3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2016; 7(4):427-435. doi:10.7150/jca.13749 This issue Cite
Research Paper
Chromobox Homolog 4 is Positively Correlated to Tumor Growth, Survival and Activation of HIF-1α Signaling in Human Osteosarcoma under Normoxic Condition
1. Department of Orthopedics, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, No.600, Yishan Road, Shanghai, 200233, China;
2. Institute of Orthopaedics, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, No.600, Yishan Road, Shanghai, 200233, China;
3. Department of Ultrasound, Shanghai Jiao Tong University Affiliated Sixth People's Hospital, No.600, Yishan Road, Shanghai, 200233, China.
* These authors contributed equally to this work.
Received 2015-9-4; Accepted 2015-12-8; Published 2016-1-28
Abstract
Objectives: The clinical significance and tumorigenesis of Chromobox homolog 4 (CBX4) have been reported in hepatocellular carcinoma. The purpose of this study is to confirm the expression, elucidate the biological function and investigate the potential mechanism of CBX4 in osteosarcoma (OS). Methods: The expression of CBX4 in OS samples and cell lines was measured by RT-PCR and western blot test. Cell cycle, CCK8 and colony-forming assays were used to detect changes of cells growth. Cell apoptosis assay was used to measure cell survival capacity. Trans-well assay was used to test the activities of migration and invasion. The expression of genes regulated by CBX4 was detected by qRT-PCT test. Results: The expression of CBX4 was up-regulated in multiple OS cell lines and clinical samples. Overexpression of CBX4 was correlated with advanced clinical stage, high degree of malignancy and low tumor necrosis rate. Moreover, knockdown of CBX4 resulted in significant inhibition of cell growth and cell survival in OS cells under normoxic condition. In addition, we found that knockdown of CBX4 lead to down-regulating of HIF-1α-targeted genes without changing HIF-1α expression itself. Conclusion: Taken together, CBX4 is up-regulated and has a pro-tumor effect in OS with an activation of HIF-1α signaling pathway under normoxic condition. Therefore, targeting CBX4 may provide a new therapeutic method for OS.
Keywords: Chromobox homolog 4, Growth, Survival, HIF-1α pathway, Osteosarcoma.
Introduction
Osteosarcoma (OS), the most frequent primary malignant bone tumor, has a peak incidence in the second decade of life (1). Though advance in the treatment, the efficacy of current strategies for the metastatic and recurrent OS is limited (2, 3). In fact, the etiology and pathogenesis intrinsic to the malignant properties of OS remain blurred.
Polycomb group (PcG) proteins, a member of Polycomb Repressive Complexes (PRCs), are important transcriptional repressors that exert pivotal role in regulating early embryogenesis and tumorigenesis in mammals (4, 5). The large variety of PcG proteins forms at least two PRCs, PRC1 and PRC2, which are of different properties (6). Some of the PRCs subunits, like BMI1 and EZH2, have been found overexpressed in a variety of tumors (7, 8). However, for the complex functions of PcG proteins, the exact regulatory events of PRCs are almost unclear.
The chromobox family (CBXs), including five members (CBX2, CBX4, CBX6, CBX7 and CBX8), interact with the core PRC1 complex to perform distinct functions in different tissues (9, 10). The most studied CBX in cancers was CBX7. On one hand, CBX7 was an oncogene that was activated in prostate cancer (11) and ovarian cancer (12). On the other, CBX7 had tumor-suppressing capabilities in certain contexts (13). However, little was known about tumor associated properties in other CBXs. The expression of CBX2 was reported to be associated with tumor stage and overall survival in some cancers (14, 15), yet no published functional tests provided evidence that CBX2 was an oncogene. Lee et al reported CBX8 was an oncogene via cooperating with SIRT1 and inhibiting Tp53 activity in breast cancer cells (16). CBX4 (also known as hPC2), besides being part of PRC1 complex, is also a small ubiquitin-related modifier (SUMO) E3 ligase, which is unique among CBXs protein (17). Recent research showed that CBX4 was correlated to prognosis and tumor growth in hepatocellular carcinoma, which indicated that CBX4 was an oncogene (18). Furthermore, Li et al observed that CBX4 promoted angiogenesis of hepatocellular carcinoma via sumoylating of HIF-1α protein (19), which confirmed HIF-1α to be a new substrate for the SUMO E3 ligase activity of CBX4.
To date, HIF-1α is overexpressed and associated to a poor prognosis in various human cancers (20, 21). It performs function by activating the downstream genes, which regulate the proliferation, apoptosis, and metastasis of tumor cells. Generally, HIF-1α pathway was activated by O2-dependent post-translational modification of prolyl hydroxylation (22). However, the HIF-1α pathway is also regulated by other signals under normoxic conditions, such as the loss of tumor-suppressor function and the gain of oncogene function (23, 24).
In this study, we showed for the first time that CBX4 was up-regulated and correlated to in vitro tumor growth and survival in OS. Moreover, knockdown of CBX4 inhibited the expression of HIF-1α targeted genes under normoxic condition. Therefore, CBX4 was an oncogene in OS, suggesting a potential target for OS treatment.
