinctive miRNA expression profiles in patients with various tumors, including lung cancer, colorectal cancer, prostate cancer and pancreatic cancer [18, 19, 27]. These findings implicate the potentials of circulating miRNAs as novel noninvasive biomarkers for cancer screening. However, no such markers have been identified yet for lung AD, and there are none of effective screening test for early-stage of lung AD till nowadays, neither. So identification of sensitive biomarkers for early detection of lung cancer is urgently needed. Therefore, we aimed to identify the circulating miRNA signatures for early diagnosis of AD, the most common type of lung cancer with a high incidence in Chinese and East Asian populations, especially in females and never smokers [6, 7].
In the current study, to profiled the expression of serum miRNAs in Chinese patients with AD, we developed a reliable strategy that combining techniques of LDA for screening the candidate miRNAs using pooled serum samples and RT-qPCR assay for validation using the other solus serum samples from 160 AD patients and 160 healthy controls. As the result, from the cohort of 180 AD patients and 180 healthy controls, we found that six serum miRNAs, including miR-103, miR-146a, miR-151-3p, miR-221, miR-222 and miR-223 were ubiquitously aberrantly up-regulated in AD patients. Our findings imply the possibility of those differentially expressed miRNAs to be used as serum miRNA signature in AD patients. Interestingly, the expression level of miR-146a, miR-222, and miR-223 in stage I/II AD patients was significantly up-regulated compared with that in healthy controls and other lung disease controls, suggesting these three miRNAs can be used as noninvasive biomarkers with high-sensitivity and high-specificity for screening tests of high-risk subjects and early-stage AD patients.
Moreover, the majority of the aberrantly expressed miRNAs identified in the serum samples from AD patients are well-documented cancer-related miRNAs. For instance, miR-222 is known as one of the most up-regulated miRNAs in TRAIL-resistant non-small cell lung cancer cells [28], and it has been confirmed that miRNA-222 elevates in the serum of patients with NSCLC [29]. miR-223 overexpression in serum has also been observed in the patients with NSCLC [30]. Moreover, miR-223 has been found up-regulated in tumor samples of gastrointestinal carcinomas, including gastric, colon, and pancreatic cancer [31, 32]. Additionally, consistent with our results, miR-146a has also been reported to be present in the serum of NSCLC patients [33], but its function in lung cancer remains unknown. Given that miR-146a is shown to promote cell proliferation and colony formation in NIH/3T3 cells [34], it might play some potent roles in the development of lung cancer. To further investigate the function of serum miRNAs in AD and explore the mechanisms underlying regulation of serum miRNA expression, more intensive studies are need in the future.
ROC of miRNA panel comprised of miR-146a, miR-222 and miR-223 by logistic regression analysis. (A) ROC of miRNA panel AD group from the healthy group, (B) IASLC stage I group from the healthy group, (C) IASLC stage II group from the healthy group, and (D) IASLC stage III group from the healthy group.
Additionally, several miRNAs (such as miR-155, miR-566 and miR-939) have been reported as serum biomarker of AD in previous studies, but they haven't been detected in our research. Gao et al. [35] found miR-155 highly expressed in the serum of AD patients compared with the healthy control. However, miR-155 was just 2.034 fold-difference expressed between AD patients and healthy control. In the serum miRNA signature identification stage of our research, miRNA expression levels showed at least a 5-fold difference between AD patients and the control groups could be considered as a differentially expressed miRNA. Maybe that's the reason why we didn't detected miR-155. In another research, Rani et al.[36] found miR-566 and miR-939 significantly highly expressed in AD patients compared with healthy groups. Considering the 4 stages of AD, they found the level of these two miRNAs increase in stage II compared with stage I, but substantially decreased in stage III. Since we performed the biomarker-selection by Exqion high-throughput PCR using pooled serum samples from 20 AD patients and 20 healthy controls, respectively, we suspect the serum from the stage III AD patients diluted the serum pool, decreasing the level of miR-566 and miR-939. As a result, we didn't detected there two miRNAs.
According to the reports published in recent years, extracellular/circulating miRNAs have been detected in a variety of biological fluids. These miRNAs clearly possess many properties of ideal biomarkers, such as stability in nuclease-rich body fluids, unique sequences, and tissue-specific expression. Accumulating data strongly suggests there are three types of membranous vesicles that contain extracellular miRNA: apoptotic bodies, shedding vesicles, and exosomes [37-39]. Besides those vesicles, extracellular miRNA can also be vesicle-free and associated with Argonaute (AGO) proteins alone or be incorporated into high-density lipoprotein(HDL) particles [40-42]. There are two theories regarding the export and function of circulating miRNA: (i) they are merely by-products of routine microvesicle secretion and cell death and (ii) they are specifically secreted and functional in intercellular communication. Although either could be true, further studies are still needed to reinforce the theory on circulating miRNAs as functional signaling molecules in vivo [43].
