3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2016; 7(8):991-1001. doi:10.7150/jca.14663 This issue Cite
Research Paper
The Long Noncoding RNA MALAT-1 is A Novel Biomarker in Various Cancers: A Meta-analysis Based on the GEO Database and Literature
1. Key Laboratory of Carcinogenesis of Ministry of Health, Xiangya Hospital, Central South University, Changsha, Hunan 410078, China
2. Key Laboratory of Carcinogenesis and Cancer Invasion of Ministry of Education, Cancer Research Institute, Central South University, Changsha, Hunan 410078, China
3. Hunan Key Laboratory of Nonresolving Inflammation and Cancer, Disease Genome Research Center, The Third Xiangya Hospital, Central South University, Changsha, Hunan 410013, China
4. School of Public Health, Central South University, Changsha, Hunan 410078, China
Yumin Wang, Dan Xue, and Yuwei Li contributed equally to this work.
Received 2015-12-12; Accepted 2016-3-21; Published 2016-5-20
Abstract
Background: MALAT-1 is significantly overexpressed in various cancers, suggesting that it might be a potential biomarker of cancer.
Methods: A meta-analysis was performed using microarray data obtained via the Affymetrix Human Genome U133 Plus 2.0 platform found in the Gene Expression Omnibus (GEO) database and data obtained through a systematic search of PubMed and Web of Science. The pooled odds ratio (OR) and hazard ratio (HR) with 95% CI (Confidence interval) were used to judge the value of biomarkers.
Results: A total of 28 studies were included in this meta-analysis, comprising a total of 3573 patients. MALAT-1 was significantly linked with over survival (OS) (HR=1.58, 95%CI: 1.12-2.23), recurrence-free survival (RFS) (HR=2.32, 95% CI: 1.68-3.19) and death-free survival (DFS) (HR=3.28, 95% CI: 1.52-7.09). We found that MALAT-1 was a risk factor in the prognoses of lung cancer (HR=1.54, 95%CI: 1.01-2.34), digestive system cancer (HR=2.16, 95% CI: 1.34-3.48) and ovarian cancer (HR=3.98, 95% CI: 1.54-10.25). In contrast, MALAT-1 was a safe factor in the prognosis of B cell lineage cancer (HR=0.45, 95% CI: 0.33-0.61). MALAT-1 was also a risk factor of RFS in breast cancer (HR=1.97, 95% CI: 1.25-3.09) and the TNM stage in pancreatic cancer (OR=3.65, 95% CI: 1.86-7.18) and glioma (OR=4.30, 95% CI: 1.90-9.73) and was a safe factor in colorectal cancer (OR=0.17, 95% CI: 0.08-0.35). MALAT-1 was significantly associated with lymph node metastasis in clear cell carcinoma (OR=5.04, 95% CI: 2.36-10.78) and distant metastasis in pancreatic cancer (OR=11.64, 95% CI: 2.13-63.78).
Conclusions: MALAT-1 can serve as a molecular marker in different types of cancers.
Keywords: Long noncoding RNA, Cancer, MALAT-1, Meta-analysis, prognosis.
Introduction
Malignant tumors are harmful to human health and have become a leading cause of mortality worldwide. According to the "global cancer report 2012" released by the World Health Organization (WHO), the global incidence of cancer is rapidly increasing [1]. Early diagnosis and treatment are critical for the research and clinical treatment of tumors. Furthermore, tumor molecular markers are of great practical value in tumor screening, diagnosis, prognosis, and evaluation of treatment efficacy, as well as for the follow-up study of high-risk populations [2]. Thus, identifying tumor molecular markers for early diagnosis and prognosis has long been a topic of research.
Long noncoding RNA (lncRNA) is a type of noncoding RNA greater than 200 nucleotides in length [3]. In recent years, lncRNA has been shown to play an important regulatory role in chromatin modification, X chromosome inactivation and transcription, translation, genetic imprinting, dosage compensation, and the regulation of protein activity and RNA alternative splicing [4-11]. Furthermore, lncRNA has important potential applications in the diagnosis and treatment of malignant tumors [12-16]. Recent studies have shown that a variety of lncRNAs can be used as tumor markers [8-12].
MALAT-1 (metastasis associated lung adenocarcinoma transcript 1) is significantly overexpressed in many types of cancer [20-24] and has been shown to promote tumor cell proliferation, invasion and metastasis through polycomb 2 (Pc2) [25], the Wnt pathway [26], the ERK/MAPK pathway [27] and the SFPQ/PTBP2 complex [28], which is involved in tumor development. Thus, high levels of MALAT-1 expression may be related to tumor prognosis. However, most studies of MALAT-1 are limited by their small sample size. The GEO database includes a massive amount of gene chip data that profiles gene expression in many tumor types; in these datasets, MALAT-1 gene expression has been measured. To determine the value of MALAT-1 as a prognostic molecular marker, we systematically reviewed expression data and performed a quantitative meta-analysis using gene chip data from the GEO database combined with published literature.
Materials and Methods
Study strategy
To identify relevant studies for this review, we obtained microarray data based on the Affymetrix Human Genome U133 Plus 2.0 platform from the GEO database. We also retrieved all relevant published literature from Pubmed and Web of Science. The literature search was limited to the English language and ended in July, 2015. To increase the sensitivity of the search, both mesh-terms and free words were used. The search terms included "MALAT-1"; "metastasis-associated lung adenocarcinoma transcript 1"; "long intergenic noncoding RNA" or "lncRNA" or "noncoding RNA"; "cancer" or "carcinoma" or "neoplasm"; and "prognosis" or "survival".
Study selection
All five authors (Zhang, Li, Pan, Kuang and Xue) independently selected all eligible articles after reading. The eligible studies met the following criteria: any type of human cancer was involved; MALAT 1-expression in human tissue was measured and analyzed; the relationship between MALAT-1 expression and overall survival (OS), the relationship between MALAT-1 expression and recurrence free survival (RFS), or the relationship between MALAT-1 expression and disease-free survival (DFS) was identified; and the survival curve or sufficient relevant data were provided to obtain hazard ratios (HR) for survival rates and their 95% confidence intervals.
Studies were excluded if they met the following criteria: they were letters, case reports, reviews, or conference reports; the required data could not be extracted or calculated from the original article; the article was not found in full or had been published repeatedly. When the same data subsets were published in more than one article, only the latest publication was included. Controversies regarding study selection were resolved via discussion with another writer (Wang).