Materials and methods
Cell lines and cell culture
Human fetal osteoblastic cell line (hFOB) and human OS cell lines (MNNG/HOS, U2OS and MG63) were purchased from the institute of Cell Bank for Biological sciences (Shanghai, China). Cells were maintained at 37℃in a humidified air atmosphere containing 5% CO2 in DMEM (MNNG/HOS, MG63) and RPIM-1640 (U2OS) supplemented with 10% fetal bovine serum (Biowest, South America Origin), 100U/ml penicillin (Sigma-Aldrich, St Louis, MO, USA) and 100mg/ml streptomycin(Sigma-Aldrich). HFOB was cultured according to ATCC protocols and references described as before (25, 26).
Patients and OS samples
The 20 paired samples of human OS and their matched adjacent non-cancerous tissues were collected at the time of surgery between 2014 and 2015 at the Department of bone and soft oncology, Shanghai Jiao Tong University affiliated Sixth People's hospital (Shanghai, China). Upon resection, human surgical specimens were immediately frozen in liquid nitrogen and stored at -80℃ in the refrigerator. All the patients agreed to participate in the study and gave informed consent. The study was approved by the Ethics committee of Shanghai Jiao Tong University.
RNA isolation and qRT-PCR assays
Total RNA of human tissue samples and cultured cells was extracted and quantified with Trizol Kit (Invitrogen, Carlsbad, CA, USA) and Nanodrop 2000 (Thermo Fisher Scientific, Waltham, MA, USA) respectively according to the manufacturer's protocol. The PrimeScript RT Reagent kit (TaKaRa, Shiga, Japan) was used to synthesize the first-strand cDNA. RT-PCR was performed with SYBR Green premix Ex Taq (TaKaRa) as follow: an initial pre-denaturation step for 30 seconds at 95℃, followed by amplification of 40 cycles at 95℃ for 5 seconds and at 60 ℃ for 20 seconds, melting curve analysis was performed at the end. All reactions were done in a 10 ul reaction volume in triplicate. The expression level of genes was measured using the comparative Ct method. The information of the primer sequences used was described in Table S1.
Cell transfection
Transfection of siRNA was performed by RNAi-max (Invitrogen) according to the manufacturer's protocol. The siRNA sequences targeting CBX4 were: forward primer: 5'-AGAUGAAGAUAGUCAAGAA-3' and reverse primer: 5'-UUCUUGACUAUCUUCAUCU-3'. In brief, transfection with the final concentration of 50 nM siRNA was conducted when the cell density was 30-50% in six-well plates. For CCK8, cell cycle, cell apoptosis, colony formation, trans-well, RNA extraction and western blot assays, cells were used 48 h after transfection.
Cell proliferation and cell cycle assay
Cell counting kit-8 (Dojindo Molecular Technologies, Inc, Kumamoto, Japan) assay was used to measure the cell viability. In short, the CBX4 siRNA-transfected MNNG/HOS or MG63 cells were seeded at the density of 3000 and 2500 per well in 96-well microplate, respectively. 10 μl CCK8 solution and 100 μl DMEM were added to each well and incubated for 2 h. The optical density was detected at 450 nm wavelength by microplate reader (Model 680, Bio-Rad Laboratories, Hercules, CA, USA). This procedure was repeated once a day for 5 days. For cell cycle assay, 48 hours after transfection, the cells were fixed in 70% ethanol at -20℃ in the refrigerator for 12-24 hours. After that, cells were treated with staining solution, which containing 50ug/ml propidium iodide (PI) (Biolegend, California, USA) and 50ug/ml RNase A (BD LSRⅡ, San Jose, CA, USA). Each experiment was repeated three times.
Colony formation assay
Transfected OS cells were collected and placed in six plates (1000 cells/well). The cells were cultured for 2 weeks to form colonies. Colonies were fixed with 95% methanol for 10 minutes and stained with 0.1% crystal violet for 10 minutes, washed, and air-dried. Colony-forming efficiency was calculated as colonies/plated cells × 100%.
Cell apoptosis analysis
For the apoptosis experiment, following 48 hours of transfection, cells were cultured in 37℃ incubator for 24 hours. The fluorescein isothiocyanate Annexin V Apoptosis Detection kit I(BD Pharmingen, San Diego, CA, USA) was used. Briefly, the cells were collected and centrifuged at 2000 ×g for 5 min. Then the cells resuspended in 500 μl binding buffer, supplemented with 5 μl Annexin V and 5 μl propidium iodide (PI), for 15 min of dark treatment at the room temperature. The flow cytometry (FC500 MPL, Beckman Coulter, Brea, CA, USA) was used to analyze the samples.
Migration and invasion assays
Cell migration and invasion assays were performed in 24-well plates with 8 cm pore size chamber inserts (Corning, NY, USA). For migration assays, 5 × 104 cells were placed into each well of the upper chamber with the non-coated membrane. For invasion assays, 1× 105 cells were placed into the upper chamber with the Matrigel-coated medium. In both assays, cells were suspended in 200 ul of DMEM without fetal bovine serum when they were seeded into the upper chamber. In the lower chamber, 800 ul of DMEM supplemented with 10% fetal bovine serum was added. After incubation for 16 h (migration assay) or 24 h (invasion assay), the chamber inserts were removed from the plate, and non-invading cells were removed from the upper surface of the membrane. Cells that moved to the bottom surface of the membrane were fixed with 100% methanol for 20 min and stained with 0.1% crystal violet for 30 min. Finally, at least 10 randomly selected fields were imaged and the cell numbers were counted under a CKX41 inverted microscope (Olympus, Tokyo, Japan). The assays were conducted with at least three independent repeats.
