3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2015; 6(3):292-301. doi:10.7150/jca.11038 This issue Cite
Research Paper
Controlling RECK miR21 Promotes Tumor Cell Invasion and Is Related to Biochemical Recurrence in Prostate Cancer
Laboratory of Medical Research, LIM55, Urology Department, University of Sao Paulo Medical School, Sao Paulo, Brazil
Received 2014-11-11; Accepted 2014-12-12; Published 2015-1-21
Abstract
The search for biomarkers to characterize prostate cancer aggressiveness has been the objective for the majority of researchers involved with the most prevalent tumor in men. MiRNAs are important for the control of many cellular functions and their deregulation is involved with tumor development and progression. To find miRNAs differentially expressed in prostate cancer and their relation to prognostic factors and biochemical recurrence we studied 53 surgical specimens from men who underwent radical prostatectomy, through a microarray analysis using the microarray platform (GeneChip® miRNA Array - Affymetrix) with more than 46,000 probes and 847 mature human miRNAs and transcripts. We defined different as an expression level greater or less than 1.1 with p<0.05. The validation study using qRT-PCR had confirmed miR21 as overexpressed in tumor that have recurred with a risk of 2.5. Transfection of miR-21 using lipid based assay in DU145 cell line, showed decrease in expression of RECK resulting in increase in expression of MMP9. Invasion assay with Matrigel showed increase in tumor cell invasion after miR-21 transfection. We conclude that miR-21 overexpression is related to increased biochemical recurrence after surgical treatment of prostate cancer. And the negative control of RECK results in overexpression of MMP9 promotes increasing tumor cell invasion supporting miR-21 as an oncomiR related to aggressiveness in prostate cancer.
Keywords: Prostate cancer, Micro RNA, Microarray, Biochemical recurrence, Prognosis, miR-21, Cell invasion, RECK, DU145
Introduction
Prostate cancer (PC) aggressiveness must be characterized because there are various options of patient management. Active surveillance and focal therapy or brachytherapy are suitable for patients with low-risk PC, whereas radical prostatectomy, external beam radiation and hormone therapy are more appropriate for high-risk PC 1. Risk is based on the clinical stage, prostate specific antigen (PSA) serum levels and Gleason score. The Gleason score is the most significant independent prognostic parameter; however, it is somewhat inaccurate 2,3. Molecular biologists have worked to identify new risk parameters but have generated disappointing results thus far.
MicroRNAs (miRNAs) are a class of small, noncoding RNAs that regulate protein-coding gene expression by triggering degradation or preventing target messenger RNA (mRNA) translation 4. miRNAs are located in chromosomal regions that are frequently altered in cancer; miRNAs that promote oncogenesis are “oncomiRs,” and those that inhibit carcinogenesis are tumor suppressor miRNAs (“tsmiRs”). Compared with messenger RNA, miRNA expression profiles better classify undifferentiated tumors, which indicates that miRNAs are more tissue-specific and more accurately reflect the characteristics of certain tumors 5.
miRNA expression profiling studies have been performed to identify cancer-specific miRNA signatures, and certain miRNAs are aberrantly expressed in PC. However, validation studies remain necessary to identify miRNAs that are altered in numerous samples and should be considered aggressiveness markers. Few reports have correlated miRNA expression with PC behavior 6,7. Identifying microRNA subsets related to PC aggressiveness would be useful in determining the best management strategy for individual patients.
In this study, we utilized a genome-wide association analysis and GeneChip® miRNA microarray, version 1 (Affymetrix, CA, USA) to identify differentially expressed miRNAs in 53 PC specimens from patients who underwent radical prostatectomy to treat localized cancer. The miRNA profiles were compared between patients who had biochemical recurrence with those who have not in a period of 46.9-month follow-up.
Methods
Ethics statement
The study was conducted according to the principles of the Helsinki Declaration. The study was approved by the local ethics committee under protocol # 0039/11 and all individuals signed an informed consent to participate in the study.
Clinical samples
From 260 patients that underwent a radical prostatectomy by the same surgeon (MS) to treat localized PC between August 2000 and June 2002, 53 were included in the study. The inclusion criteria were the absence of neoadjuvant anti-androgen treatment and the presence of at least 75% of tumor in the frozen specimen stored in the bio repository. The mean age of patients was 65 years (SD: 7.5, range: 49 - 77). Gleason score was graded according to the recommendations of the 2005 consensus meeting of the International Society of Urological Pathology (ISUP)8. The mean Gleason score was 7 (SD: 1.1, range: 5 - 10); 15 tumors were scored at 5 or 6, and 38 were scored between 7 and 10. Twenty-four (45.3%) patients were staged pT2 when tumor was confined to the prostate gland, and 29 (54.7%) were staged pT3 when tumor extended to extraprostatic tissue or infiltrated seminal vesicles. Thirteen (24.5%) patients recurred with a 33.24-month mean with a 3.25 ng/mL mean PSA serum level (SD: 5.8 ng/mL). One cm3 of fresh prostate tissue was excised immediately after gland removal and stored at 80°C. For each specimen, a mirror fragment was formalin-fixed, routinely processed and stained with hematoxylin and eosin to ensure that at least 75% of the sample was tumor tissue. The entire gland was fixed in formalin and analyzed for the Gleason score and pathological stage (TNM 2010). The specimens were examined by the same uropathologist (KRML); the patients were followed for a 46.9-month mean (median: 34.3 months, range: 2.0 - 120.2 months). PSA serum levels over 0.2 ng/mL indicated biochemical recurrence. To validate the microarray data, tumors from the 51 patients in the test group were analyzed through quantitative real-time polymerase chain reaction (qRT-PCR). An additional validation group with 76 patients, 61.5-year mean age (SD: 7.6, range: 46 - 79) was also analyzed. The same inclusion criteria were used for this group. Their Gleason score was <7 in 29 (38.2%) tumors, and 38 (50%) tumors were at stage pT2. Thirty-seven (48.7%) patients exhibited biochemical recurrence. The control group was the same used for the microarray analysis and comprises of three patients with lower urinary tract symptoms due to benign prostatic hyperplasia (BPH) treated through open surgery with a 42.7-g mean prostate gland size. The mean age of this group was 55.3-year. It is important to mention that both techniques require a normal control for comparative analysis and this may be considered critics for the interpretation of the results. The use of normal prostate tissue from the same patient with cancer is questionable since the majority of prostate cancer are multifocal and molecular alterations may be already present without phenotypical alterations. Also, Lexander et al have published the same protein profile between peripheral and central zone of the prostate that we believe endorses our procedure 9.