In conclusion, a panel of miRNAs including miR-146a, miR-222 and miR-223 has both enough sensitivity and specificity to distinguish AD from health control and other lung disease. It could be used as potential noninvasive biomarkers for early detection and a complementary of routine screening strategy for AD.
Supplementary Material
Supplementary figures.
Acknowledgements
We thank our colleagues and technicians for their great assistance and support. The study was supported by The National Natural Science Foundation of China (81372193), The project of National Science and technology cooperation (2014DFA33010), The Scientific Research project of Shanghai Science and Technology Committee (14411950800), Feasible scheme of the Scientific Research project of Shanghai Science and Technology Committee (13JC1405502), The major project of science and technology development foundation of Shanghai Chest Hospital (2014YZDC10100), and The Municipal hospital clinical auxiliary department capacity building project of Shanghai Shen Kang Hospital Development Center (SHDC22014011).
Authorship and contributors
Dr. Shaogang Lv and Jian Xue performed the research, collected data, analyzed the data and wrote the paper. Dr. Chuanyong Wu, Lin Wang, Jing Wu and Shujun Xu, performed the research and analyzed the data. Dr. Xiaohui Liang contributed important reagents and collected data. Dr. Jiatao Lou designed the research, analyzed the data and wrote the paper.
Ethics
The study was approved by the Ethics Committee of Shanghai Jiao Tong University, China.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Jemal A, Siegel R, Xu J. et al. Cancer statistics, 2010. CA: a cancer journal for clinicians. 2010;60:277-300
2. Jemal A, Bray F, Center MM. et al. Global cancer statistics. CA: a cancer journal for clinicians. 2011;61:69-90
3. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA: a cancer journal for clinicians. 2013;63:11-30
4. Herbst RS, Heymach JV, Lippman SM. Lung cancer. The New England journal of medicine. 2008;359:1367-1380
5. Imielinski M, Berger AH, Hammerman PS. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell. 2012;150:1107-1120
6. Toh CK, Gao F, Lim WT. et al. Never-smokers with lung cancer: Epidemiologic evidence of a distinct disease entity. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 2006;24:2245-2251
7. Subramanian J, Govindan R. Lung cancer in never smokers: A review. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 2007;25:561-570
8. Gavelli G, Giampalma E. Sensitivity and specificity of chest x-ray screening for lung cancer: Review article. Cancer. 2000;89:2453-2456
9. Bach PB, Kelley MJ, Tate RC. et al. Screening for lung cancer: A review of the current literature. Chest. 2003;123:72s-82s
10. Coskun E, von der Heide EK, Schlee C. et al. The role of microrna-196a and microrna-196b as erg regulators in acute myeloid leukemia and acute t-lymphoblastic leukemia. Leukemia research. 2011;35:208-213
11. Li Y, Guessous F, Zhang Y. et al. Microrna-34a inhibits glioblastoma growth by targeting multiple oncogenes. Cancer research. 2009;69:7569-7576
12. Ding J, Huang S, Wu S. et al. Gain of mir-151 on chromosome 8q24.3 facilitates tumour cell migration and spreading through downregulating rhogdia. Nature cell biology. 2010;12:390-399
13. Bartel DP. Micrornas: Genomics, biogenesis, mechanism, and function. Cell. 2004;116:281-297
14. Garzon R, Marcucci G, Croce CM. Targeting micrornas in cancer: Rationale, strategies and challenges. Nature reviews Drug discovery. 2010;9:775-789
15. Lv H, Zhang Z, Wang Y. et al. Microrna-92a promotes colorectal cancer cell growth and migration by inhibiting klf4. Oncology research. 2016;23:283-290
16. Gu JJ, Zhang JH, Chen HJ. et al. Microrna-130b promotes cell proliferation and invasion by inhibiting peroxisome proliferator-activated receptor-gamma in human glioma cells. International journal of molecular medicine. 2016;37:1587-1593
17. Zhou YW, Zhang H, Duan CJ. et al. Mir-675-5p enhances tumorigenesis and metastasis of esophageal squamous cell carcinoma by targeting reps2. Oncotarget. 2016
18. Chen X, Ba Y, Ma L. et al. Characterization of micrornas in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008;18:997-1006
19. Mitchell PS, Parkin RK, Kroh EM. et al. Circulating micrornas as stable blood-based markers for cancer detection. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:10513-10518
20. Wang H, Wang L, Wu Z. et al. Three dysregulated micrornas in serum as novel biomarkers for gastric cancer screening. Medical oncology (Northwood, London, England). 2014;31:298
