3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2016; 7(15):2333-2340. doi:10.7150/jca.15812 This issue Cite
Research Paper
An In Vitro Inverted Vertical Invasion Assay to Avoid Manipulation of Rare or Sensitive Cell Types
1. Department of Biomedical Engineering, University of Minnesota - Twin Cities;
2. Stem Cell Institute, University of Minnesota - Twin Cities;
3. Masonic Cancer Center, University of Minnesota - Twin Cities;
4. Lillehei Heart Institute, University of Minnesota - Twin Cities;
5. Institute for Engineering in Medicine, University of Minnesota - Twin Cities;
6. Department of Biology, Jackson State University.
Received 2016-4-11; Accepted 2016-8-18; Published 2016-11-26
Abstract
The ability to quantify cell migration and invasion is critical in the study of cancer metastasis. Current invasion assays, such as the Boyden Chamber, present difficulties in the measurement of the invasion of cells that are few in number and are intrinsically tied to the cell microenvironment. There exists a need for a three-dimensional invasion assay that is easily reproduced, accessible for most laboratories, and requires no displacement of cells from their original microenvironment. Here we present a simple design for an inverted vertical invasion assay able to assess the invasion capabilities of cells in a three dimensional, extracellular matrix-based environment without displacement from the original culture location. We used the assay to determine the migratory capacity of hybrids between mesenchymal/multipotent stem/stroma cells (MSCs) and breast cancer cells MCF7. These hybrids are formed reliably but rarely (1 in 1,000 cells) and for this reason require an invasion assay that does not involve extensive cell manipulation. Using this assay, we found that MSCs, breast cancer cells, and corresponding fusion products are able to migrate and invade through the extracellular matrix and that hybrids invade in a manner more similar to stromal cells than cancer cells. Thus, this assay can aid the study of the invasive capacity of both cancerous cells and associated fusion hybrids and could augment testing of therapeutic strategies to inhibit metastatic spread.
Keywords: invasion, hybrids, metastasis, 3D assay, collagen.
Introduction
Approximately 90% of cancer-related deaths are caused by local invasion and distant metastasis of tumor cells. To successfully relocate in the body cancer cells must invade local tissues by degrading extracellular matrix (ECM) components and ultimately, transverse the basement membrane. Once in circulation, these cells can form metastatic colonies at secondary locations. Thus, increased invasion and migration capabilities of cancer cells often indicate increased metastatic potential [1]. Different assays have been designed to quantify the metastatic capability of tumor cells. These assays include wound healing or scratch assay, time-lapse microscopy, and the transwell or Boyden Chamber assay. In the body, cells exist in a 3D ECM environment rich in type I collagen. The frequently used wound healing assay and time lapse microscopy are two dimensional systems and do not closely mimic the in vivo environment especially with respect to cell adhesion and associated cell motility [2-7]. The more physiologically relevant transwell or Boyden Chamber assay, which is a 3D system, requires the cells to be removed from their original environment and seeded on a layer of ECM in the upper chamber of the transwell. The cells then invade through the gel into a lower chamber containing a chemoattractant [6]. This technique, although valuable, appears challenging and unsuitable for cells susceptible to the microenvironment and/or significantly limited in number. As one example cell hybrids, formed as result of fusion between cancer cells and cells of the tumor microenvironment, are rare and are significantly influenced by the local microenvironment.
In previous studies including ours, hybrids arising from fusion between cancer cells and cells of the tumor microenvironment (mesenchymal stem/stroma cells, macrophages) have been proposed to contribute to tumor metastasis [8-18]. In particular, hybrids might acquire the migratory capability of the stroma cell parent and the proliferative property of the cancer cell parent leading to dissemination and new tumor growth at a distant site. However, traditional cell-based assays are not suitable to quantify the migration and invasion capability of hybrids since hybrids are vulnerable to the microenvironment in culture and the pool of hybrid cells is very small occurring at a frequency of 1 in 1000 cells [18] or less. These features of hybrids have hindered the study of their role in the development of metastases. Therefore, designing of a customized assay to quantify migration and invasion capability of hybrids is imperative. This assay should function at a per cell scale and should limit disruption to the cell microenvironment. In order to fulfill these criteria, we have developed an inverted vertical invasion assay. Using this newly developed assay, we efficiently analyzed the migration and invasion capacity of fusion products and parental lines. This assay could be used in different laboratories to study other complex cell types or to screen for pharmacological agents affecting cell migration and invasion.