RNA isolation, quantification, and quality control
Frozen prostate tissue fragments were placed in a 1.5 mL microtube with 500 µl of lysis buffer from the mirVana miRNA isolation kit (Ambion, Grand Island, NY, USA) and 2 5-mm stainless steel beads; the fragments were macerated in a TissueLyser LT (Qiagen, Germantown, MD, USA) for 2 min. The mRNA and miRNA were isolated using a mirVana Kit® (Applied Biosystems, CA, USA) in accordance with the manufacturer's instructions, and the nucleic acid concentrations were calculated based on absorbance at 260/280 nM using a Nanodrop® ND-1000 spectrophotometer (Thermo Scientific, West Palm Beach, FL, USA).
Affymetrix miRNA labeling, array hybridization and data pre-processing
The total RNA, which includes low molecular weight RNA, was labeled using the FlashTag RNA labeling kit (Genisphere, Hatfield, PA, USA) in accordance with the manufacturer's instructions. For each sample, 2 µg of total RNA was used in a tailing reaction (incubated in 2.5 mM MnCl2, ATP and poly(A) polymerase for 15 min at 37°C) followed by biotinylated signal molecule ligation to the target RNA sample (1X Flash Tag ligation mix, biotin and T4 DNA ligase, which were incubated for 30 min at RT) and stop solution addition. Each sample was hybridized to a GeneChip® miRNA Array (Affymetrix, Santa Clara, CA, USA) at 48°C and 60 rpm for 16 h then washed and stained using a Fluidics Station 450 (Fluidics script FS450_0003) and, finally, scanned on a GeneChip® Scanner 3000 7G (Affymetrix, Santa Clara, CA, USA). The image data were analyzed using the miRNA QC Tool software for quality control (www.affymetrix.com/products_services/arrays/specific/mi_rna.affx#1_4).
Microarray data normalization and statistical analysis
Affymetrix chips were normalized using the robust multichip analysis (RMA) procedure 10. Median-centric normalization was used for the custom miRNA oligonucleotide chips. The genes with significant expression differences were identified based on the fold change and p values; biochemical recurrence were considered when PSA>0.2 ng/mL. For each comparison, we only considered human miRNA. The microarray data were deposited in the NCBI Gene Expression Omnibus under accession number GSE46738 (http://www.ncbi.nlm.nih.gov/geo).
Quantitative real-time PCR for miRNAs
miR-21, 92, 885, 450, 1226, 16 and 1202 expression was validated using the commercially available pre-designed TaqMan RT-qPCR assay (Applied Biosystems, Foster City, CA, USA). The RNA extraction procedure was similar to the above description, and the TaqMan® MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) was used to prepare the cDNA. The reverse transcription reactions (20 µL) were performed at 16°C for 30 min, 42°C for 30 min, and at 85°C for 5 min to terminate the reaction. Each reaction included 20 ng of total RNA, multiple (heptaplex) stem-loop miRNA-specific primers from the TaqMan MicroRNA Assays (12.5 nM each), 2 mM dNTPs, 100 U MultiScribe™ Reverse Transcriptase, 1X Reverse Transcription buffer, and 5 U RNase Inhibitor. The hsa-miRs target sequences were amplified in a 10 μL reaction with 5 μL of TaqMan Universal PCR Master Mix, 0.5 μL from the TaqMan miRNA expression assays, 1 μL of cDNA and 3.5 μL of DNase-free water. The PCR cycling conditions were as follows: 2 min at 50°C, 10 min at 95°C, 40 cycles at 15 sec and 95°C and 1 min at 60°C. RNU43 was the endogenous control. The ΔΔCT method was used to calculate the relative miRNA expression: ΔΔCT = (PC sample CT miRNA - PC sample CT endogenous control) - (BPH sample CT miRNA - BPH sample CT endogenous control). The expression fold change was calculated as 2-ΔΔCT.
miR-21 and anti-miR-21 transfection, qRT-PCR for RECK and MMP9 expression
The prostate cancer cell line DU145 was maintained in McCoy's medium with 20% FBS and 1% antibiotic/antimycotic solution (Sigma Co, St. Louis, MO, USA) at 37°C in an atmosphere with 5% CO2. The cells were grown in 24-well plates, 5.0 x 104 cells were seeded into each well. The transfections were performed in Opti-MEM I with Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) and diluted 1:100 in accordance with the manufacturer's protocol. Synthetic miR-21 and antimiR-21 as well as their respective scrambled controls were purchased from Ambion (Austin, TX, USA). The negative controls and miRs were used at 50 nM. The total RNA for quantitative PCR was isolated using the Ambion mirVana kit (Austin, TX, USA) as recommended by the manufacturer. qRT-PCR was performed using the TaqMan assays from Applied Biosystems (Foster City, CA, USA) and 2 mg RNA that was reverse transcribed with random primers and MultiScribe reverse transcriptase (Applied Biosystems, Foster City, CA, USA). After reverse transcription, the samples were analyzed using the comparative DDCt method with human ß2 microglobulin as the internal standard. The gene expression fold change was 2-ΔΔCT. The data shown represent three independent experiments.