Data extraction
Data extraction was repeated independently by each of the six investigators, and a consensus was reached for each data set. For the 18 Affymetrix Human Genome U133 Plus 2.0 platform microarray datasets available in the GEO database that included MALAT-1 expression and related survival data, patient data such as OS, RFS, lymph node metastasis, TNM stage, survival outcome, and MALAT-1 expression value were extracted. For each article obtained from Pubmed and Web of Science, the following items were extracted: author; publication date; tumor type; country of the study population; sample size; MALAT-1 expression; cut-off value; length of follow-up; survival analysis methodology; OS, DFS, RFS, HR values and confidence intervals; and patient data such as lymphatic metastasis, TNM staging and distant metastasis. According to the requirements of our meta-analysis, a database was established after the selected data were arranged and verified, and the lnHR and SelnHR were presented as combined indexes. If the articles only provided survival curves without offering HR and Se (Standard Error) directly, appropriate data were extracted from the survival curves using Engauge Digitizer 4.1 software and the lnHR and SelnHR were calculated according to Tierney et al [29].
Characteristics of articles included in the meta-analysis
| Study | Year | Region | Tumor type | Sample size | Clinical stage of tumor | Test methods of MALAT-1 expression | Elevated MALAT-1 | Outcome measures | Survival analysis | Method* | Quality score |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Ji et al. | 2003 | Germany | Lung Cancer | 225 | Stage I, II and IIIA | qRT-PCR | Significantly higher (P=0.03) | OS | No | 1 | 25.0 |
| Schmidt et al. | 2011 | Germany | Lung Cancer | 352 | Stage I-III | qRT-PCR | Significantly higher (P<0.05) | OS | Univariate analysis and multivariate analysis | 2 | 28.5 |
| Lai et al. | 2012 | China | Hepatocellular Carcinoma | 164 | None | qRT-PCR | Up-regulation of MALAT-1 was observed in 38 (73.1%) cases (P = 0.012) | RFS | Univariate analysis and multivariate analysis | 2 | 34.5 |
| Cho et al. | 2014 | China | Multiple Myeloma | 144 | International staging system stage 1-3 | qRT-PCR | Significantly higher(p<0.001) | OS | No | 1 | 33.5 |
| Zheng et al. | 2014 | China | Colorectal Cancer | 169 | Stage II-III, N0-N2, T2-T4 | qRT-PCR | 2.26 times higher | OS DFS | Univariate analysis and multivariate analysis | 2 | 33.5 |
| Liu et al. | 2014 | China | Pancreatic Cancer | 70 | TNM: T1-T4 | qRT-PCR | Significantly higher (P=0.009) | OS | Univariate analysis and multivariate analysis | 2 | 36.0 |
| Zhang et al. | 2014 | China | Clear Cell Carcinoma | 212 | Stage I-IV, TNM: T1-T4 | qRT-PCR | Significantly higher (P < 0.05) | OS | Univariate analysis and multivariate analysis | 2 | 36.5 |
| Pang et al. | 2015 | China | Pancreatic Cancer | 141 | Early stages and advanced stages | qRT-PCR | Significantly higher (P<0.001) | OS | Univariate analysis and multivariate analysis | 2 | 32.0 |
| Ma et al. | 2015 | China | Glioma | 118 | Stage I-IV | qRT-PCR | Significantly higher (P<0.001) | OS | Univariate analysis and multivariate analysis | 2 | 32 |
*1 denoted as extracting HRs from Kaplan-Meier curves; 2 denoted as obtaining HRs directly from publications
OS: overall survival. DFS: death free survival. RFS: recurrence free survival
Quality assessment of the primary studies
A quality assessment of all included articles was performed independently by four authors (Wang, Li, Xue, Pan), according to the method of Steels et al [30,31]. The final scores are expressed as a percentage, with a higher percentage denoting a better methodological quality. The final scores are shown in Table 1.
Statistical analysis
(1) We calculated HR values and their 95% CI OS and RFS using univariate cox analysis, then drew Kaplan-Merier curves for MALAT-1 expression and the survival data using GraphPad Prism 5. The cut-off value for differentially expressed lncRNA was set to be a ≥1.5-fold difference, and the false discovery ratio (FDR) was <0.05. (2) We analyzed the obtained data for OS, DFS, RFS, clinical diagnostic criteria and other factors using Review Manager 5.3. For OS, RFS and DFS, we merged the HR values and performed heterogeneity tests. I2 > 50% was defined as significant heterogeneity [32]. If significant heterogeneity existed between studies, we used a random effects model and performed subgroup analyses, or used the fixed effects model to combined HR values and 95% confidence intervals (95% CI). All graphical representations were generated as forest plots. HR is the ratio of the prognosis in the case of high MALAT-1 expression to the prognosis in the case of low MALAT-1 expression. An HR > 1 indicates that the patients with high MALAT-1 expression have a good prognosis and the patients with low MALAT-1 expression have a poor prognosis. (3) For the studies from which we could obtain OS and DFS data, we made a funnel plot to describe publication bias using SelnHR as the abscissa and lnHR as the ordinate. We then tested the funnel plot and evaluated publication bias via a linear regression model using Stata software statistical software version 12.0 (Begg's test and Egger's test). Sensitivity analyses were conducted for the HR values and OS values of MALAT-1 extracted from the individual studies using Stata12.0. (4) For the studies from which we could obtain clinicopathological characteristics, we combined the OR values and performed heterogeneity tests to analyze the relationship between high MALAT-1 expression and lymph node metastasis, TNM stage, and distant metastasis in different types of cancers. P <0.05 was considered statistically significant.
Result
Included studies and characteristics
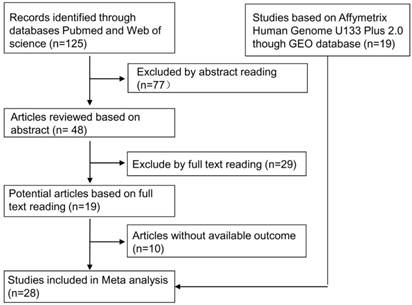
As shown in Fig. 1, we downloaded 19 studies based on Affymetrix Human Genome U133 Plus 2.0 data. We also identified 125 relevant articles, including 77 that were excluded by abstract reading. After a careful full-text reading, a total of 19 articles were reviewed in detail. Ten of these articles were excluded due to insufficient data. After this review process, 28 studies were included in the meta-analysis.