Protein extraction and western blot analysis
These procedures were performed as described previously (27). In short, total protein was isolated from cell lines or fresh tissues and was separated by 6 and 8% SDS-PAGE gels. Then we followed standard procedures using primary antibodies: CBX4 (Bioworld Technology, Inc.), HIF-1α (Santa Cruz Biotechnology), β-Actin (Sigma-Aldrich). The secondary antibody was horseradish peroxidase-conjugated anti-rabbit IgG (Sigma-Aldrich). SuperSignal West Femto Maximun Sensitivity Substrate (Thermo Fisher Scientific) was used in the subsequent visualization.
Statistical analysis
Data were imaged with GraphPad Prism 5 software (Graphpad Software, Inc, La Jolla, CA, USA). Quantitative variables were presented as means ± standard deviation. A two-tailed student's t-test was used to compare the differences between two groups. The SPSS version 16.0 (SPSS, Inc., Chicago, IL, USA) software was used to analyze the data. P<0.05 was considered statistically significant different.
Results
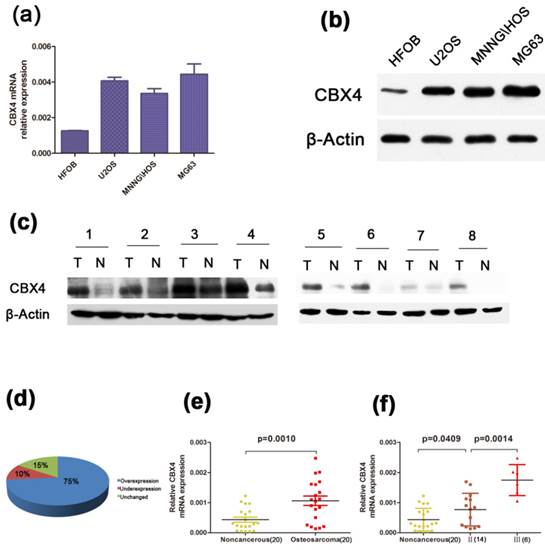
CBX4 expression is up-regulated in OS clinical samples and associated with the progress of OS
To identify the expression level of CBX4 in OS, we examined the CBX4 expression at its mRNA and Protein level, respectively. A higher expression of CBX4 was observed in osteosarcoma cell lines (U2OS, MNNG/HOS, MG63) compared with human osteoblast cell lines (HFOB) (Figure 1 a, b). Moreover, by using fresh OS specimens, we also demonstrated that CBX4 protein was up-regulated in all eight OS compared with corresponding non-cancerous tissues (Figure 1 c). Meanwhile, we detected the mRNA expression of CBX4 in 20 pairs of human OS and their corresponding non-cancerous tissues. The correlation between XBP1 expression and the data of OS patients was shown in Table 1. The XBP1 mRNA expression was not correlated to age, gender, anatomic location, or tumor size statistically significant. However, the correlation with clinical stage (p<0.01), degree of malignancy (p<0.05), tumor necrosis rate (p<0.05) was significant. In addition, CBX4 was overexpressed in 75% of OS tissues compared with non-cancerous tissues (Figure 1 d), and the expression was more than twofold (Figure 1 e) in OS compared with non-cancerous tissues. We also observed a significantly increase in the expression of CBX4 mRNAs in advanced stage (Ⅲ) compared with early stage (Ⅱ) (Figure 1 f).Taken together, these results indicated that CBX4 was up-regulated and potentially had a pivotal role in the tumorigenesis of OS.
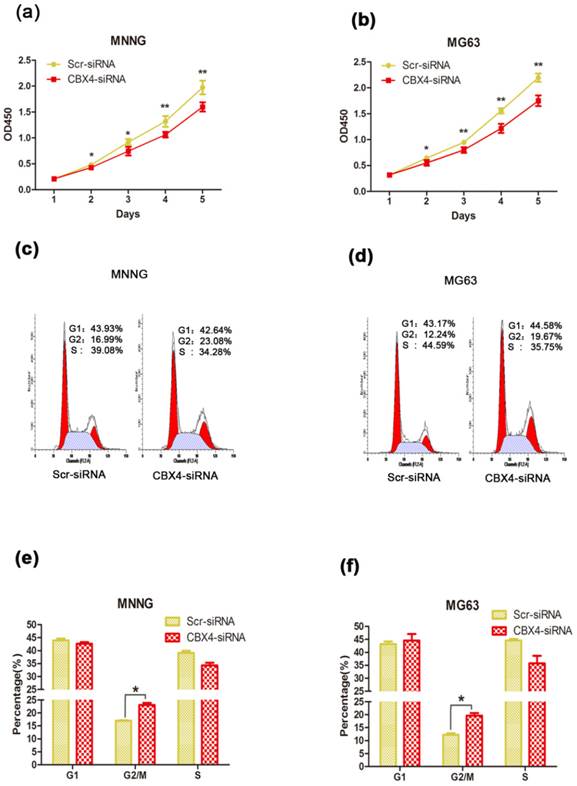
Knockdown of CBX4 inhibited cell proliferation through G2/M phase arrest
To explore the potential role of CBX4 in tumorigenesis, we first evaluated the effect of CBX4 on the growth of OS cells in vitro. After knockdown of CBX4 by specifically targeted siRNA, the viabilities of MNNG/HOS and MG63 cells were obviously inhibited by CCK8 assays (Figure 2 a, b). Furthermore, knockdown of CBX4 delayed the cell cycle progression by arresting G2/M phase in both MNNG/HOS (Figure 2 c, e) and MG63 cell (Figure 2 d, f). We then detected the effect of CBX4 on OS cells migration and invasion, however, the effect was weak with no statistical significance (Figure S1). To summarize, these data showed that CBX4 was essential for the proliferation of OS cells.