21. Dou H, Wang Y, Su G. et al. Decreased plasma let-7c and mir-152 as noninvasive biomarker for non-small-cell lung cancer. International journal of clinical and experimental medicine. 2015;8:9291-9298
22. Hur K, Toiyama Y, Okugawa Y. et al. Circulating microrna-203 predicts prognosis and metastasis in human colorectal cancer. Gut. 2015
23. Wang X, Ning Y, Yang L. et al. Diagnostic value of circulating micrornas for osteosarcoma in asian populations: A meta-analysis. Clinical and experimental medicine. 2016
24. Wang Q, Zhang H, Shen X. et al. Serum microrna-135a-5p as an auxiliary diagnostic biomarker for colorectal cancer. Annals of clinical biochemistry. 2016
25. Roth C, Stuckrath I, Pantel K. et al. Low levels of cell-free circulating mir-361-3p and mir-625* as blood-based markers for discriminating malignant from benign lung tumors. PloS one. 2012;7:e38248
26. Shen J, Todd NW, Zhang H. et al. Plasma micrornas as potential biomarkers for non-small-cell lung cancer. Laboratory investigation; a journal of technical methods and pathology. 2011;91:579-587
27. Wang J, Chen J, Chang P. et al. Micrornas in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer prevention research (Philadelphia, Pa). 2009;2:807-813
28. Garofalo M, Romano G, Di Leva G. et al. Egfr and met receptor tyrosine kinase-altered microrna expression induces tumorigenesis and gefitinib resistance in lung cancers. Nature medicine. 2012;18:74-82
29. Mao KP, Zhang WN, Liang XM. et al. Microrna-222 expression and its prognostic potential in non-small cell lung cancer. TheScientificWorldJournal. 2014;2014:908326
30. Sanfiorenzo C, Ilie MI, Belaid A. et al. Two panels of plasma micrornas as non-invasive biomarkers for prediction of recurrence in resectable nsclc. PloS one. 2013;8:e54596
31. Volinia S, Calin GA, Liu CG. et al. A microrna expression signature of human solid tumors defines cancer gene targets. Proceedings of the National Academy of Sciences of the United States of America. 2006;103:2257-2261
32. Bloomston M, Frankel WL, Petrocca F. et al. Microrna expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. Jama. 2007;297:1901-1908
33. Wang RJ, Zheng YH, Wang P. et al. Serum mir-125a-5p, mir-145 and mir-146a as diagnostic biomarkers in non-small cell lung cancer. International journal of clinical and experimental pathology. 2015;8:765-771
34. Xu T, Zhu Y, Wei QK. et al. A functional polymorphism in the mir-146a gene is associated with the risk for hepatocellular carcinoma. Carcinogenesis. 2008;29:2126-2131
35. Gao F, Chang J, Wang H. et al. Potential diagnostic value of mir-155 in serum from lung adenocarcinoma patients. Oncology reports. 2014;31:351-357
36. Rani S, Gately K, Crown J. et al. Global analysis of serum micrornas as potential biomarkers for lung adenocarcinoma. Cancer biology & therapy. 2013;14:1104-1112
37. Valadi H, Ekstrom K, Bossios A. et al. Exosome-mediated transfer of mrnas and micrornas is a novel mechanism of genetic exchange between cells. Nature cell biology. 2007;9:654-659
38. Hunter MP, Ismail N, Zhang X. et al. Detection of microrna expression in human peripheral blood microvesicles. PloS one. 2008;3:e3694
39. Zernecke A, Bidzhekov K, Noels H. et al. Delivery of microrna-126 by apoptotic bodies induces cxcl12-dependent vascular protection. Science signaling. 2009;2:ra81
40. Arroyo JD, Chevillet JR, Kroh EM. et al. Argonaute2 complexes carry a population of circulating micrornas independent of vesicles in human plasma. Proceedings of the National Academy of Sciences of the United States of America. 2011;108:5003-5008
41. Turchinovich A, Weiz L, Langheinz A. et al. Characterization of extracellular circulating microrna. Nucleic acids research. 2011;39:7223-7233
42. Vickers KC, Palmisano BT, Shoucri BM. et al. Micrornas are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nature cell biology. 2011;13:423-433
43. Turchinovich A, Weiz L, Burwinkel B. Extracellular mirnas: The mystery of their origin and function. Trends in biochemical sciences. 2012;37:460-465
Author contact
![]() Corresponding author: Jiatao Lou, MD, Department of Laboratory Medicine, Shanghai Chest Hospital, 241 West Huaihai Road, Shanghai, 200030, China. Tel.: +86-21-2220000-1503, Fax: +86-21-62808279, E-mail: loujiataocom.
Corresponding author: Jiatao Lou, MD, Department of Laboratory Medicine, Shanghai Chest Hospital, 241 West Huaihai Road, Shanghai, 200030, China. Tel.: +86-21-2220000-1503, Fax: +86-21-62808279, E-mail: loujiataocom.

 Global reach, higher impact
Global reach, higher impact