Among these 28 studies, a total of 3573 patients were represented, with a maximum sample size of 414 and a minimum sample size of 28. The mean sample size was 128. The most recent publication date was July 2015. The regions represented in the studies include America (9), China (8), Canada (6), Germany (4), Sweden (3), Japan (1), and Denmark (1). A total of 15 types of cancer were included in the meta-analysis (6 lung cancer, 4 breast cancer, 3 ovarian cancer, 3 DLBCL, 1 hepatocellular carcinoma, 1 multiple myeloma, 1 colorectal cancer, 2 pancreatic cancer, 1 clear cell carcinoma, 1 glioma, 1 meningioma, 1 melanoma, 1 lymphocytic leukemia, 1 colon cancer, and 1 CN-AML).
All studies were comprised of a high MALAT-1 expression group and a low MALAT-1 expression group. Nine studies [17, 19, 20, 23, 24, 33-36] analyzed the expression level of MALAT-1 by RT-PCR, while 19 utilized gene chips. OS, RFS and DFS were estimated as a survival outcome in 83% (24), 21% (6) and 3% (1) of the studies, respectively. 10, 7 and 3 studies estimated the association between MALAT-1 and clinicopathological characteristics of cancers (TNM stage, lymph node metastasis and distant metastasis, respectively).
GEO data analysis
We analyzed the relationship between MALAT-1 and survival (OS/RFS). The Kaplan-Meier curve (Supplemental Fig. S1), HR and 95%CI (Table 1) were derived. The cut-off value for differentially expressed lncRNA was set at ≥1.5-fold change, and the false discovery ratio (FDR) was < 0.05. We then calculated the OR between MALAT-1 and the clinicopathological characteristics of cancers (i.e., TNM stage, lymph node metastasis and distant metastasis) (Table 2).
The workflow of the meta-analysis. We identified 19 RNA microarray data sets in the GEO database. Among 125 published studies, we selected 9 by reading abstracts and full texts and by extracting data. A total of 28 studies were included in this meta-analysis.
Survival characteristics of studies based on Affymetrix Human Genome U133 Plus 2.0
| Type of cancer | GEO number | Region | No. of patients | Outcome measure | Cut off value | HR | 95%CI |
|---|---|---|---|---|---|---|---|
| Lung Cancer | GSE31210 | Japan | 226 | OS | 43503 | 3.009 | 1.165-7.767 |
| DLBCL | GSE11318 | USA | 200 | OS | 13.856 | 0.465 | 0.189-1.141 |
| Glioblastoma | GSE7696 | Canada | 80 | OS | 12.948 | 2.032 | 1.105-3.734 |
| DLBCL | GSE10846 | USA | 414 | OS | 13.986 | 0.461 | 0.333-0.637 |
| Meningioma | GSE16581 | USA | 67 | OS | 12919 | 0.274 | 0.057-1.331 |
| Ovarian Cancer | GSE18520 | USA | 53 | OS | 845.01 | 4.763 | 0.645-35.152 |
| Melanoma | GSE19234 | USA | 44 | OS | 6532.4 | 0.348 | 0.044-2.729 |
| Ovarian Cancer | GSE19829 | USA | 28 | OS | 15.339 | 3.153 | 0.876-11.348 |
| Breast Cancer | GSE20711 | Canada | 88 | OS | 12.25 | 7.208 | 0.972-53.462 |
| Lymphocytic Leukemia | GSE22762 | Germany | 107 | OS | 12.185 | 2.781 | 1.648-4.692 |
| DLBCL | GSE23501 | USA | 69 | OS | 15.193 | 0.223 | 0.049-1.009 |
| Ovarian Cancer | GSE30161 | USA | 58 | OS | 10.475 | 5.847 | 0.794-43.074 |
| CN-AML | GSE12417 | Germany | 79 | OS | 14.045 | 3.488 | 1.45-8.386 |
| Lung Cancer | GSE37745 | Sweden | 196 | OS | 11.812 | 0.9057 | 0.6503-1.261 |
| Lung Cancer | GSE29013 | USA | 55 | OS | 14.90 | 0.455 | 0.1592-1.299 |
| Lung Cancer | GSE50081 | Canada | 181 | OS | 10.39 | 1.964 | 1.084-3.560 |
| Breast Cancer | GSE6532 | Canada | 87 | RFS | 9.4118 | 1.892 | 0.896-3.997 |
| Breast Cancer | GSE9195 | Canada | 77 | RFS | -0.25977 | 2.491 | 0.8347-7.437 |
| Breast Cancer | GSE20711 | Canada | 88 | RFS | 12.564 | 1.866 | 0.9671-3.601 |
| Colon Cancer | GSE31595 | Denmark | 33 | RFS | 11.754 | 4.822 | 1.010-23.02 |
| Lung Cancer | GSE37745 | Sweden | 96 | RFS | 9.6769 | 1.947 | 0.759-4.995 |
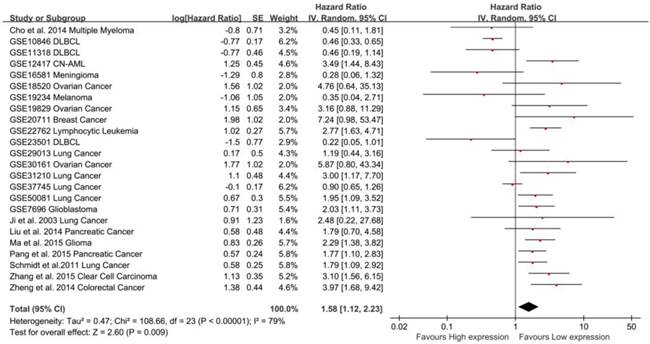
Meta-analysis of the independent role of MALAT-1 in the OS of different cancers. Twenty-two studies were used in the meta-analysis to obtain a pooled HR of OS (pooled HR: 1.57, 95%CI: 1.07-2.29).
Association between MALAT-1 and survival in 14 types of cancers
In total, 24 studies used Cox analysis to obtain the HR of OS. A significant association was founded between MALAT-1 and OS in cancer patients (pooled HR: 1.58, 95% CI: 1.12-2.23, Fig. 2). Significant heterogeneity existed between studies (Tau² = 0.47; Chi²=108.66, df=23, P<0.00001; I²=79%), and there was no significant publication bias (Egg's test: P>|t|=0.059>0.05, Begg's test: Pr>|z|=0.728) (Supplemental Fig. S2A). We used both the fixed effect model and random effect model and found that the result did not differ.