The relationship between CBX4 mRNA expression and their clinicopathologic parameters in 20 of osteosarcoma patients
| Parameters | N (%) | Mean ± s. d. | p-value |
|---|---|---|---|
| Total | 20(100%) | ||
| Age (years) | |||
| <20 | 12(60%) | 0.0009±0.0006 | 0.2418 |
| ≥20 | 8(40%) | 0.0013±0.0008 | |
| Gender | |||
| Male | 14(70%) | 0.0011±0.0007 | 0.5306 |
| Female | 6(30%) | 0.0009±0.0007 | |
| Anatomic location | |||
| Tibia/ Femur | 14(70%) | 0.0027±0.0021 | 0.5421 |
| Elsewhere | 6(30%) | 0.0034±0.0025 | |
| Ennecking stage | |||
| Ⅱ | 14(70%) | 0.0008±0.0005 | 0.0014** |
| Ⅲ | 6(30%) | 0.0017±0.0005 | |
| Tumor size (cm3) | |||
| <50 | 10(50%) | 0.0009±0.0008 | 0.4051 |
| ≥50 | 10(50%) | 0.0012±0.0006 | |
| Degree of malignancy | |||
| Low | 9(45%) | 0.0006±0.0004 | 0.0012** |
| High | 11(55%) | 0.0015±0.0006 | |
| Tumor necrosis rate (%) | |||
| <90 | 11(55%) | 0.0014±0.0007 | 0.0317* |
| ≥90 | 9(45%) | 0.0007±0.0005 |
P-value represents the probability from a Student's t-test for CBX4 mRNA expression between variable subgroups. *P<0.05, **P<0.01, which was considered to have a significant difference.
CBX4 was up-regulated in OS and correlated with advanced clinical stage. (a) (b) The mRNA and protein expression of CBX4 were measured by qRT-PCR and western blot assays in OS cell lines (U2OS, MNNG/HOS, MG63) and human normal osteoblastic cell line (hFOB); (c) Western blot test indicated CBX4 protein level was higher in 8 OS tumor tissues (T) than their noncancerous counterparts (N); β-Actin was used as the internal control. (d) (e) Relative expression of CBX4 was detected using qRT-PCR in 20 pairs of OS samples and their corresponding noncancerous samples. (f) CBX4 mRNA expression in different clinical stages; Patients were staged in accordance with the Ennecking musculoskeletal tumor staging system. Error bars represent mean ± s. d triplicate experiments. The statistical analysis was performed using paired t-test (e) and student's t-test (f).
Knockdown of CBX4 inhibited OS cells growth. (a) (b) Knockdown of CBX4 inhibited the cell proliferation in OS cells: The viability of MNNG/HOS and MG63 cell was examined by CCK8 assay at different time points as indicated. (c) (d) Flow cytometry of cell distribution in OS cells after transfection. (e) (f) Histograms of each phase in cell cycle of OS cells: Knockdown of CBX4 decreased cell proportion of G2/M phrase in MNNG/HOS and MG63 cells. Data represent the mean of three replicates. Each performed in triplicate. *p<0.05, **P<0.01, by student's t test.
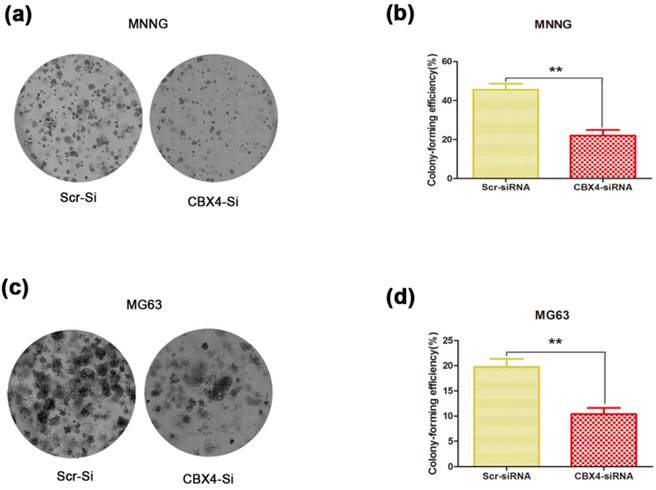
Decreased CBX4 impaired OS cell colony-forming capacity
To determine whether CBX4 knockdown affected the colony-forming ability of OS cells, the colony formation assay was performed. We observed that both the size and number of colonies were dramatically reduced in the CBX4 knockdown group compared with control group (Figure 3 a, c). The number of colonies was significantly reduced by 51.6% and 47.3% in MNNG/HOS (Figure 3 b) and MG63 (Figure 3 d) cells, respectively. In general, these results demonstrated that CBX4 knockdown could significantly impaired the colony-forming ability of OS cells, thus indicating the crucial role of CBX4 in OS cells growth.