Materials and Methods
Cell lines and culture
To optimize and validate our inverted vertical invasion assay design, we used MSCs and the breast cancer cells MDA-MB-231 and MCF7.
MSCs were a generous gift from Dr. Peiman Hematti (University of Wisconsin, Madison, WI, USA). They were derived from human embryonic stem cells in accordance with guidelines of the University of Wisconsin Institutional Review Board (Trivedi and Hematti, 2007) and maintained in α-minimum essential medium (Sigma-Aldrich, St. Louis, MO, USA), supplemented with 10% heat-inactivated fetal bovine serum (Hyclone, Logan, UT, USA). We reconfirmed the identity of the MSCs in our lab by flow cytometry for specific MSC markers, CD73, CD90 and CD105.
The human breast cancer cells MDA-MB-231 and MCF7 were obtained from American Type Culture Collection (ATCC; Manassas, VA, USA) and maintained in accordance with the recommendations of ATCC; the cells were not passaged for more than 6 months from the time of receipt. MDA-MB-231 cells were tested for: 1) mycoplasma by DNA stain and agar culture, 2) species determination by STR and COI assay, 3) sterility by BacT/ALERT 3D, and 4) the human pathogens. MCF7s were tested for: 1) mycoplasma by DNA stain and agar culture, 2) species determination by STR, 3) sterility by BacT/ALERT 3D, and 4) the human pathogens.
Transfection and Coculture Protocol. We used the BiFC approach to identify fusion products [18]. In brief, MSCs were transfected with the BiFC halve construct VNH3.1 and the breast cancer cells MCF7 were transfected with the corresponding BiFC halve construct YCH3.1. Transfected MSCs and MCF7 cells were cocultured in a MatTek dish at the ratio of 35,000 MSCs for 75,000 MCF7 cells. At this cell density, the surface area coverage for each cell type was approximately 50% at 24 h after the cocultures were initiated. The cocultures were initiated and maintained in α-MEM (Sigma-Aldrich, St. Louis, MO, USA) media supplemented with 10% heat inactivated defined FBS (Hyclone, Logan, UT, USA), 1% NEAA (Sigma-Aldrich, St. Louis, MO, USA), and 1% L-Glutamine (Sigma-Aldrich, St. Louis, MO, USA). At the end of the experiment, the cells were fixed, stained with DAPI and the fusion products were identified by fluorescence microscopy as previously described [18]. Figure 1C displays the timeline followed for this protocol.
Preparation of culture plates. The inverted vertical invasion assay was performed in MatTek 35 mm plates with 10 mm glass-bottom wells (MatTek P35G-1.5-10-C). Before initiating the cocultures, each plate was washed with 1 M hydrochloric acid to lower the hydrophobicity of the glass-bottom and to allow the collagen gel to adhere evenly to the surface. The plates were subsequently washed twice with phosphate buffered saline solution to remove any trace of acid, twice with 70% ethanol, and once with cell culture medium (Dulbecco's modified eagle's medium or α minimum essential medium).
Preparation of collagen gel. Collagen used for the inverted vertical invasion assay was prepared by combining on ice, rat tail collagen I (Corning, Corning, NY, USA), 10x DMEM (Thermo Fisher Scientific Inc., Waltham, MA, USA), and 1x cell culture medium containing heat inactivated fetal bovine serum (FBS) (Thermo Fisher Scientific Inc, Waltham, MA, USA) and the appropriate chemoattractant to a final concentration of collagen of 2.4 mg/ml. Sodium bicarbonate solution, 4.5%, was used to neutralize the collagen gel for its proper polymerization. Three chemoattractants were tested; FBS, platelet-derived growth factor BB (PDGF-BB), and insulin-like growth factor 1 (IGF-1). Eighty microliters of the prepared solution was laid over the cells within the well of MatTek dishes. The gel was allowed to polymerize at 37°C in a CO2 incubator with humidifier for thirty minutes, and then 2 mL of cell culture medium was added to the plate to cover the gel. The added medium used for experiments involving the chemoattractant IGF-1 or PDGF-BB contained 2% FBS, whereas it contained 10% FBS for experiments using FBS as the chemoattractant.