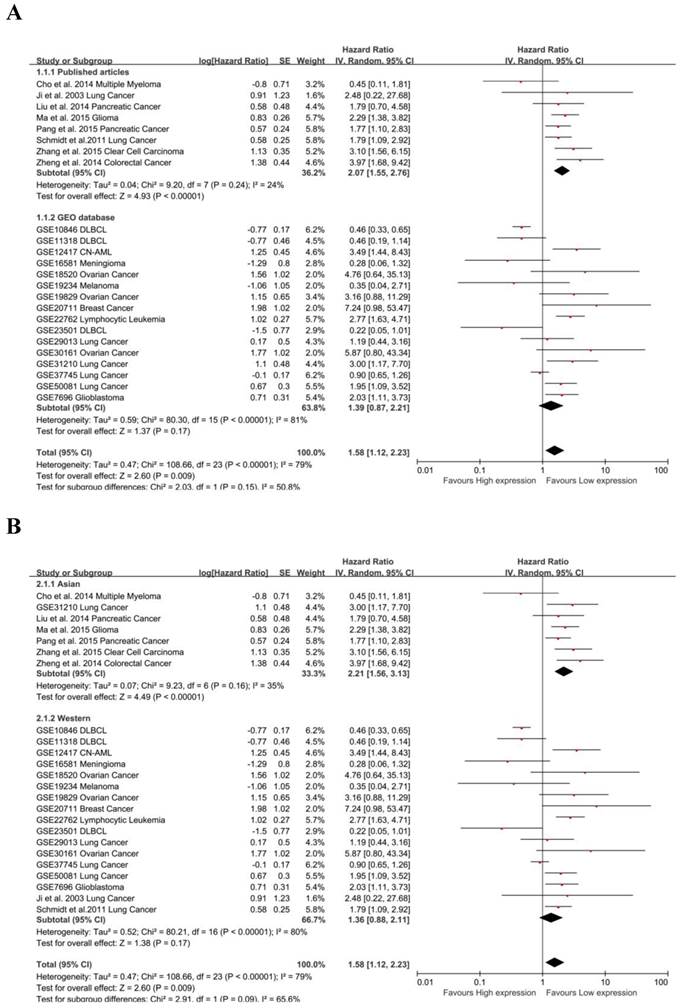
Due to the presence of heterogeneity, subgroups were analyzed for data resource and region. MALAT-1 was found to be significantly associated with the OS of cancer patients in data from published articles (pooled HR: 1.39, 95% CI: 0.87-2.21), but the association was not significant for data from the GEO database (Fig. 3A). We found a significant association between MALAT-1 and the OS of cancer patients in Asian countries (pooled HR: 2.06, 95% CI: 1.62-2.61). However, the association was not significant in western regions (pooled HR: 1.36, 95% CI: 0.88-2.11) (Fig. 3B). There was no significant heterogeneity across studies in the Asian subgroup [Chi²=9.23, df=6 (P=0.16); I²=35%] and the source of the published articles [Chi²=9.20, df=7 (P=0.24); I²=24%]. More heterogeneity existed in western subgroup [Tau²=0.52; Chi²=80.21, df=16 (P<0.00001); I²=80%] and GEO source [Tau²=0.59; Chi²=80.30, df=15 (P<0.00001); I²=81%].
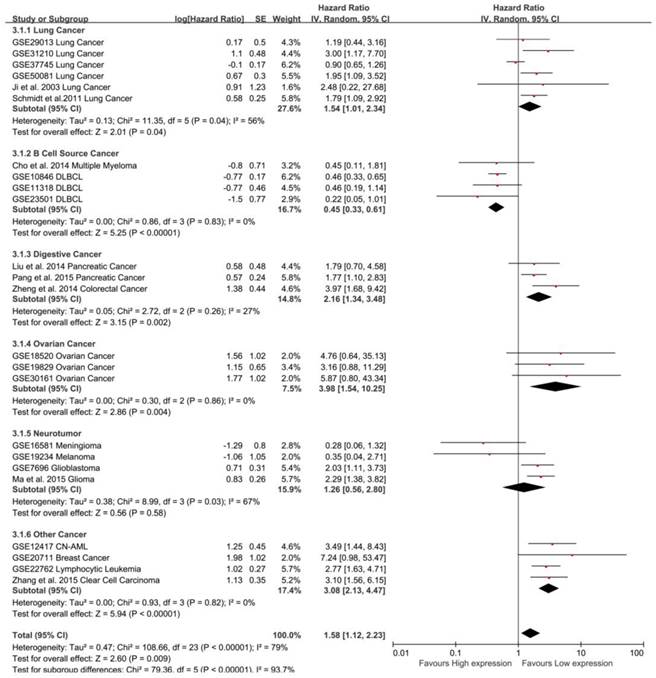
To maximize clinical relevance, subgroups were analyzed based on tumor types (Table 3). We found that MALAT-1 was a risk factor in the prognosis of lung cancer (pooled HR: 1.54, 95%CI: 1.01-2.34), digestive system cancer (pooled HR: 2.16, 95%CI: 1.34-3.48) and ovarian cancer (pooled HR: 3.98, 95% CI: 1.54-10.25). In contrast, MALAT-1 was a safe factor in the prognosis of B cell cancer (pooled HR: 0.45, 95%CI: 0.33-0.61). However, there was no significant association between MALAT-1 and OS for neurotumors. This result was strengthened by the low heterogeneity between the studies (Fig. 4).