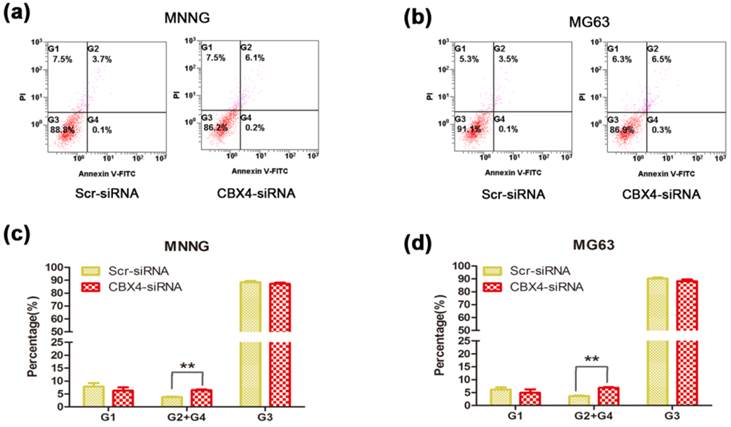
Silencing CBX4 attenuated OS cells survival by enhancing cells apoptosis rate
To determine the role of CBX4 in cell survival, we conduct the cell apoptosis assay to measure the changes of apoptosis rate. Knockdown of CBX4 remarkably increased the apoptosis rates in both MNNG/HOS cells (Figure 4 a) and MG63 cells (Figure 4 b). The apoptosis rates were significantly enhanced by 72.0% and 88.9% in MNNG/HOS cells (Figure 4 c) and MG63 cells (Figure 4 d), respectively. Taken together, these results demonstrated that CBX4 was involved in and required for OS cells survival.
Knockdown of CBX4 impaired colony-forming ability of OS cells. (a) (c) Representative photomicrographs of MNNG/HOS (a) and MG63 (c) colonies in culture plates. (b) (d) Significant reduction in the colony-forming efficacy in MNNG/HOS (b) and MG63 (d) cells following CBX4 knockdown. Data are expressed as mean ± s. d of three independent experiments, **P<0.01, by student's t test.
Knockdown of CBX4 increased the apoptosis rate of OS cells. (a)(b) Comparison of apoptosis between Scramble and CBX4-knockdown group in MNNG/HOS cells. (c)(d) Comparison of apoptosis between Scramble and CBX4-knockdown group in MG63 cells. The data represent mean ± s. d. of triplicate (b, d). *p<0.05, **p<0.01 by student's t-test. The cells were transfected with CBX4 siRNA (50nM) for 48 hours, and then cultured for 24 hours. All the apoptotic rates were measured by using the fluorescein isothiocyanate Annexin V apoptosis assay. PI, propidium iodide.
CBX4 was correlated with HIF-1α transactivation activity in OS cells. Cells were transfected with either Scramble or CBX4-siRNA under normoxic condition. (a) (b) Knockdown of CBX4 didn't change the expression of HIF-1α in MNNG/HOS (a) and MG63 (b) cells. (c) (d) Analyses of expression level of HIF-1α-targeted genes by qRT-PCR test: VEGFA, PDK1, HK2 in MNNG/HOS and MG63 cells (c, d); BNIP3 in MNNG/HOS cells (c); ANGPTL4 in MG63 cells (d). The data are representative of three independent experiments. Error bars represent mean ± s. d. **p<0.01 by student's t test.
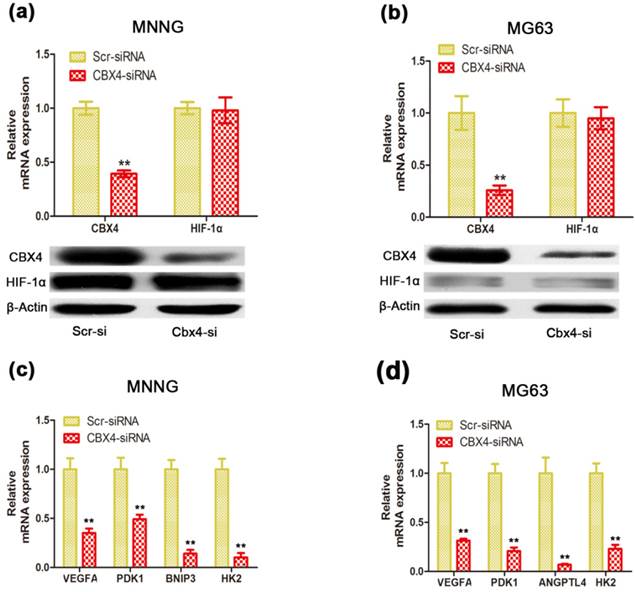
CBX4 knockdown suppressed the expression of HIF-1α-targeted genes without changing HIF-1α level under normoxic condition
Recent evidence showed that elevated CBX4, via its SUMO E3 ligase activity, contributed to hepatocellular carcinoma processes by regulating transactivation of HIF-1α (19). To investigate whether the decreased CBX4 inhibited cell growth and cell survival of OS cells via interaction with HIF-1α signaling, we found that knockdown of CBX4 didn't change the HIF-1α mRNA and protein expression in MNNG/HOS (Figure 4 a) and MG63 (Figure 4 b) cells. However, the mRNA levels of vascular endothelial growth factor (VEGFA), Angiopoietin-like 4 (ANGPTL4), Pyruvate dehydrogenase kinase 1 (PDK1), Hexokinase 2 (HK2), and BCL2/adenovirus E1B 19kDa interacting protein 3 (BNIP3), five classical HIF-1α downstream target genes, were suppressed in MNNG/HOS (Figure 4 c) and MG63 (Figure 4 d) cells under normoxic condition. These results indicated a distinctive role of CBX4 in OS, which facilitated HIF-1α signaling activation under normoxic condition.