Invasion assay
The BD BioCoat Matrigel Invasion Chamber (Becton Dickinson, Bedford, MA) included an 8 μm pore size PET membrane with a Matrigel Basement Membrane matrix. The transfection method was similar the above description. The cells were collected through trypsinization followed by seeding 2 × 104 cells in the internal chamber with FBS-free media. The lower chamber was filled with media that contained 10% FBS, which is a chemo attractant. The cells were incubated for 24 h at 37 °C in a 5% CO2 atmosphere. After incubation, the non-invading cells were removed from the upper surface of the membrane using a cotton-tipped swab. The cells on the lower surface of the membrane were embedded using paraformaldehyde and stained with crystal violet at 0.2%; the inserts were then desiccated to room temperature. The cells were counted under a microscope at 200X magnification to determine the number of colonies and 400X to determine the total number of cells and number of cells per colony. The experiments were performed in triplicate.
Statistical Analysis
Fold change and p-value were used to analyze the microarray experiments. These values were determined by statistical T tests. Differential in gene expression was considered higher as lower was p and higher was the fold change. The choice for statistical tests was based in the determination of the groups of samples to be analyzed. For independent samples, simple T test was used. For groups with differences in variance, Welch T test was used. The fold change value was obtained by the equation FC=Xcase/Xctrl were Xcase and Xctrl corresponding to the mean of the case and control groups. To analyze more than two groups, the ANOVA test was used. For quantitative variables we used T test and Mann-Whitney. miR-21 expression and biochemical recurrence was analyzed by Mann-Whitney test. A Kaplan-Meier curve was designed and the Cox regression model was used for multivariate analysis. For the miR-21 and anti-miR-21 transfections as well as invasion assay, the statistical significance was calculated by Student's T test. The statistical analyses were performed using SPSS 15.0 for Windows with 5% significance.
Results
Differentially expressed microRNAs in prostate cancer
Compared with the three BPH tissue controls, 20 miRNAs were underexpressed, and 57 were overexpressed in prostate cancer specimens. The data are presented in Table 1. miRNAs were considered overexpressed or underexpressed when the fold change was greater or than 1.1 with p<0.05, respectively.
miRNA expression, prognostic factors and biochemical recurrence in prostate cancer
The results comparing different prognostic factors and miRNA expression are exposed in table 2. Dividing Gleason score in <7 and ≥7, 5 miRNAs were underexpressed and 9 overexpressed. Considering pre-operatory PSA levels ≤10ng/mL and >10ng/mL, 21 miRNAs were underexpressed and 29 overexpressed. Considering pathological stage, divided as organ confined (pT2) and non-organ confined (pT3), 21 miRNAs were underexpressed and 22 overexpressed.
PSA levels greater than 0.2 ng/mL indicate biochemical recurrence, and the differentially expressed miRNAs under such conditions are presented in Table 3.
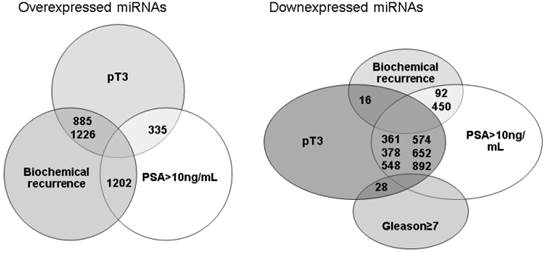
When all results were compared, there were few miRNAs commonly under or overexpressed by different unfavorable prognostic factors and related to biochemical recurrence (Figure 1). miRNAs 92, 450 and 16 were downexpressed by tumors that have recurred and were non organ-confined or when the pre-operatory PSA was >10ng/mL. Whereas miRNAs 885, 1226 and 1202 were overexpressed by tumors that have recurred and were non organ-confined or the pre-operatory PSA was >10ng/mL. The validation by qRT-PCR of these miRNAs in the same group of patients previously studied using the microarray did not confirm a relationship between their expression, prognostic factors or biochemical recurrence.
miRNAs simultaneously under or overexpressed considering different unfavorable prognostic factors and biochemical recurrence after radical prostatectomy.