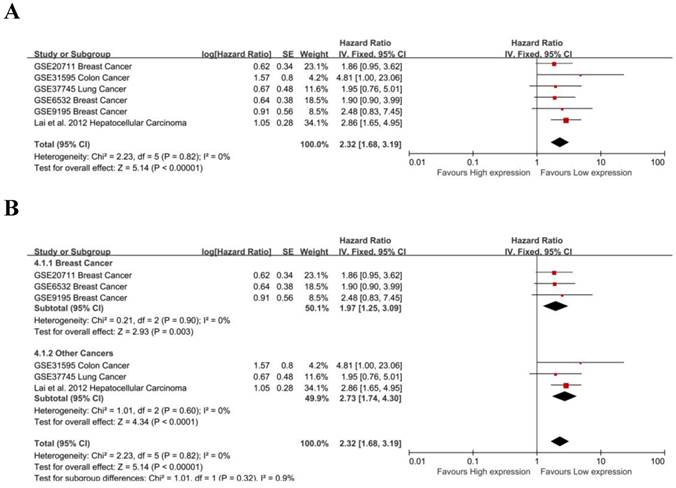
The prognostic value of MALAT-1 in RFS was evaluated for 6 studies. MALAT-1 was significantly associated with RFS (pooled HR: 2.32, 95% CI: 1.68-3.19) (Fig. 5A). There was no significant heterogeneity across the studies [Chi²=2.23, df=5(P=0.82); I²=0%], nor was there significant publication bias (Begg's test:Pr>|z|=0.260, Egg's test:P>|t|=0.656>0.05)(Supplemental Fig. S2B). Three studies focused on breast cancer, and a subgroup analysis found that MALAT-1 was significantly associated with RFS in breast cancer (pooled HR=1.97, 95%CI: 1.25-3.09; heterogeneity: Chi² = 0.21, df = 2 (P = 0.90); I² = 0%) (Fig. 5B). We used both a fixed effect model and random effect model to calculate the effect and found that the results were not markedly different.
Subgroup analysis of independent regions and sources in OS. (A) MALAT-1 was found to be significantly associated with the OS in cancer patients based on data from published articles (pooled HR: 3.97, 95%CI: 1.68-9.42), but not based on data from the GEO database; (B) Subgroups were analyzed based on the data resource and by considering the presence of heterogeneity. Founding association was significant between MALAT-1 and the OS of cancer patients in Asian (pooled HR: 2.06, 95%CI: 1.62-2.61) but not western (pooled HR: 1.02, 95%CI: 0.86-1.21) populations.
Results of subgroup analysis of pooled hazard ratios of OS and RFS of different types of cancer with increased HOTAIR expression
| Subgroup analysis | No. of studies | No. of patients | Pooled HR(95%CI) | Heterogeneity | ||
|---|---|---|---|---|---|---|
| Fixed | Random | I2 | P value | |||
| OS | 24 | 3376 | 1.34 [1.17, 1.54] | 1.58 [1.12, 2.23] | 79% | < 0.00001 |
| Data source | ||||||
| Published articles | 8 | 1431 | 2.06 [1.62, 2.61] | 2.07 [1.55, 2.76] | 24% | 0.24 |
| GEO | 16 | 1945 | 1.07 [0.90, 1.27] | 1.39 [0.87, 2.21] | 81% | < 0.00001 |
| Region | ||||||
| Asian countries | 7 | 1080 | 2.20 [1.69, 2.86] | 2.21 [1.56, 3.13] | 35% | 0.16 |
| Western countries | 17 | 2296 | 1.10 [0.94, 1.30] | 1.36 [0.88, 2.11] | 80% | < 0.00001 |
| Tumor type | ||||||
| Lung Cancer | 6 | 1235 | 1.31 [1.04, 1.66] | 1.54 [1.01, 2.34] | 56% | 0.04 |
| B Cell Source Cancer | 4 | 827 | 0.45 [0.33, 0.61] | 0.45 [0.33, 0.61] | 0% | 0.83 |
| Digestive System Cancer | 3 | 380 | 2.07 [1.42, 3.02] | 2.16 [1.34, 3.48] | 27% | 0.26 |
| Ovarian Cancer | 3 | 139 | 3.98 [1.54, 10.25] | 3.98 [1.54, 10.25] | 0% | 0.86 |
| Neurotumor | 4 | 309 | 1.83 [1.26, 2.65] | 1.26 [0.56, 2.80] | 67% | 0.03 |
| Other Cancer | 4 | 486 | 3.08 [2.13, 4.47] | 3.08 [2.13, 4.47] | 0% | 0.82 |
| RFS | 6 | 545 | 2.32 [1.68, 3.19] | 2.32 [1.68, 3.19] | 0% | 0.82 |
| Tumor type | ||||||
| Breast Cancer | 3 | 252 | 1.97 [1.25, 3.09] | 1.97 [1.25, 3.09] | 0% | 0.90 |
| Other Cancer | 3 | 293 | 2.73 [1.74, 4.30] | 2.73 [1.74, 4.30] | 0% | 0.60 |
Subgroup analysis by tumor type in OS. MALAT-1 was a risk factor for the OS of lung cancer (pooled HR: 1.54, 95%CI: 1.01-2.34), digestive system cancer (pooled HR: 2.16, 95%CI: 1.34-3.48) and ovarian cancer (pooled HR: 3.98, 95%CI: 1.54-10.25). In contrast, MALAT-1 was a safe factor for the OS of B cell cancer (pooled HR: 0.45, 95%CI: 0.33-0.61).
Meta-analysis of the independent role of MALAT-1 in the RFS of different cancers. (A) 6 studies were used in the meta-analysis to obtain a pooled HR of RFS (pooled HR: 2.32, 95%CI: 1.68-3.19); (B) MALAT-1 was a risk factor for the RFS of breast cancer (HR=1.97, 95%CI: 1.25-3.09).
Association between MALAT-1 and the clinicopathological characteristics of cancers
As shown in Table 4, 17 studies examined the association between MALAT-1 and the clinicopathological characteristics of 14 types of cancer. Nine studies examined the association between TNM stage and MALAT-1 in different cancers, including pancreatic cancer (2), glioma (1), lung cancer (2), colorectal cancer (2), clear cell carcinoma (1) and DLBCL (2). There was a significant association in pancreatic cancer (pooled OR=3.65, 95% CI: 1.86-7.18) and glioma (OR=4.30, 95% CI: 1.90-9.73), and there was no significant association in lung cancer (OR=1.01, 95% CI: 0.35-2.94), clear cell carcinoma (OR=0.61, 95% CI: 0.27-1.38) and DLBCL (pooled OR=1.04, 95% CI: 0.72-1.52). In particular, MALAT-1 is a safe factor in colorectal cancer (pooled OR=0.17, 95% CI: 0.08-0.35). There were 7 studies examining lymph node metastases from breast cancer (3), pancreatic cancer (2), clear cell carcinoma (1) and colorectal cancer (1). We observed a significant association for clear cell carcinoma (OR=5.04, 95%CI: 2.36-10.78), but no significant association for breast cancer (pooled OR=1.32, 95% CI 0.34-5.21), pancreatic cancer (pooled OR=1.95, 95% CI: 0.55-6.92) and colorectal cancer (OR=0.76, 95% CI: 0.37-1.57). There were 3 studies that examined distant metastases, including pancreatic cancer (2) and clear cell carcinoma (1). MALAT-1 was significantly associated with distant metastasis in pancreatic cancer (pooled OR=11.64, 95%CI: 2.13, 63.78) but not in clear cell carcinoma (OR=0.69, 95%CI: 0.21-2.22). Subgroup analysis, sensitivity analysis and an appraisal of publication bias were not performed due to the limited number and relative homogeneity of the studies.