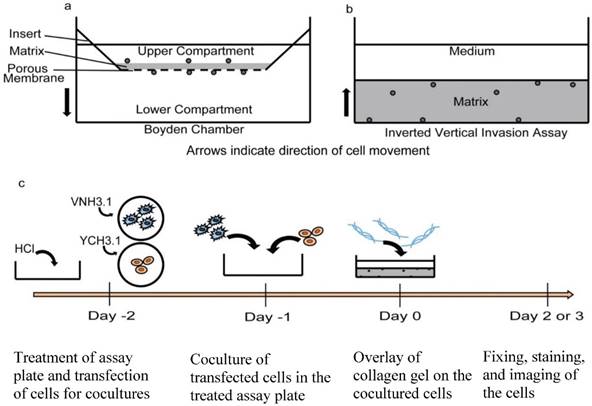
Invasion assay designs. a) Traditional Boyden Chamber or transwell invasion assay. In this assay, cells are collected from their original culture environment and plated onto the transwell insert. The arrow indicates the direction of cell movement during the assay. b) Inverted vertical invasion assay currently proposed. In this design, cells remain in their original microenvironment and the matrix protein is laid over the cell layer. The arrow indicates the direction of cell movement during the assay. c) Schematic detailing timeline of the inverted vertical invasion assay procedure. Day -2: MatTek dishes are treated with HCl and the two cell lines to be cocultured are transfected with the BiFC constructs as previously described [18]. If fused cells are not the focus of the invasion assay, the BiFC step is eliminated. Day -1: Transfected cells are cocultured on the treated MatTek dish. Day 0: Collagen gel mixture is made and laid over the cells. Day 2 or 3: cells are fixed, stained and subsequently imaged.
Imaging and assay analysis. The ability of cells to invade vertically was investigated at various time points during our study. At the end of each experiment, the cells were fixed with 4% paraformaldehyde for fifteen minutes and subsequently stained with a DAPI solution (100 ng/ml) at room temperature for twenty minutes. The fixed cells were imaged using a confocal fluorescent microscope (Olympus IX81; Olympus America, Center Valley, PA, USA). The total cell populations were imaged in the DAPI channel, while the GFP+ fusion products were imaged in the FITC channel. A fluorescence signal was considered positive if it was localized to the nucleus, fluoresced above background fluorescence of unaltered cocultures and did not fluoresce with excitation/emission at alternate wavelengths (Rhodamine, Cy5) [18]. Each plate was imaged in eight unique locations, taking 400 µm z-stack images of each location. Images were then processed using ImageJ software. The 3D image depicted in Figure 2C was captured after staining an assay of MSCs with DAPI and phalloidin. Images were taken using a mode-locked Ti:Sapphire multi-photon laser scanning microscope (MPLSM) (Prarie Technologies, Sioux Falls, SD, USA), with a 40x (NA 0.8) objective. 3D reconstructions were generated using Imaris 7.5.2 software (Bitplane Inc., Zurich, CH).
Results
Optimization of culture substrate for the inverted vertical invasion assay design
The inverted vertical invasion assay was optimized using a variety of cell culture dishes including 24-well plates, glass bottom chamber slides, and MatTek dishes. The 35 mm MatTek dish with a 10 mm glass bottom well in its center was found to be the most effective culture dish for the inverted vertical invasion assay (Fig. 1B). The MatTek dish has been used in a previous collagen based assay, the design of which was used as a basis for the inverted vertical invasion assay proposed in this study [19]. The advantages of this MatTek dish are threefold: 1) the thin glass bottom of the MatTek dish allows effective imaging and accurate assessment of cell position within the gel, 2) the 10 mm well within the MatTek dish restricts the gel to a thickness appropriate for oxygen diffusion and cell survival, and 3) most importantly the cultured cells can be analyzed within their microenvironment during the experiment (Fig 1B) and are not displaced as is the case in the Boyden Chamber assay (Fig 1A). The glass bottom chamber slides and the 24-well plates yielded uneven gel layers as a function of the meniscus formed at the edge of the well or chamber. The thickened gel at the edge of the wells limited gas and nutrient diffusion to the cells causing substantial cell death. In addition, the 24-well plates were not suitable for imaging due to the thickness of the plate and associated long working distance.
Cell viability and imaging potential within the MatTek dish
As a result of the MatTek design, the collagen gel layer of the assay was thin with even distribution. The concentration of collagen was 2.4 mg/mL and thus was thick enough to remain stable within the assay chamber during the course of the assay. The gels were thicker than the 200 μm nutrient diffusion limit in static conditions (350 +/- 75 μm). However, the wall height is substantially shorter than standard well plates and we predicted nutrient and gas exchange might occur readily from beneath and to the media-exposed sides of the gel. To test cell viability, especially in the gel center, we assessed cell morphology and numbers three days following gel polymerization. We found cells were able to survive and proliferate throughout the gel matrix. In particular, MCF7, MSCs, and the resulting cocultures maintained healthy morphologies to at least three days following initial gel formation (Fig 2A). In addition, the inverted design allowed for clear distinction of cells at distinct z planes within the collagen gel (Fig 2B, 2C). In this way, precise quantification of migratory distance into the gel could be assessed. We associate this migratory Z distance with in vitro invasive capacity.