Differentially expressed micro RNA between prostate cancer and BPH
| Underexpressed miRNAs | Overexpressed miRNAs | ||||
|---|---|---|---|---|---|
| Gene name | fold change | p-value | Gene name | fold change | p-value |
| hsa-let-7d_st | -1.29 | 0.015 | hsa-let-7b-star_st | 1.65 | <0.001 |
| hsa-miR-1178_st | -1.18 | 0.005 | hsa-miR-1182_st | 1.33 | 0.001 |
| hsa-miR-1206_st | -1.17 | 0.036 | hsa-miR-1226-star_st | 3.79 | 0.006 |
| hsa-miR-146a-star_st | -1.63 | 0.017 | hsa-miR-1231_st | 11.4 | 0.022 |
| hsa-miR-195_st | -1.28 | 0.050 | hsa-miR-124_st | 1.86 | 0.001 |
| hsa-miR-320c_st | -1.23 | 0.001 | hsa-miR-1258_st | 1.23 | 0.013 |
| hsa-miR-376a-star_st | -1.43 | 0.042 | hsa-miR-1276_st | 1.11 | 0.023 |
| hsa-miR-380-star_st | -1.17 | 0.012 | hsa-miR-128_st | 2.68 | 0.017 |
| hsa-miR-384_st | -1.23 | <0.001 | hsa-miR-1303_st | 1.49 | 0.039 |
| hsa-miR-450a_st | -1.41 | <0.001 | hsa-miR-1307_st | 8.91 | 0.032 |
| hsa-miR-518e_st | -1.22 | 0.032 | hsa-miR-138_st | 9.73 | <0.001 |
| hsa-miR-523-star_st | -1.32 | 0.005 | hsa-miR-145-star_st | 1.75 | <0.001 |
| hsa-miR-566_st | -1.41 | 0.029 | hsa-miR-148a_st | 5.62 | 0.035 |
| hsa-miR-591_st | -1.95 | 0.006 | hsa-miR-148b_st | 2.11 | <0.001 |
| hsa-miR-633_st | -2.46 | 0.024 | hsa-miR-183_st | 6.36 | 0.001 |
| hsa-miR-758_st | -1.22 | 0.026 | hsa-miR-183-star_st | 2.71 | 0.022 |
| hsa-miR-877_st | -1.55 | 0.026 | hsa-miR-195-star_st | 3.07 | <0.001 |
| hsa-miR-886-5p_st | -3.10 | <0.001 | hsa-miR-19a_st | 2.12 | 0.038 |
| hsa-miR-923_st | -5.10 | 0.009 | hsa-miR-200a-star_st | 5.14 | 0.010 |
| hsa-miR-96-star_st | -1.69 | 0.023 | hsa-miR-206_st | 1.47 | 0.041 |
| hsa-miR-212_st | 3.6 | 0.015 | |||
| hsa-miR-22-star_st | 1.77 | <0.001 | |||
| hsa-miR-27a-star_st | 2.70 | 0.007 | |||
| hsa-miR-29c-star_st | 2.48 | <0.001 | |||
| hsa-miR-300_st | 1.36 | <0.001 | |||
| hsa-miR-330-5p_st | 1.21 | 0.032 | |||
| hsa-miR-339-5p_st | 7.83 | 0.036 | |||
| hsa-miR-33b-star_st | 1.71 | 0.047 | |||
| hsa-miR-346_st | 4.78 | <0.001 | |||
| hsa-miR-34b-star_st | 2.02 | 0.010 | |||
| hsa-miR-363-star_st | 4.16 | <0.001 | |||
| hsa-miR-373-star_st | 2.34 | <0.001 | |||
| hsa-miR-429_st | 1.29 | <0.001 | |||
| hsa-miR-431-star_st | 1.36 | <0.001 | |||
| hsa-miR-449a_st | 1.62 | 0.012 | |||
| hsa-miR-484_st | 1.46 | 0.020 | |||
| hsa-miR-498_st | 2.38 | 0.005 | |||
| hsa-miR-508-5p_st | 1.22 | 0.014 | |||
| hsa-miR-509-5p_st | 1.64 | 0.040 | |||
| hsa-miR-541_st | 1.79 | <0.001 | |||
| hsa-miR-557_st | 1.88 | 0.022 | |||
| hsa-miR-596_st | 1.82 | 0.004 | |||
| hsa-miR-602_st | 4.87 | 0.020 | |||
| hsa-miR-606_st | 2.31 | 0.036 | |||
| hsa-miR-624_st | 1.31 | 0.034 | |||
| hsa-miR-629_st | 2.13 | 0.002 | |||
| hsa-miR-638_st | 2.03 | 0.025 | |||
| hsa-miR-663b_st | 4.86 | 0.001 | |||
| hsa-miR-668_st | 1.52 | 0.009 | |||
| hsa-miR-7-1-star_st | 1.90 | 0.020 | |||
| hsa-miR-769-5p_st | 4.27 | 0.042 | |||
| hsa-miR-885-5p_st | 2.87 | 0.006 | |||
| hsa-miR-890_st | 2.00 | 0.006 | |||
| hsa-miR-92a-1-star_st | 5.06 | <0.001 | |||
| hsa-miR-92a-2-star_st | 1.74 | 0.002 | |||
| hsa-miR-943_st | 2.03 | <0.001 | |||
| hsa-miR-98_st | 1.67 | 0.024 | |||
Differentially expressed micro RNAs considering the prognostic parameters in prostate cancer
| Underexpressed micro RNAs | Overexpressed micro RNAs | ||||
|---|---|---|---|---|---|
| Gene name | fold change | p-value | Gene name | fold change | p-value |
| Gleason score <7 vs ≥7 | |||||
| hsa-miR-122-star_st | -1.52 | 0.049 | hsa-miR-539_st | 1.20 | 0.013 |
| hsa-miR-20a-star_st | -1.46 | 0.026 | hsa-miR-1197_st | 1.20 | 0.020 |
| hsa-miR-1244_st | -1.35 | 0.044 | hsa-miR-613_st | 1.21 | 0.003 |