Discussion
This meta-analysis used the RNA gene chip data from the GEO database to evaluate the relationship between MALAT-1 expression and survival time grouped by clinical indicators. A total of 28 studies comprising 3573 samples were included in this meta-analysis. A total of 19 studies were identified via the GEO database, representing 2142 samples. At present, very few clinical studies have focused on MALAT-1, and the sample size is therefore small. Furthermore, negative results are generally less likely to be published [37,38]. Thus, a classical meta-analysis may overestimate the effect of MALAT-1 in tumors. The inclusion of GEO data will likely result in a more extensive data source and more realistic results.
Results of meta-analysis of increased MALAT-1 expression and clinicopathological features in various cancers
| Cancer types | No. of studies | No. of patients | Pooled OR | Heterogeneity | |
|---|---|---|---|---|---|
| Fixed | Random | I2 | |||
| TNM stage | |||||
| Pancreatic Cancer | 2 | 97 | 3.65 [1.86, 7.18] | 3.78 [1.70, 8.39] | 19% |
| Glioma | 1 | 126 | 4.30 [1.90, 9.73] | 4.30 [1.90, 9.73] | Not applicable |
| Lung Cancer | 2 | 422 | 0.89 [0.45, 1.74] | 1.01 [0.35, 2.94] | 56% |
| Clear Cell Carcinoma | 1 | 106 | 0.61 [0.27, 1.38] | 0.61 [0.27, 1.38] | Not applicable |
| Colorectal Cancer | 2 | 173 | 0.29 [0.16, 0.53] | 0.50 [0.05, 4.86] | 87% |
| Lymph node metastasis | |||||
| Breast Cancer | 3 | 252 | 1.28 [0.77, 2.12] | 1.32 [0.34, 5.21] | 85% |
| Pancreatic Cancer | 2 | 152 | 2.06 [0.95, 4.45] | 1.95 [0.55, 6.92] | 60% |
| Clear Cell Carcinoma | 1 | 126 | 5.04 [2.36, 10.78] | 5.04 [2.36, 10.78] | Not applicable |
| Colorectal Cancer | 1 | 146 | 0.76 [0.37, 1.57] | 0.76 [0.37, 1.57] | Not applicable |
| Distant metastasis | |||||
| Pancreatic Cancer | 2 | 171 | 11.64 [2.13, 63.78] | 9.57 [1.66, 55.01] | 0% |
| Clear Cell Carcinoma | 1 | 106 | 0.69 [0.21, 2.22] | 0.69 [0.21, 2.22] | Not applicable |
In recent years, numerous studies have demonstrated that MALAT-1 is an independent prognostic marker for various tumors, including glioma, pancreatic cancer and colon cancer. However, the sample size of these studies is small. This meta-analysis assesses the value of MALAT-1 as a marker of tumor prognosis. Through systematic analysis, we found that MALAT-1 was highly expressed in many types of tumors. By combining the HRs from Cox analysis, we found that MALAT-1 was an independent risk factor for OS in tumor patients (HR=1.58, 95%CI: 1.12-2.23). In addition, MALAT-1 can be regarded as an independent prognostic risk factor (HR=2.32, 95%CI: 1.68-3.19) for RFS in patients. We also performed an analysis of DFS, which indicated that MALAT-1 is an independent prognostic marker for DFS (HR=3.28, 95%CI: 1.52-7.09).
Due to the large difference among various types of tumors, the clinical relevance of our findings for all tumor types is limited. To explore this point in greater detail, we analyzed different tumor subgroups. We found that there was a significant association between high MALAT-1 expression and poor prognosis in lung cancer (HR=1.54, 95%CI: 1.01-2.34), digestive system cancer (HR=2.16, 95%CI: 1.34-3.48) and ovarian cancer (HR=3.98, 95%CI: 1.54-10.25). However, for B cell source cancer (HR=0.45, 95%CI: 0.33-0.61), high expression of MALAT-1 is related to a good prognosis. The heterogeneity of each subgroup was significantly decreased in our subgroup analysis of different tumor types, thereby strengthening our results. Subgroup analysis revealed that the expression of MALAT1 can be used as a prognostic factor for OS and RFS, but for individual tumor types such as DLBCL, the high expression of MALAT-1 is a safety factor. This suggested that the effect of MALAT-1 in different types of tumors varies. In the prognosis of RFS, we found that the effect of MALAT-1 on the prognosis of breast cancer was very significant (HR=1.97, 95%CI: 1.25-3.09) and that heterogeneity [Chi2=0.21, df=2(P=0.90); I2=0%] was small, making our conclusion more reliable. Both Begg's test and Egg's test failed to identify a significant publication bias regarding the independent prognostic role of MALAT-1 in OS and RFS. Through a sensitivity analysis, the results of both the fixed effect model and random effect model were found to exhibit no obvious differences, indicating that our analysis is stable and that the overexpression of MALAT-1 is associated with poor prognosis.
TNM stage is associated with cancer prognosis. We found a significant association in pancreatic cancer (pooled OR=3.65, 95% CI: 1.86-7.18) and glioma (OR=4.30, 95% CI: 1.90-9.73), but there was no significant difference in lung cancer (OR=1.01, 95% CI: 0.35-2.94), clear cell carcinoma (OR=0.61, 95%CI: 0.27-1.38) or DLBCL (OR=1.04, 95%CI: 0.72-1.52). Interestingly, MALAT-1 was a safe factor in colorectal cancer (OR=0.17, 95% CI: 0.08-0.35). In the case of lymph node metastasis, we observed a significant association in clear cell carcinoma (OR=5.04, 95%CI: 2.36-10.78) but not breast cancer (OR=1.32, 95%CI: 0.34-5.21), pancreatic cancer (OR=1.95, 95%CI: 0.55-6.92) or colorectal cancer (OR=0.76, 95%CI: 0.37-1.57). The overexpression of MALAT-1 was significantly associated with distant metastasis in pancreatic cancer (OR=11.64, 95%CI: 2.13-63.78), but held no significance for clear cell carcinoma (OR=0.69, 95%CI: 0.21-2.22). The value of MALAT-1 as a biomarker is therefore different for different types of cancers.