Discussion
Polycomb group proteins (PcGs) have been reported to be linked with the occurrence of various cancers (5). Misregulation of PcGs always leads to activation of developmental pathways, thus enhancing the proliferation ability of cells (28). CBX4, a member of CBXs, is a unique PcG protein for its small ubiquitin-related modifier (SUMO) E3 ligase activity. Previous studies have reported that CBX4 regulated multiple key proteins, such as centrin-2 (29) and Bmi-1 (30), in the biological functions of cells via its SUMO E3 ligase activity. However, the study of CBX4 is far from enough, especially in the field of cancer research. Though studies of CBX7 have been widely reported in various cancers, little is known of CBX4. Previous studies reported that CBX7 was inactivated in various cancers, such as cancer of thyroid (31), breast (32), pancreas (33) et al. Interestingly, O'Loghlen et al (34) observed that the expression of CBX4 was directly repressed by CBX7 during embryonic stem cell differentiation, which indicated activation of CBX4 but loss of CBX7 in some cancers. Recent studies in hepatocellular cancer (18, 19, 35) have verified the speculation. In the present study, we observed that CBX4 was up-regulated and correlated to clinical progress in OS. To our best knowledge, we are the first to report the activation of CBX4 in OS.
The role of CBX4 in cancers was first identified in hepatocellular cancer. Wang et al (18) showed that knockdown of CBX4 resulted in inhibition of in vitro tumor growth, which indicated CBX4 to be an oncogene. More recently, Li et al (19) showed that CBX4 promoted the angiogenesis of hepatocellular cancer by regulating the proliferation, invasion and migration of tumor cells. Our in vitro data showed that knockdown of CBX4 resulted in significant inhibition of cellular growth and survival in OS cells. However, little effect of CBX4 on the metastasis of OS cells was observed in vitro.
Recent studies have demonstrated that the polycomb- and SUMO E3 ligase-dependent function of CBX4 were linked with its N terminus and two SIMs respectively (36, 37). Luis et al (38) reported that the anti-senescence and anti-differentiation functions of CBX4 were, respectively, regulated by the polycomb activity and SUMO ligase activity in human epidermal stem cells. Since Kagey et al (17) first identified CtBP1 as substrate for the SUMO E3 ligase activity of CBX4, increasing new substrates were conformed. Recent study reported that CBX4, via its SUMO ligase activity on HIF-1α, enhanced hypoxia-induced angiogenesis in hepatocellular carcinoma (19). In the present study, down-regulating of HIF-1α-targeted genes (with no change of HIF-1α) was observed following knockdown of CBX4 under normoxic condition in OS cells. The result was similar with previous study in hepatocellular carcinoma under hypoxic condition (19). However, it was not clear whether CBX4 exert its effect on OS via the polycomb-dependent or the SUMO E3 ligase-dependent function. Further investigation is wanted to unveil the deeper mechanism between CBX4 and HIF-1α pathway in OS.
HIF-1α, an oxygen-sensitive subunit of HIF-1, induces plenty of targeted genes expression, promoting tumor angiogenesis, growth and survival (21, 39-41). VEGFA, a well-known HIF-1-regulated gene, which is involved in regulating cell proliferation and blood vessel formation of both normal and cancer cells. ANGPTL4, best known for its role as an adipokine regulating the lipid and glucose metabolism, plays a critical role in cancer growth, anoikis resistance and metastasis. Other genes, including PDK1, HK2, and BNIP3, are also important HIF-1-targeted genes regulating tumor growth and survival. In the present study, we observed the changes of these genes regulated by CBX4, mechanically suggesting a pivotal role of CBX4 in regulating the biological progress of OS.
Generally, HIF-1α expression is relatively low under normoxic condition for the oxygen-dependent degradation. However, normoxic activation of HIF-1α pathway was observed in various cancers (24, 42-44), which seemed to have a more profound significance in cancer research. In this study, the activation of HIF-1α signaling was influenced by CBX4 under normoxic condition. For OS, HIF-1α expression was correlated to the tumor progress and prognosis in the clinic (45, 46). In vitro studies have showed that HIF-1α signaling promoted the cell growth and cell metastasis of human OS cells (47, 48). Therefore, it's undoubtedly that HIF-1α signaling is an attractive potential target for OS treatment.