| hsa-miR-876-3p_st | -1.22 | 0.047 | hsa-miR-429_st | 1.24 | 0.043 |
| hsa-miR-338-3p_st | -1.14 | 0.016 | hsa-miR-28-5p_st | 1.27 | 0.014 |
| hsa-miR-597_st | 1.28 | 0.015 | |||
| hsa-miR-483-3p_st | 1.28 | 0.048 | |||
| hsa-miR-708-star_st | 1.33 | 0.017 | |||
| hsa-miR-200b-star_st | 1.51 | 0.009 | |||
| PSA ≤10 ng/mL vs > 10 ng/mL | |||||
| hsa-miR-146b-5p_st | -1.78 | 0.016 | hsa-miR-892a_st | 1.12 | 0.029 |
| hsa-miR-146a_st | -1.77 | 0.031 | hsa-miR-548j_st | 1.13 | 0.032 |
| hsa-miR-183_st | -1.74 | 0.022 | hsa-miR-655_st | 1.13 | 0.049 |
| hsa-miR-25_st | -1.66 | 0.022 | hsa-miR-1283_st | 1.14 | 0.019 |
| hsa-miR-335_st | -1.66 | 0.033 | hsa-miR-632_st | 1.14 | 0.025 |
| hsa-miR-363_st | -1.66 | 0.035 | hsa-miR-490-3p_st | 1.14 | 0.038 |
| hsa-miR-199b-5p_st | -1.57 | 0.048 | hsa-miR-875-5p_st | 1.15 | 0.011 |
| hsa-miR-200c-star_st | -1.55 | 0.025 | hsa-miR-1206_st | 1.17 | 0.027 |
| hsa-miR-501-5p_st | -1.55 | 0.037 | hsa-miR-338-3p_st | 1.17 | 0.028 |
| hsa-miR-30b_st | -1.55 | 0.038 | hsa-miR-218_st | 1.17 | 0.033 |
| hsa-miR-629_st | -1.54 | 0.029 | hsa-miR-450a_st | 1.18 | 0.011 |
| hsa-miR-487a_st | -1.53 | 0.036 | hsa-miR-1277_st | 1.18 | 0.040 |
| hsa-miR-1202_st | -1.51 | 0.023 | hsa-miR-301b_st | 1.20 | 0.003 |
| hsa-miR-376a-star_st | -1.35 | 0.019 | hsa-miR-566_st | 1.20 | 0.014 |
| hsa-miR-1294_st | -1.29 | 0.043 | hsa-miR-551a_st | 1.21 | 0.038 |
| hsa-miR-130b_st | -1.28 | 0.049 | hsa-miR-361-5p_st | 1.22 | 0.011 |
| hsa-miR-548b-5p_st | -1.25 | 0.033 | hsa-let-7e_st | 1.22 | 0.021 |
| hsa-miR-873_st | -1.23 | 0.010 | hsa-miR-1182_st | 1.23 | 0.036 |
| hsa-miR-369-5p_st | -1.21 | 0.038 | hsa-miR-644_st | 1.29 | <0.001 |
| hsa-miR-425_st | -1.19 | 0.047 | hsa-miR-196a_st | 1.29 | 0.049 |
| hsa-miR-507_st | -1.16 | 0.033 | hsa-miR-132_st | 1.31 | 0.016 |
| hsa-miR-187-star_st | 1.34 | 0.034 | |||
| hsa-miR-92b_st | 1.35 | 0.028 | |||
| hsa-miR-378_st | 1.36 | 0.037 | |||
| hsa-miR-548a-3p_st | 1.38 | 0.042 | |||
| hsa-miR-574-5p_st | 1.38 | 0.048 | |||
| hsa-miR-652_st | 1.39 | 0.037 | |||
| hsa-miR-1323_st | 1.49 | 0.017 | |||
| hsa-miR-1_st | 1.61 | 0.021 | |||
| pT2 vs pT3 | |||||
| hsa-miR-885-3p_st | -2.49 | 0.010 | hsa-miR-553_st | 1.12 | 0.049 |
| hsa-miR-1231_st | -2.18 | 0.001 | hsa-miR-526b-star_st | 1.13 | 0.023 |
| hsa-miR-129-star_st | -1.95 | 0.004 | hsa-miR-548n_st | 1.14 | 0.013 |
| hsa-miR-335_st | -1.82 | 0.026 | hsa-miR-892a_st | 1.14 | 0.016 |
| hsa-miR-606_st | -1.81 | 0.002 | hsa-miR-361-5p_st | 1.16 | 0.027 |
| hsa-miR-129-3p_st | -1.69 | 0.025 | hsa-miR-522_st | 1.18 | 0.041 |
| hsa-miR-637_st | -1.58 | 0.028 | hsa-miR-16-1-star_st | 1.18 | 0.048 |
| hsa-miR-1226-star_st | -1.53 | 0.032 | hsa-miR-1259_st | 1.21 | 0.002 |
| hsa-miR-576-3p_st | -1.52 | 0.012 | hsa-miR-151-3p_st | 1.22 | 0.041 |
| hsa-miR-92b-star_st | -1.51 | 0.039 | hsa-miR-28-5p_st | 1.24 | 0.010 |
| hsa-miR-96_st | -1.45 | <0.001 | hsa-miR-485-3p_st | 1.24 | 0.021 |
| hsa-miR-559_st | -1.41 | 0.005 | hsa-miR-372_st | 1.25 | 0.006 |
| hsa-miR-1228-star_st | -1.38 | 0.030 | hsa-miR-30d_st | 1.27 | 0.028 |
| hsa-miR-15a-star_st | -1.35 | 0.029 | hsa-miR-574-5p_st | 1.34 | 0.033 |
| hsa-miR-330-5p_st | -1.33 | <0.001 | hsa-miR-378_st | 1.37 | 0.011 |
| hsa-miR-1302_st | -1.33 | 0.023 | hsa-miR-616-star_st | 1.37 | 0.022 |
| hsa-miR-202-star_st | -1.31 | 0.022 | hsa-miR-652_st | 1.37 | 0.024 |
| hsa-miR-518d-3p_st | -1.17 | 0.020 | hsa-miR-766_st | 1.51 | 0.005 |
| hsa-miR-362-3p_st | -1.17 | 0.036 | hsa-miR-1263_st | 1.53 | 0.026 |
| hsa-miR-1258_st | -1.16 | 0.030 | hsa-miR-106b-star_st | 1.54 | 0.041 |