In recent years, many studies have demonstrated that lncRNAs can regulate the expression of genes in and are widely involved in tumor progression and metastasis. MALAT-1 is highly expressed in different types of tumors and can influence the prognosis of tumor patients by promoting tumor growth, invasion and metastasis. In addition, MALAT-1 can promote the proliferation of tumors. MALAT-1 combines with SFPQ to break the SFPQ/PTBP2 complex [28], thereby releasing PTBP2. It can also act as a molecular scaffold in combination with the demethylated polycomb protein 2 (PC2) to activate growth control genes regulated by E2F at the transcriptional level [18]. In parallel, MALAT-1 regulates the expression of Bcl-2 to inhibit apoptosis [36].
MALAT-1 can also promote the invasion and metastasis of tumors. Down-regulation of MALAT-1 can arrest the cell cycle in the G2/M phase and G0/G1 phase, thereby promoting cell apoptosis and inhibiting tumor invasion and metastasis [39,40]. The combination of MALAT-1 and Ezh2 up-regulates β-catenin, activating the Wnt pathway to promote tumor invasion and metastasis [26]. It has also been reported that MALAT-1 can activate the ERK/MAPK pathway [27].
In addition, MALAT-1 can facilitate epithelial mesenchymal transition (EMT). MALAT-1 interacts with suppressor of zeste 12 (Suz12), thereby down-regulating the expression of E-cadherin and up-regulating the expression of N-cadherin and fibronectin to enhance TGF-β-induced EMT [41]. High expression MALAT-1 can also up-regulate CCL5 through the MALAT-1/Snail pathway to promote the EMT, tumor invasion and metastasis [42]. At the same time, MALAT-1 enhances the stem cell-like biological activity of tumor cells by up-regulating the expression of self-renewal related factors through Sox2 [43]
Our study has several limitations: (1) There is a significant correlation between the length of survival and the treatment of patients, and these differences might have a great impact on HR and thus result in heterogeneity. (2) There are 2 articles whose HRs and 95%CI could not be directly obtained from the primary studies and which were estimated by software; this could lead to error in the HR value. (3) Differences in paper quality across studies could lead to bias in the meta-analysis. (4) Heterogeneity in the relationship between MALAT-1 expression and clinical diagnosis was large. Therefore, the study conclusions must be confirmed for a larger sample in a multicenter and randomized controlled prospective study.
In conclusion, MALAT-1 overexpression was significantly associated with survival (OS, RFS and DFS) in cancer patients. The association between MALAT-1 and clinical indicators such as TNM stage, lymph node metastasis, and distant metastasis was different in different types of cancers. Thus, MALAT-1 is a novel biomarker in various cancers. In the future, additional studies will be necessary to explore the role of MALAT-1 in human cancer.
Supplementary Material
Supplementary figures.
Acknowledgements
This work was supported by grants from The Students' Innovation and Entrepreneurship Training Program of Central South University (CY14283, CY14284, CX2015466, CX2015473), the Free Exploration Program of Central South University (ZY2015772), and the Natural Science Foundation of Hunan Province (14JJ1010, 2015JJ1022).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87-108
2. DeSantis CE, Lin CC, Mariotto AB, Siegel RL, Stein KD, Kramer JL. et al. Cancer treatment and survivorship statistics, 2014. CA Cancer J Clin. 2014;64:252-71
3. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007;316:1484-8
4. Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10:155-9
5. Zhou Y, Liao Q, Li X, Wang H, Wei F, Chen J. et al. HYOU1, Regulated by LPLUNC1, Is Up-Regulated in Nasopharyngeal Carcinoma and Associated with Poor Prognosis. J Cancer. 2016;7:367-76
6. Xiao K, Yu Z, Li X, Li X, Tang K, Tu C. et al. Genome-wide Analysis of Epstein-Barr Virus (EBV) Integration and Strain in C666-1 and Raji Cells. J Cancer. 2016;7:214-24
7. Gong Z, Yang Q, Zeng Z, Zhang W, Li X, Zu X. et al. An integrative transcriptomic analysis reveals p53 regulated miRNA, mRNA, and lncRNA networks in nasopharyngeal carcinoma. Tumour Biol. 2015 [Epub ahead of print]
8. Yan Q, Zeng Z, Gong Z, Zhang W, Li X, He B. et al. EBV-miR-BART10-3p facilitates epithelial-mesenchymal transition and promotes metastasis of nasopharyngeal carcinoma by targeting BTRC. Oncotarget. 2015;6:41766-82
9. Xu K, Xiong W, Zhou M, Wang H, Yang J, Li X. et al. Integrating ChIP-sequencing and digital gene expression profiling to identify BRD7 downstream genes and construct their regulating network. Mol Cell Biochem. 2016;411:57-71
10. Bo H, Gong Z, Zhang W, Li X, Zeng Y, Liao Q. et al. Upregulated long non-coding RNA AFAP1-AS1 expression is associated with progression and poor prognosis of nasopharyngeal carcinoma. Oncotarget. 2015;6:20404-18
11. Zeng Z, Bo H, Gong Z, Lian Y, Li X, Li X. et al. AFAP1-AS1, a long noncoding RNA upregulated in lung cancer and promotes invasion and metastasis. Tumour Biol. 2015 [Epub ahead of print]
12. Zhang W, Huang C, Gong Z, Zhao Y, Tang K, Li X. et al. Expression of LINC00312, a long intergenic non-coding RNA, is negatively correlated with tumor size but positively correlated with lymph node metastasis in nasopharyngeal carcinoma. J Mol Histol. 2013;44:545-54
13. Gong Z, Zhang S, Zeng Z, Wu H, Yang Q, Xiong F. et al. LOC401317, a p53-regulated long non-coding RNA, inhibits cell proliferation and induces apoptosis in the nasopharyngeal carcinoma cell line HNE2. Plos One. 2014;9:e110674