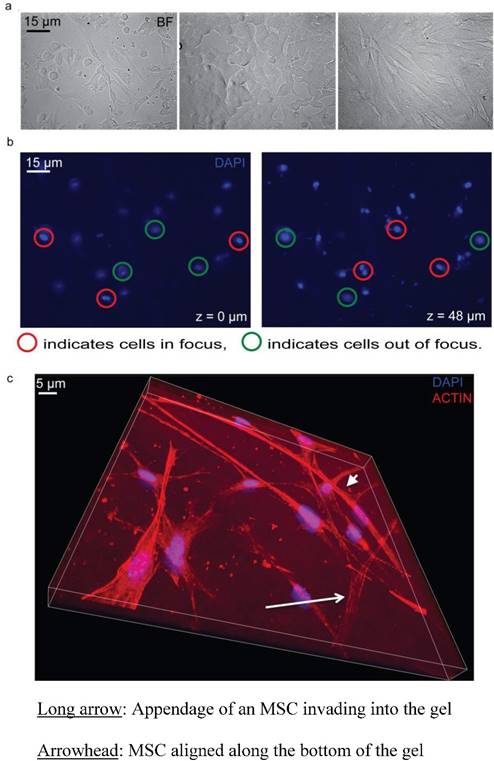
Cell viability and Z series of the inverted vertical invasion assay. a) 20x bright field images of MCF7 and MSC coculture, MCF7, and MSC (left to right) showing healthy cells under the collagen gel after two days, indicating that the assay environment maintained adequate conditions for cell survival. b) Representation of z series showing one location within a dish from the bottom (Z=0 µm) to the top (Z=48 µm). 3.8% FBS was used as a chemoattractant. Red circles indicate cells in focus. Green circles indicate cells out of focus. c) Representation of z series showing one location within a dish of MSCs from the bottom (Z=0 µm) to the Z=20 µm. 3.8% FBS was used as a chemoattractant. The long arrow indicates the appendage of an MSC invading into the gel. The arrowhead indicates an MSC, which is aligned along the bottom of the gel, and not invading. The blue shows the nuclear DAPI stain while the red shows the phalloidin stained actin filaments.
Reproducibility of the inverted vertical invasion assay
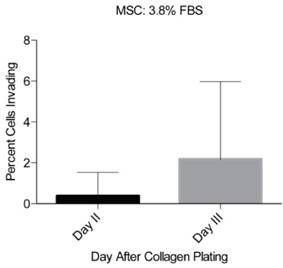
The reproducibility of the inverted vertical invasion assay was assessed by measuring the in vitro invasive capability of MSCs into the collagen gel in two separate experiments using 3.8% FBS as a chemoattractant. MSCs were previously shown to be highly migratory and invasive [20-23] and are therefore appropriate for testing of the reproducibility of our assay design. Each set of experiments consisted of three cultures of MSCs, fixed at days one, two, and three after collagen gel plating and the cells were imaged at eight different locations in each plate. No invasion was observed at day 1, little invasion was seen at day 2 (0.4 +/- 1.1% of the cells), and at day 3 substantial invasion was observed (2.2 +/- 3.8% of the cells) (Fig 3). However, due to the relatively low fraction invading cells, the variability appears large. To increase the percentage of cells able to invade the collagen gel, we tested stronger chemoattractants including PDGF- BB and IGF-1.