| hsa-miR-645_st | -1.15 | 0.034 | hsa-miR-1287_st | 1.58 | 0.037 |
| hsa-miR-378-star_st | 1.69 | 0.011 | |||
MicroRNA down an up-regulated in men with vs. without recurrent disease
| Underexpressed micro RNAs | Overexpressed micro RNAs | |||||
|---|---|---|---|---|---|---|
| micro RNA | Fold change | p | micro RNA | Fold change | p | |
| hsa-miR-129-3p_st | -1.64 | 0.041 | hsa-miR-1202_st | 1.75 | 0.019 | |
| hsa-miR-450b-5p_st | -1.63 | 0.004 | hsa-miR-1238_st | 1.78 | 0.022 | |
| hsa-miR-606_st | -1.59 | 0.014 | hsa-miR-31_st | 1.79 | 0.017 | |
| hsa-miR-16-2-star_st | -1.57 | 0.003 | hsa-miR-210_st | 1.80 | 0.014 | |
| hsa-miR-340_st | -1.56 | <0.001 | hsa-miR-33b-star_st | 1.80 | 0.014 | |
| hsa-miR-541_st | -1.56 | <0.001 | hsa-miR-1226-star_st | 1.81 | 0.014 | |
| hsa-miR-510_st | -1.55 | 0.005 | hsa-miR-181a-2-star_st | 1.81 | 0.029 | |
| hsa-miR-570_st | -1.54 | 0.006 | hsa-miR-370_st | 1.87 | 0.003 | |
| hsa-miR-550-star_st | -1.52 | 0.005 | hsa-miR-181c-star_st | 1.92 | 0.042 | |
| hsa-miR-890_st | -1.51 | 0.005 | hsa-miR-671-5p_st | 1.93 | <0.001 | |
| hsa-miR-22-star_st | -1.47 | 0.008 | hsa-miR-602_st | 1.93 | 0.018 | |
| hsa-miR-145-star_st | -1.46 | 0.001 | hsa-miR-191-star_st | 1.95 | 0.014 | |
| hsa-miR-624_st | -1.41 | <0.001 | hsa-miR-34c-3p_st | 2.00 | 0.020 | |
| hsa-miR-92a-2-star_st | -1.41 | 0.047 | hsa-miR-498_st | 2.07 | 0.008 | |
| hsa-miR-21-star_st | 2.12 | 0.050 | ||||
| hsa-miR-373-star_st | 2.27 | 0.006 | ||||
| hsa-miR-885-3p_st | 2.54 | <0.001 | ||||
The Kaplan-Meier curve shows that miR-21 expression levels greater than 0.64 are significantly related to biochemical recurrence (p=0.003). The Cox regression model revealed that only miR-21 expression is independently related to tumor recurrence with a 2.5 risk.
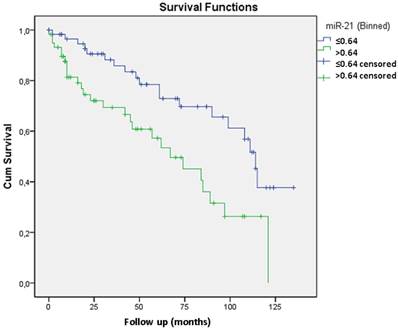
Considering that miR-21 was overexpressed in recurrent tumors and its role has already been described in other neoplasias we also proceed with the validation study of its expression by qRT-PCR 11. In prostate cancer there are only two clinical studies relating miR-21 with biochemical recurrence and with resistance to docetaxel-based chemotherapy 12 13. We analyzed miR-21 through qRT-PCR in 51 cases from the test group (included in the microarray analysis) and in 76 additional cases. Validation through qRT-PCR confirmed that miR-21 overexpression was related to biochemical recurrence. The mean expression for miR-21 in recurrent tumors was 7.20 compared with 2.21 in non-recurrent tumors (p=0.014). A Kaplan-Meier curve was generated, which indicated that expression levels greater than 0.64 were significantly related to biochemical recurrence (p=0.003) (Figure 2). Including Gleason grade <7 and ≥7, preoperatory PSA level < 10 ng/ml and ≥ 10 ng/ml and pT2 and pT3 tumors the Cox regression model revealed that only miR-21 expression was independently related to tumor recurrence with a 2.5 risk.
MMP9 and RECK expression after miR-21 transfection in a DU145 cell line
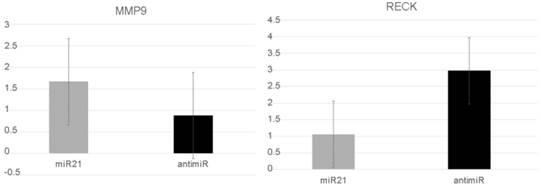
In vitro assays were performed to explore miR-21's role in more aggressive PC behavior. After miR-21 transfection, the RECK levels were significantly reduced compared with cells transfected using anti-miR-21. The mean level was 1.06 (SD: 0.55) compared with 2.97 (SD:2.2) (p=0.05). In contrast, the MMP9 levels increased after miR-21 transfection from 1.67 (SD:0.8) to 0.88 (SD:0.13), but this difference was not significantly different (p=0.127) (Figure 3).
RECK and MMP9 expression levels after the prostate cancer cells DU145 were transfected with miR21 and anti-miR21.