14. Gong Z, Zhang S, Zhang W, Huang H, Li Q, Deng H. et al. Long non-coding RNAs in cancer. Sci China Life Sci. 2012;55:1120-4
15. Panzitt K, Tschernatsch MM, Guelly C, Moustafa T, Stradner M, Strohmaier HM. et al. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology. 2007;132:330-42
16. Wang J, Liu X, Wu H, Ni P, Gu Z, Qiao Y. et al. CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res. 2010;38:5366-83
17. Zhang S, Chen S, Yang G, Gu F, Li M, Zhong B. et al. Long noncoding RNA HOTAIR as an independent prognostic marker in cancer: a meta-analysis. Plos One. 2014;9:e105538
18. Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX. et al. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology. 2011;54:1679-89
19. Lai MC, Yang Z, Zhou L, Zhu QQ, Xie HY, Zhang F. et al. Long non-coding RNA MALAT-1 overexpression predicts tumor recurrence of hepatocellular carcinoma after liver transplantation. Med Oncol. 2012;29:1810-6
20. Zhang HM, Yang FQ, Chen SJ, Che J, Zheng JH. Upregulation of long non-coding RNA MALAT1 correlates with tumor progression and poor prognosis in clear cell renal cell carcinoma. Tumour Biol. 2015;36:2947-55
21. Ma KX, Wang HJ, Li XR, Li T, Su G, Yang P. et al. Long noncoding RNA MALAT1 associates with the malignant status and poor prognosis in glioma. Tumour Biol. 2015;36:3355-9
22. Dong Y, Liang G, Yuan B, Yang C, Gao R, Zhou X. MALAT1 promotes the proliferation and metastasis of osteosarcoma cells by activating the PI3K/Akt pathway. Tumour Biol. 2015;36:1477-86
23. Cho SF, Chang YC, Chang CS, Lin SF, Liu YC, Hsiao HH. et al. MALAT1 long non-coding RNA is overexpressed in multiple myeloma and may serve as a marker to predict disease progression. BMC Cancer. 2014;14:809
24. Zheng HT, Shi DB, Wang YW, Li XX, Xu Y, Tripathi P. et al. High expression of lncRNA MALAT1 suggests a biomarker of poor prognosis in colorectal cancer. Int J Clin Exp Pathol. 2014;7:3174-81
25. Yang L, Lin C, Liu W, Zhang J, Ohgi KA, Grinstein JD. et al. ncRNA- and Pc2 methylation-dependent gene relocation between nuclear structures mediates gene activation programs. Cell. 2011;147:773-88
26. Hirata H, Hinoda Y, Shahryari V, Deng G, Nakajima K, Tabatabai ZL. et al. Long Noncoding RNA MALAT1 Promotes Aggressive Renal Cell Carcinoma through Ezh2 and Interacts with miR-205. Cancer Res. 2015;75:1322-31
27. Wu XS, Wang XA, Wu WG, Hu YP, Li ML, Ding Q. et al. MALAT1 promotes the proliferation and metastasis of gallbladder cancer cells by activating the ERK/MAPK pathway. Cancer Biol Ther. 2014;15:806-14
28. Ji Q, Zhang L, Liu X, Zhou L, Wang W, Han Z. et al. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br J Cancer. 2014;111:736-48
29. Tierney JF, Stewart LA, Ghersi D, Burdett S, Sydes MR. Practical methods for incorporating summary time-to-event data into meta-analysis. Trials. 2007;8:16
30. Xing X, Tang YB, Yuan G, Wang Y, Wang J, Yang Y. et al. The prognostic value of E-cadherin in gastric cancer: a meta-analysis. Int J Cancer. 2013;132:2589-96
31. Steels E, Paesmans M, Berghmans T, Branle F, Lemaitre F, Mascaux C. et al. Role of p53 as a prognostic factor for survival in lung cancer: a systematic review of the literature with a meta-analysis. Eur Respir J. 2001;18:705-19
32. Egger M, Davey SG, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997;315:629-34
33. Ji P, Diederichs S, Wang W, Boing S, Metzger R, Schneider PM. et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003;22:8031-41
34. Liu JH, Chen G, Dang YW, Li CJ, Luo DZ. Expression and prognostic significance of lncRNA MALAT1 in pancreatic cancer tissues. Asian Pac J Cancer Prev. 2014;15:2971-7
35. Pang EJ, Yang R, Fu XB, Liu YF. Overexpression of long non-coding RNA MALAT1 is correlated with clinical progression and unfavorable prognosis in pancreatic cancer. Tumour Biol. 2015;36:2403-7
36. Schmidt LH, Gorlich D, Spieker T, Rohde C, Schuler M, Mohr M. et al. Prognostic impact of Bcl-2 depends on tumor histology and expression of MALAT-1 lncRNA in non-small-cell lung cancer. J Thorac Oncol. 2014;9:1294-304
37. Xue Y, Teng YQ, Zhou JD, Rui YJ. Prognostic value of long noncoding RNA MALAT1 in various carcinomas: evidence from nine studies. Tumour Biol. 2015 [Epub ahead of print]
38. Wu Y, Lu W, Xu J, Shi Y, Zhang H, Xia D. Prognostic value of long non-coding RNA MALAT1 in cancer patients. Tumour Biol. 2015 [Epub ahead of print]
39. Jiao F, Hu H, Yuan C, Wang L, Jiang W, Jin Z. et al. Elevated expression level of long noncoding RNA MALAT-1 facilitates cell growth, migration and invasion in pancreatic cancer. Oncol Rep. 2014;32:2485-92
40. Ren S, Liu Y, Xu W, Sun Y, Lu J, Wang F. et al. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J Urol. 2013;190:2278-87
41. Fan Y, Shen B, Tan M, Mu X, Qin Y, Zhang F. et al. TGF-beta-induced upregulation of malat1 promotes bladder cancer metastasis by associating with suz12. Clin Cancer Res. 2014;20:1531-41
42. Kan JY, Wu DC, Yu FJ, Wu CY, Ho YW, Chiu YJ. et al. Chemokine (C-C Motif) Ligand 5 is Involved in Tumor-Associated Dendritic Cell-Mediated Colon Cancer Progression Through Non-Coding RNA MALAT-1. J Cell Physiol. 2015;230:1883-94
43. Jiao F, Hu H, Han T, Yuan C, Wang L, Jin Z. et al. Long noncoding RNA MALAT-1 enhances stem cell-like phenotypes in pancreatic cancer cells. Int J Mol Sci. 2015;16:6677-93
Author contact
![]() Corresponding author: Zhaoyang Zeng, E-mail: zengzhaoyangedu.cn; Tubao Yang, Email: yangtbcsucom
Corresponding author: Zhaoyang Zeng, E-mail: zengzhaoyangedu.cn; Tubao Yang, Email: yangtbcsucom

 Global reach, higher impact
Global reach, higher impact