In conclusion, we demonstrated for the first time that CBX4 was up-regulated in human OS tissues and cell lines. High level of CBX4 was correlated to advanced clinical stage, high degree of malignancy, and low tumor necrosis rate in OS, which might be illustrated by the fact that CBX4 affected the cell growth and survival capacity of OS cells. More importantly, we showed that CBX4 exert oncogenic function on OS via activating HIF-1α signaling. These data provided compelling evidence that CBX4 might provide a therapeutic target for OS.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of Shanghai Jiao Tong University and with the principles of Declaration of Helsinki.
Supplementary Material
Table S1 and Figure S1.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ottaviani G, Jaffe N. The epidemiology of osteosarcoma. Cancer Treat Res. 2009;152:3-13
2. Kager L, Zoubek A, Potschger U. et al. Primary metastatic osteosarcoma: presentation and outcome of patients treated on neoadjuvant Cooperative Osteosarcoma Study Group protocols. J Clin Oncol. 2003;21:2011-2018
3. Bielack SS, Kempf-Bielack B, Delling G. et al. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. J Clin Oncol. 2002;20:776-790
4. Sauvageau M, Sauvageau G. Polycomb group proteins: multi-faceted regulators of somatic stem cells and cancer. Cell Stem Cell. 2010;7:299-313
5. Richly H, Aloia L, Di Croce L. Roles of the Polycomb group proteins in stem cells and cancer. Cell Death Dis. 2011;2:e204
6. Morey L, Helin K. Polycomb group protein-mediated repression of transcription. Trends Biochem Sci. 2010;35:323-332
7. Gil J, Bernard D, Peters G. Role of polycomb group proteins in stem cell self-renewal and cancer. DNA Cell Biol. 2005;24:117-125
8. Martin-Perez D, Piris MA, Sanchez-Beato M. Polycomb proteins in hematologic malignancies. Blood. 2010;116:5465-5475
9. Bracken AP, Helin K. Polycomb group proteins: navigators of lineage pathways led astray in cancer. Nat Rev Cancer. 2009;9:773-784
10. Vandamme J, Volkel P, Rosnoblet C. et al. Interaction proteomics analysis of polycomb proteins defines distinct PRC1 complexes in mammalian cells. Mol Cell Proteomics. 2011;10:M110.002642
11. Bernard D, Martinez-Leal JF, Rizzo S. et al. CBX7 controls the growth of normal and tumor-derived prostate cells by repressing the Ink4a/Arf locus. Oncogene. 2005;24:5543-5551
12. Shinjo K, Yamashita Y, Yamamoto E. et al. Expression of chromobox homolog 7 (CBX7) is associated with poor prognosis in ovarian clear cell adenocarcinoma via TRAIL-induced apoptotic pathway regulation. Int J Cancer. 2014;135:308-318
13. Koppens M, van Lohuizen M. Context-dependent actions of Polycomb repressors in cancer. Oncogene. 2015
14. Clermont PL, Sun L, Crea F. et al. Genotranscriptomic meta-analysis of the Polycomb gene CBX2 in human cancers: initial evidence of an oncogenic role. Br J Cancer. 2014;111:1663-1672
15. Parris TZ, Aziz L, Kovacs A. et al. Clinical relevance of breast cancer-related genes as potential biomarkers for oral squamous cell carcinoma. BMC Cancer. 2014;14:324
16. Lee SH, Um SJ, Kim EJ. CBX8 suppresses Sirtinol-induced premature senescence in human breast cancer cells via cooperation with SIRT1. Cancer Lett. 2013;335:397-403
17. Kagey MH, Melhuish TA, Wotton D. The polycomb protein Pc2 is a SUMO E3. Cell. 2003;113:127-137
18. Wang B, Tang J, Liao D. et al. Chromobox homolog 4 is correlated with prognosis and tumor cell growth in hepatocellular carcinoma. Ann Surg Oncol. 2013;20(Suppl 3):S684-692
19. Li J, Xu Y, Long XD. et al. Cbx4 governs HIF-1alpha to potentiate angiogenesis of hepatocellular carcinoma by its SUMO E3 ligase activity. Cancer Cell. 2014;25:118-131
20. Talks KL, Turley H, Gatter KC. et al. The expression and distribution of the hypoxia-inducible factors HIF-1alpha and HIF-2alpha in normal human tissues, cancers, and tumor-associated macrophages. Am J Pathol. 2000;157:411-421
21. Semenza GL. Targeting HIF-1 for cancer therapy. Nat Rev Cancer. 2003;3:721-732
22. Bruick RK, McKnight SL. A conserved family of prolyl-4-hydroxylases that modify HIF. Science. 2001;294:1337-1340
23. Yang H, Xiong Y, Guan K. Metabolic alteration in tumorigenesis. Sci China Life Sci. 2013;56:1067-1075
24. Zhang M, Ma Q, Hu H. et al. Stem cell factor/c-kit signaling enhances invasion of pancreatic cancer cells via HIF-1alpha under normoxic condition. Cancer Lett. 2011;303:108-117