Invasion assay showing higher number of cells pos-transfection of miR-21
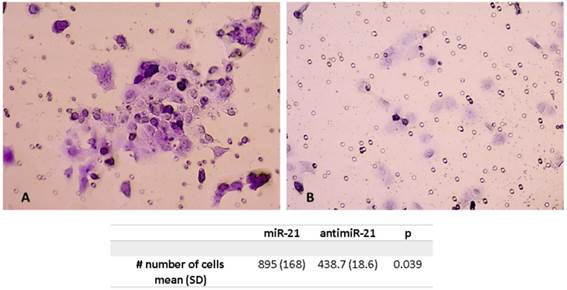
Invasion Assay
The mean number of cells after miR-21 transfection were significantly higher compared to the scramble control. The mean number of tumor cells was 895 (SD 168) after miR21 transfection compared to 438.7 (SD 18.6) for the control (p=0.039) (Figure 4).
Discussion
By analyzing 53 men with localized prostate cancer that was surgically treated, we demonstrated that tumors are more often correlated with overexpressed than underexpressed miRNAs. Additional published genome-wide association prostate cancer analyses have yielded conflicting results. Similar to our study, Volinia et al. 14 primarily identified overexpressed miRNAs, whereas Porkka et al. 15 and Ozen et al. 16 predominantly observed underexpressed miRNAs. The study by Volinia et al. was similar to the study herein, including the number of cases, but the remaining two studies used few clinical samples and included cell lines and xenografts that may have compromised the results. We also found a prevalence of overexpressed miRNAs by potentially more aggressive tumors. A result different from those published by Ozen et. al 16 and Tong et al 17 that found more underexpressed miRNAs.
miRNAs have hundreds of target genes and each gene have 3'UTR seed regions for many different miRNAs. miRNAs overexpressed in cancer and related to unfavorable prognosis are considered oncomiRs and have as target tumor suppressor genes being negatively controlled. Oppositely, underexpressed miRNAs are considered tumor suppressor miRNAs or tsmiRs, and they theoretically control oncogenes negatively18. The search of miRNAs profiles related to tumor behavior can bring to light new molecules to be used as prognostic or predictive markers. Moreover, offers an opportunity to design new forms of treatment given that inhibiting overexpressed molecules is easier than supplementing decreased molecules, enzymes and proteins.
The most relevant result of our study was the identification of the miRNAs that were differentially expressed in recurrent tumors. Again, there were more overexpressed than underexpressed miRNAs, meaning that they act more as oncomiRs than tsmiRs. Schaefer et al 19 also demonstrated overexpressed miRNAs related to biochemical recurrence in PC, validating miR-96 as a potential marker for disease behavior. Distinguishing aggressive PC is clinically important to determining initial therapy and, subsequently, an appropriate post-surgery adjuvant treatment, which may entail radiotherapy and anti-androgen therapy, both of which can notably affect quality of life. It is well known that the Gleason score and tumor stage are the primary means to determine tumor behavior 20, but they frequently incorrectly classify the disease aggressiveness, which results in patient under- or over-treatment. Recently Haffner et al have published tumor progression in tumors classified as Gleason 6 (3+3), promoting a debate in the benign behavior of the well differentiated PC 21,22.
In this study we have split tumors in low grade, Gleason 6 or less, and high grade, Gleason 7 or higher. However it is import to mention that recent reviews show that the percentage of Gleason pattern 4 and the presence of pattern 5 have important influence in the behavior of PC23. The same occurs with tumor stage, where pT3b disease (infiltration of seminal vesicles) behaves more aggressively than extraprostatic extension only (pT3a)24.
In this study, we confirmed that miR-21 was overexpressed in recurrent prostate cancer. miR-21 expression levels greater than 0.64 were independently related to biochemical recurrence after RP with a 2.5 risk. We have previously reported high levels of miR-21 in high grade prostatic intraepithelial neoplasia7. Together with the upregulation demonstrated in recurrent prostate cancer we could hypothesize, that miR-21 is important for tumor invasion and has a role in tumor dissemination. miR-21 is a well-known oncogenic miRNA, and previous studies have shown that it is overexpressed in lung, ovarian, and colon cancer as well as in astrocytoma 11. The role of miR-21 in tumor cell dissemination has already been demonstrated in hepatocellular carcinoma (HCC), and it is related to decreased RECK, PTEN and PDCD4 expression 25,26.
Ribas et al. were the first to describe a role for miR-21 in PC showing that miR-21 expression regulation by androgen increases cell proliferation and androgen-independent cell growth 27 but there is only one study similar to ours relating miR-21 and biochemical recurrence in PC 12. miR-21 up-regulation was also related to androgen-independent cell growth in vitro, and its inhibition promotes apoptosis in DU145 cells. More recently, studies have shown increased levels of miR21 in castration resistant PC related to resistance to docetaxel-based chemotherapy illustrating the possibility of transforming miR21 in a serum biomarker for PC aggressiveness 13,28,29.
miR-21 targets important tumor suppressor genes, such as PTEN, RECK and TIMP3. Reversion-inducing cysteine-rich protein with Kazal motifs (RECK) is a tumor and metastasis suppressor gene, which is critical for regulating invasive and metastatic tumor cell activity. RECK which is an enzyme that regulates MMP9, is down-regulated in certain malignancies, and its expression has been positively correlated with cancer patient survival 30. For surgical specimens from men with PC that was surgically treated, we previously reported that miR-21 expression increased relative to decreased RECK expression resulting in unfavorable behavior for this tumor 31. Herein, after miR-21 transfection in DU145 cells, we show decreased RECK expression associated with increased tumor cell invasiveness characterized by increased tumor cells using Matrigel assays.