Optimization of chemoattractants in the inverted vertical invasion assay
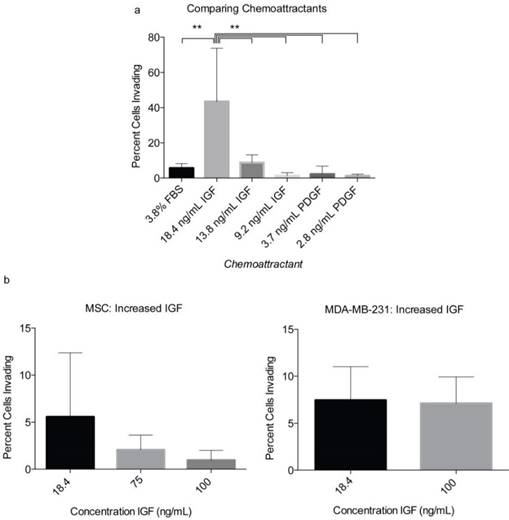
In order to optimize the level of observable invasion, the chemoattractants PDGF-BB and IGF-1 were tested at various concentrations within the collagen gel (Fig 4). The initial concentrations of IGF-1 and PDGF-BB were selected based on previously optimized concentrations used to achieve maximum MSC migration [24]. We found that IGF-1 at 18.6 ng/mL could induce up to 43.5% of MSCs to invade into the collagen gel. IGF-1 tested at 9.2, 75, and 100 ng/mL had a less substantial effect on MSC invasion into the collagen (Fig 4A-B). PDGF-BB produced levels of invasion similar to that of FBS and significantly less than that of IGF-1. At 2.8 ng/ml, PDGF-BB induced 1.21% of MSCs to invade into the collagen gel and at 3.1 ng/mL induced 6% MSC invasion (Fig 4A). Therefore, of the chemoattractants and concentrations used in the assay, IGF-1 at 18.6 ng/mL showed the most profound effect on MSC migration and invasion (Fig 4A). In order to confirm the chemoattractant effects of IGF-1 and further optimize the assay, the breast cancer cells MDA-MB-231 were tested in the assay. This breast cancer cell line is highly metastatic and characterized by elevated migratory and invasive ability [25, 26]. We observed that 8.4% of the MDA-MB-231 cells could invade into the gel in two days when IGF-1 was used at a concentration of 18.6 ng/mL (Fig 4B). A higher concentration of IGF-1 (100 ng/ml) did not increase the percentage of invading MDA-MB-231 cells (Fig 4B).
Reproducibility of the inverted vertical invasion assay. The percentage of invading MSCs was assessed in two separate experiments, in eight fields of view on days two and three following the collagen gel placement. While little invasion was seen on days one and two, day three showed substantial levels of invasion throughout the gel and in both experiments.
Optimization of chemoattractants using MSCs and MDA-MB-231. a) Various concentrations of PDGF-BB and IGF-1 were tested initially with MSCs. The percentage of invading MSCs was assessed 3 days following the layering of the collagen gel over the cells. b) After identifying IGF-1 as the more effective chemoattratant, its concentration was optimized using both MSCs and MDA-MB-231. Each number represents the average of the percentages of invading cells in eight unique locations within a plate. Experiments were performed in triplicate. A two-tailed unpaired ANOVA with Tukey's HSD post hoc test. **P < 0.01.
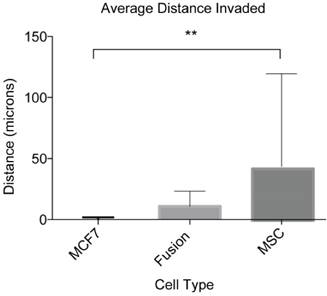
Invasion capability of cancer cell fusion products. We validated our assay by quantifying the invasion capacity of cancer cell fusion products. These numbers represent the average distance covered by four independent fusion products from three separate cocultures and their parental cell lines analyzed from eight different locations within each assay. This experiment was performed in triplicate. ANOVA with Tukey's HSD post hoc test. **P < 0.01.
Validation of the inverted vertical invasion assay using fusion products
The ultimate goal of this study was to develop an assay suitable for quantifying the migratory and invasive capabilities of rare and sensitive cell types including cancer cell fusion products. As a result of the optimizing experiments with MSCs and MDA-MB-231s, IGF-1 at 18.6 ng/mL was selected as the appropriate chemoattractant and the optimum concentration to investigate the in vitro invasive capacity of cancer cell fusion products. We looked specifically at hybrid cells arising from spontaneous fusion between MSCs and the nonmetastatic breast cancer cells MCF7. For this assay, MSCs were transfected with the BiFC halve construct VNH3.1 and MCF7 with the BiFC halve counterpart, YCH3.1. Cocultures were initiated and covered with collagen gel as described in materials and methods. Two days following the collagen overlay, the cells were fixed, stained with DAPI, and imaged. Z-stack image analysis of the cells showed that cancer cell fusion products were able to invade into the collagen gel to an average distance of 10.7 +/- 12.4 μm (Fig 5). The MCF7 cells showed an average invasion distance of 0.04 +/- 1.8 μm, and MSCs an average invasion distance of 41.3 +/- 76.0 μm (Fig 5). The difference in the invasion levels between MCF7 and MSCs was shown to be significantly different (P<0.01), while the fusion product invasion was shown to be insignificant when compared to both the MCF7 (P=0.85) and MSCs (P=0.21). These results indicate that our inverted vertical invasion assay not only is efficient and suitable to quantify in vitro invasive capacity of cancer cell fusion products, but also suggests that the fusion products have a higher migratory and invasive capability than the parental cancer cells, MCF7, as suggested by our previous study using 2-D time lapse microscopy [18].