25. Yen ML, Chien CC, Chiu IM. et al. Multilineage differentiation and characterization of the human fetal osteoblastic 1.19 cell line: a possible in vitro model of human mesenchymal progenitors. Stem Cells. 2007;25:125-131
26. Kapinas K, Kessler C, Ricks T. et al. miR-29 modulates Wnt signaling in human osteoblasts through a positive feedback loop. J Biol Chem. 2010;285:25221-25231
27. Hu T, He N, Yang Y. et al. DEC2 expression is positively correlated with HIF-1 activation and the invasiveness of human osteosarcomas. J Exp Clin Cancer Res. 2015;34:22
28. Mills AA. Throwing the cancer switch: reciprocal roles of polycomb and trithorax proteins. Nat Rev Cancer. 2010;10:669-682
29. Klein UR, Nigg EA. SUMO-dependent regulation of centrin-2. J Cell Sci. 2009;122:3312-3321
30. Ismail IH, Gagne JP, Caron MC. et al. CBX4-mediated SUMO modification regulates BMI1 recruitment at sites of DNA damage. Nucleic Acids Res. 2012;40:5497-5510
31. Pallante P, Federico A, Berlingieri MT. et al. Loss of the CBX7 gene expression correlates with a highly malignant phenotype in thyroid cancer. Cancer Res. 2008;68:6770-6778
32. Mansueto G, Forzati F, Ferraro A. et al. Identification of a New Pathway for Tumor Progression: MicroRNA-181b Up-Regulation and CBX7 Down-Regulation by HMGA1 Protein. Genes Cancer. 2010;1:210-224
33. Karamitopoulou E, Pallante P, Zlobec I. et al. Loss of the CBX7 protein expression correlates with a more aggressive phenotype in pancreatic cancer. Eur J Cancer. 2010;46:1438-1444
34. O'Loghlen A, Munoz-Cabello AM, Gaspar-Maia A. et al. MicroRNA regulation of Cbx7 mediates a switch of Polycomb orthologs during ESC differentiation. Cell Stem Cell. 2012;10:33-46
35. Jiao HK, Xu Y, Li J. et al. Prognostic significance of Cbx4 expression and its beneficial effect for transarterial chemoembolization in hepatocellular carcinoma. Cell Death Dis. 2015;6:e1689
36. Fischle W, Wang Y, Jacobs SA. et al. Molecular basis for the discrimination of repressive methyl-lysine marks in histone H3 by Polycomb and HP1 chromodomains. Genes Dev. 2003;17:1870-1881
37. Merrill JC, Melhuish TA, Kagey MH. et al. A role for non-covalent SUMO interaction motifs in Pc2/CBX4 E3 activity. PLoS One. 2010;5:e8794
38. Luis NM, Morey L, Mejetta S. et al. Regulation of human epidermal stem cell proliferation and senescence requires polycomb- dependent and -independent functions of Cbx4. Cell Stem Cell. 2011;9:233-246
39. Harris AL. Hypoxia-a key regulatory factor in tumour growth. Nat Rev Cancer. 2002;2:38-47
40. Semenza GL. Defining the role of hypoxia-inducible factor 1 in cancer biology and therapeutics. Oncogene. 2010;29:625-634
41. Schwab LP, Peacock DL, Majumdar D. et al. Hypoxia-inducible factor 1alpha promotes primary tumor growth and tumor-initiating cell activity in breast cancer. Breast Cancer Res. 2012;14:R6
42. Zou C, Yu S, Xu Z. et al. ERRalpha augments HIF-1 signalling by directly interacting with HIF-1alpha in normoxic and hypoxic prostate cancer cells. J Pathol. 2014;233:61-73
43. Cao Y, Eble JM, Moon E. et al. Tumor cells upregulate normoxic HIF-1alpha in response to doxorubicin. Cancer Res. 2013;73:6230-6242
44. Agani F, Jiang BH. Oxygen-independent regulation of HIF-1: novel involvement of PI3K/AKT/mTOR pathway in cancer. Curr Cancer Drug Targets. 2013;13:245-251
45. Yang QC, Zeng BF, Dong Y. et al. Overexpression of hypoxia-inducible factor-1alpha in human osteosarcoma: correlation with clinicopathological parameters and survival outcome. Jpn J Clin Oncol. 2007;37:127-134
46. Mizobuchi H, Garcia-Castellano JM, Philip S. et al. Hypoxia markers in human osteosarcoma: an exploratory study. Clin Orthop Relat Res. 2008;466:2052-2059
47. Kuzmanov A, Wielockx B, Rezaei M. et al. Overexpression of factor inhibiting HIF-1 enhances vessel maturation and tumor growth via platelet-derived growth factor-C. Int J Cancer. 2012;131:E603-613
48. Guan G, Zhang Y, Lu Y. et al. The HIF-1alpha/CXCR4 pathway supports hypoxia-induced metastasis of human osteosarcoma cells. Cancer Lett. 2015;357:254-264
Author contact
![]() Corresponding authors: E-mail: tjyqccom; yingtaomailnet. Tel: +86-21-64369181 Fax: +86-21-64701361.
Corresponding authors: E-mail: tjyqccom; yingtaomailnet. Tel: +86-21-64369181 Fax: +86-21-64701361.

 Global reach, higher impact
Global reach, higher impact