Discussion
Although cell fusion has been proposed for many decades as a mechanism of cancer metastasis, relevant cell-based assays to investigate the metastatic potential of hybrids are still limited. Traditional cell-based invasion assays like the transwell or Boyden Chamber assay require excess manipulation of the cells that affect cell recovery and survival. While acceptable for many cell lines, such features of traditional assays make impossible the quantification of invasive qualities of sensitive and low frequency cell types including cancer cell hybrids. Therefore, there is a need to develop a cell-based assay, which avoids manipulation of cells and is suitable for cells, which are sensitive and/or limited in number. In this study, we developed and validated a collagen gel-based, inverted vertical invasion assay as a viable tool that can be used to quantify in vitro invasive capacity of cancer cell fusion products and other such scarce and sensitive cell types. The assay was shown to be cell-compatible and enabled easy identification and tracking of rare cell types.
We have shown here that while the inverted assay design results in easy and reliable cell invasion measurements, it does require a stronger chemoattractant than FBS. We observed that IGF-1 at a concentration of 18.6 ng/mL constitutes a sufficiently strong chemoattractant for the use of the assay with MSCs and cancer cells lines [27]. However, the use of other chemoattractants appropriate for the cell type being tested could be used in other studies. In the current experiment, for example, cytokines CSCL12/SDF-1α or CCL21/6Ckine could have been used, as both have shown to promote metastasis of breast cancer cell lines [28]. Furthermore, PDGF-BB may have been a sufficiently strong chemoattractant should we have further optimized the concentration. However, as IGF-1 was already found to promote sufficient invasion of the MSCs and MDA-MB-231s, this was not done. Chemoattractants could also be chosen in order to mimic a specific bodily environment. For example, osteonectin has been shown to promote the invasiveness of bone-metastasizing prostate carcinoma [29]. The inverted vertical invasion assay could be used to develop a model system of such organ-specific cancer metastasis. The need for a strong chemoattractant like IGF-1 in this assay may result from the preference of the cells for their environment on the glass-bottom of the well [1], and/or from the effect of gravity.
While producible and efficient itself, the major advantage of the inverted vertical assay is the circumvention of issues presented by the Boyden chamber assay and better control of the culture substrate. Cells are not displaced from the original cell culture environment, thus ensuring better control of cells that are intrinsically linked to the microenvironment. This is especially important for the case of cancer/stromal cell co-cultures, which are frequently used in order to mimic the tumor microenvironment. Studies that aim to quantify the invasiveness of the co-cultured cells in a 3D environment typically use a 3D spheroid assay, or displace the cells into Boyden chamber assay [30,31]. It has also been shown that the fate of stem cells is in part determined by previous culture substrate [21]. The inverted vertical invasion assay design that we propose here avoids the complications of a spheroid assay, can maintain the co-culture microenvironment, and better controls the culture substrate, all within a 3D environment.
While the inverted design does offer many benefits, there exist some drawbacks when compared to other 3D invasion assays. First, the MatTek dish design requires a relatively large amount of collagen to produce only a single invasion assay. For our purposes this was sufficient. However, in order to expand the versatility of the inverted assay design, it could be optimized for smaller culture wells, allowing for multiple simultaneous assays in a multi-well plate. Second, we found that there exists a need to balance the collagen gel strength while keeping the gel sufficiently thin for cell survival. This results in a collagen gel layer that is sensitive to large movements; thus, the assay must be handled delicately in order to prevent lifting of the gel. In addition to the small technical difficulties presented by the inverted assay design, it also provides less mimicry of the in vivo conditions compared to the dynamic system of microfluidic invasion assays. These assays provide a similar 3D environment as the inverted invasion assay, yet can maintain cells in an environment with flow similar to that of a bodily microenvironment [32]. While the static nature of the inverted invasion assay limits the degree to which it can mimic bodily microenvironments, it offers far greater simplicity and cost-effectiveness when compared to a microfluidic design.
The purpose of this study was to develop and standardize an assay that could be readily used to quantify the invasion capabilities of sensitive and rare cell types such as cancer cell fusion products. The need to develop this assay was highlighted by the lack of 3D systems that enable the quantification of cell invasion without affecting the integrity of the cell microenvironment [33,34]. And, although we used the inverted invasion assay to study the invasive capabilities of stem/cancer cell hybrids, invasion is a biological phenomenon that is not unique to cancer metastasis. Our proposed assay could also assist in the study of embryonic development, wound healing, and/or inflammatory responses, all of which require some form of cell migration/invasion [2]. The assay also could be customized to address many other questions including the effect of ECM engagement on cell invasion, the role of different chemoattractants on cell invasion, as well as the contribution of cancer stem cells or tumor circulating cells in cancer progression and metastasis. This assay could also be a useful tool for screening of pharmacological agents effective in inhibiting metastasis. Investigating these questions would not only increase understanding of cell interaction with the 3D environment, but could help in the design of therapeutic strategies to prevent metastatic spread.
Acknowledgements
The authors thank Dr. Peiman Hematti (Division of Hematology and Oncology, University of Wisconsin, Madison, WI, USA) for the gift of MSCs, Dr. Thomas Kerppola (Professor of Biological Chemistry, University of Michigan, Ann Arbor, MI, USA) for the gift of BiFC constructs, Dr. Philip Jung (University of Minnesota, Minneapolis, MN, USA) for help with ANOVA with Tukey's HSD post hoc test, the MPSCM, and the 3D image reconstruction. MPLSM was performed in the lab of Dr. Paulo Provenzano and the confocal microscope imaging in the lab of Dr. Patrick Alford (University of Minnesota, Minneapolis, MN, USA). This work was supported by U.S. Department of Defense, Idea Award 11-1-0205, the University of Minnesota Undergraduate Research Opportunities Program, and the National Institutes of Health (NIMHD-G12MD007581) through the RCMI-Center for Environmental Health at Jackson State University.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Minguell JJ, Erices A. Mesenchymal stem cells and the treatment of cardiac disease. Exp Biol Med. 2006;231(1):39-49
2. Kramer N, Walzl A, Unger C. et al. In vitro cell migration and invasion assays. Mutat Res. 2013;752(1):10-24
3. Birgersdotter A, Sandberg R, Ernberg I. Gene expression perturbation in vitro-a growing case for three-dimensional (3D) culture systems. Semin Cancer Biol. 2005;15(5):405-412
4. Weaver VM, Petersen OW, Wang F. et al. Reversion of the malignant phenotype of human breast cells in three-dimensional culture and in vivo by integrin blocking antibodies. J Cancer Biol. 1997;137(1):231-245
5. Bhadriraju K, Chen CS. Engineering cellular microenvironments to improve cell-based drug testing. Drug Discov Today. 2002;7(11):612-620
6. Valster A, Tran NL, Nakada M. et al. Cell migration and invasion assays. Methods. 2005;37(2):208-215
7. Baharvand H, Hashemi SM, Kazemi Ashtiani S, Farrokhi A. Differentiation of human embryonic stem cells into hepatocytes in 2D and 3D culture systems in vitro. Int J Dev Biol. 2006;50(7):645-652
8. Aichel O. Vortrage und Aufsatze uber Entvickelungsmechanik Der Organismen Leipzig. Wilhelm Engelmann. 1911
9. Pawelek JM, Chakraborty AK. The cancer cell-leukocyte fusion theory of metastasis. Adv Cancer Res. 2008;101:397-444
10. Chakraborty AK, Sodi S, Rachkovsky M. et al. A spontaneous murine melanoma lung metastasis comprised of host x tumor hybrids. Cancer Res. 2000;60(9):2512-2519
11. Yilmaz Y, Lazova R, Qumsiyeh M, Cooper D, Pawelek J. Donor Y chromosome in renal carcinoma cells of a female BMT recipient: visualization of putative BMT-tumor hybrids by FISH. Bone Marrow Transpl. 2005;35(10):1021-1024
12. Mekler LB. Hybridization of transformed cells with lymphocytes as 1 of the probable causes of the progression leading to the development of metastatic malignant cells. Vestn Akad Med Nauk+. 1971;26(8):80-89
13. Goldenberg DM. On the progression of malignity: a hypothesis. Klin Wochenschr. 1968;46(16):898-899
14. Rappa G, Mercapide J, Lorico A. Spontaneous formation of tumorigenic hybrids between breast cancer and multipotent stromal cells is a source of tumor heterogeneity. Am J Pathol. 2012;180(6):2504-2515
15. Xu MH, Gao X, Luo D. et al. EMT and acquisition of stem cell-like properties are involved in spontaneous formation of tumorigenic hybrids between lung cancer and bone marrow-derived mesenchymal stem cells. PloS One. 2014;9(2):e87893
16. Chakraborty A, Lazova R, Davies S. et al. Donor DNA in a renal cell carcinoma metastasis from a bone marrow transplant recipient. Bone Marrow Transplant. 2004;34(2):183-186
17. Lazova R, Laberge GS, Duvall E. et al. A Melanoma Brain Metastasis with a Donor-Patient Hybrid Genome following Bone Marrow Transplantation: First Evidence for Fusion in Human Cancer. PloS One. 2013;8(6):e66731
18. Noubissi FK, Harkness T, Alexander CM, Ogle BM. Apoptosis-induced cancer cell fusion: a mechanism of breast cancer metastasis. Faseb J. 2015
19. Hooper S, Marshall JF, Sahai E. Tumor cell migration in three dimensions. Method Enzymol. 2006;406:625-643
20. Steingen C, Brenig F, Baumgartner L. et al. Characterization of key mechanisms in transmigration and invasion of mesenchymal stem cells. J Mol Cell Cardiol. 2008;44(6):1072-1084
21. Yang C, Tibbitt MW, Basta L, Anseth KS. Mechanical memory and dosing influence stem cell fate. Nat Mater. 2014;13(6):645-652
22. Ponte AL, Marais E, Gallay N. et al. The in vitro migration capacity of human bone marrow mesenchymal stem cells: comparison of chemokine and growth factor chemotactic activities. Stem Cells. 2007;25(7):1737-1745
23. Ries C, Egea V, Karow M. et al. MMP-2, MT1-MMP, and TIMP-2 are essential for the invasive capacity of human mesenchymal stem cells: differential regulation by inflammatory cytokines. Blood. 2007;109(9):4055-4063
24. Ozaki Y, Nishimura M, Sekiya K. et al. Comprehensive analysis of chemotactic factors for bone marrow mesenchymal stem cells. Stem Cells Dev. 2007;16(1):119-129
25. Farina AR, Tacconelli A, Cappabianca L, Gulino A, Mackay AR. Inhibition of human MDA-MB-231 breast cancer cell invasion by matrix metalloproteinase 3 involves degradation of plasminogen. Eur J Biochem. 2002;269(18):4476-4483
26. Munoz R, Man S, Shaked Y. et al. Highly efficacious nontoxic preclinical treatment for advanced metastatic breast cancer using combination oral UFT-cyclophosphamide metronomic chemotherapy. Cancer Res. 2006;66(7):3386-3391
27. Zahorec P, Koller J, Danisovic L, Bohac M. Mesenchymal stem cells for chronic wounds therapy. Cell Tissue Bank. 2015;16(1):19-26
28. Müller A, Homey B, Soto H. et al. Involvement of chemokine receptors in breast cancer metastasis. Nature. 2001;410:50-56
29. Jacob K, Webber M, Benayahu D, Kleinman HK. Osteonectin promotes prostate cancer cell migration and invasion: a possible mechanism for metastasis to bone. Cancer Res. 1999;59:4453-4457
30. Knuchel S, Anderle P, Werfelli P, Diamantis E, Rüegg C. Fibroblast surface-associated FGF-2 promotes contact-dependent colorectal cancer cell migration and invasion through FGFR-SRC signalling and integrin αVβ5-mediated adhesion. Oncotarget. 2015;6(16):14300-14317
31. Chen D, Liu S, Ma H. et al. Paracrine factors from adipose-mesenchymal stem cells enhance metastatic capactiy through Wnt signalling pathway in colon cancer cell co-culture model. Cancer Cell Int. 2015;15:42-55
32. Sakolish CM, Esch MB, Hickman JJ. et al. Modeling barrier tissues in vitro: methods, achievements, and challenges. EBioMedicine. 2016 in press
33. Li F, Tiede B, Massague J, Kang Y. Beyond tumorigenesis: cancer stem cells in metastasis. Cell Res. 2007;17(1):3-14
34. Li H, Feng Z, Tsang TC. et al. Fusion of HepG2 cells with mesenchymal stem cells increases cancerassociated and malignant properties: an in vivo metastasis model. Oncol Rep. 2014;32(2):539-547
Author contact
![]() Corresponding author: phone: 601-979-2184 fax: 601-979-5853 email: felicite.noubissi_kamdemedu
Corresponding author: phone: 601-979-2184 fax: 601-979-5853 email: felicite.noubissi_kamdemedu

 Global reach, higher impact
Global reach, higher impact