In conclusion, through a genome-wide association analysis we showed that miRNAs are most frequently overexpressed in PC, acting preferentially as oncogenes. Future studies are necessary to validate all these findings, searching for profiles that could help define aggressiveness for PC. We can also affirm that miR-21 can be considered a key miRNA related to prostate cancer behavior which overexpression is associated to biochemical recurrence likely targeting RECK, promoting tumor cell invasiveness.
Abbreviations
BPH: Benign prostate hyperplasia
BR: Biochemical recurrence
FBS: Fetal bovine serum
HCC: hepatocellular carcinoma
miRNA: micro RNA
MMP9: Metalloproteinase 9
OncomiR: Oncogenic micro RNA
PC: prostate cancer
PCR: Polymerase chain reaction
PDCD4: programmed cell death 4
PSA: Prostate specific antigen
PTEN: Phosphatase and tensin homolog
qRT-PCR: Quantitative real time polymerase chain reaction
RECK: Reversion-inducing cysteine-rich protein with Kazal motifs
RP: Radical prostatectomy
SD: standard deviation
TIMP : Tissue inhibitor of metalloproteinases
tsmiR: Tumor suppressor micro RNA
Acknowledgements
This study was sponsored by CNPq #473540/2010-0.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Kollmeier MA, Zelefsky MJ. How to select the optimal therapy for early-stage prostate cancer. Critical reviews in oncology/hematology. 2012;84(Suppl 1):e6-e15
2. Moreira Leite KR, Camara-Lopes LH, Dall'Oglio MF. et al. Upgrading the Gleason score in extended prostate biopsy: implications for treatment choice. International journal of radiation oncology, biology, physics. 2009;73:353-6
3. Epstein JI, Feng Z, Trock BJ, Pierorazio PM. Upgrading and downgrading of prostate cancer from biopsy to radical prostatectomy: incidence and predictive factors using the modified Gleason grading system and factoring in tertiary grades. European urology. 2012;61:1019-24
4. Osman A. MicroRNAs in health and disease--basic science and clinical applications. Clinical laboratory. 2012;58:393-402
5. Lu J, Getz G, Miska EA. et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435:834-8
6. Leite KR, Tomiyama A, Reis ST. et al. MicroRNA-100 expression is independently related to biochemical recurrence of prostate cancer. The Journal of urology. 2011;185:1118-22
7. Leite KR, Sousa-Canavez JM, Reis ST. et al. Change in expression of miR-let7c, miR-100, and miR-218 from high grade localized prostate cancer to metastasis. Urologic oncology. 2011;29:265-9
8. Epstein JI, Allsbrook WC Jr, Amin MB, Egevad LL, Committee IG. The 2005 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma. The American journal of surgical pathology. 2005;29:1228-42
9. Lexander H, Franzen B, Hirschberg D. et al. Differential protein expression in anatomical zones of the prostate. Proteomics. 2005;5:2570-6
10. Irizarry RA, Hobbs B, Collin F. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4:249-64
11. Nair VS, Maeda LS, Ioannidis JP. Clinical outcome prediction by microRNAs in human cancer: a systematic review. Journal of the National Cancer Institute. 2012;104:528-40
12. Li T, Li RS, Li YH. et al. miR-21 as an independent biochemical recurrence predictor and potential therapeutic target for prostate cancer. The Journal of urology. 2012;187:1466-72
13. Zhang HL, Yang LF, Zhu Y. et al. Serum miRNA-21: elevated levels in patients with metastatic hormone-refractory prostate cancer and potential predictive factor for the efficacy of docetaxel-based chemotherapy. The Prostate. 2011;71:326-31
14. Volinia S, Calin GA, Liu CG. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proceedings of the National Academy of Sciences of the United States of America. 2006;103:2257-61
15. Porkka KP, Pfeiffer MJ, Waltering KK, Vessella RL, Tammela TL, Visakorpi T. MicroRNA expression profiling in prostate cancer. Cancer research. 2007;67:6130-5
16. Ozen M, Creighton CJ, Ozdemir M, Ittmann M. Widespread deregulation of microRNA expression in human prostate cancer. Oncogene. 2008;27:1788-93
17. Tong AW, Fulgham P, Jay C. et al. MicroRNA profile analysis of human prostate cancers. Cancer Gene Ther. 2009;16:206-16
18. Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Developmental biology. 2007;302:1-12
19. Schaefer A, Jung M, Mollenkopf HJ. et al. Diagnostic and prognostic implications of microRNA profiling in prostate carcinoma. International journal of cancer Journal international du cancer. 2010;126:1166-76
20. Kattan MW, Wheeler TM, Scardino PT. Postoperative nomogram for disease recurrence after radical prostatectomy for prostate cancer. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 1999;17:1499-507
21. Haffner MC, De Marzo AM, Yegnasubramanian S, Epstein JI, Carter HB. Diagnostic Challenges of Clonal Heterogeneity in Prostate Cancer. Journal of clinical oncology: official journal of the American Society of Clinical Oncology. 2014
22. Haffner MC, Mosbruger T, Esopi DM. et al. Tracking the clonal origin of lethal prostate cancer. J Clin Invest. 2013;123:4918-22
23. Lucca I, Shariat SF, Briganti A. et al. Validation of tertiary Gleason pattern 5 in Gleason score 7 prostate cancer as an independent predictor of biochemical recurrence and development of a prognostic model. Urologic oncology. 2014
24. Algarra R, Barba J, Merino I. et al. Prognostic value of seminal vesicle involvement due to prostate cancer in radical prostatectomy specimens. Actas urologicas espanolas. 2014
25. Zhu Q, Wang Z, Hu Y. et al. miR-21 promotes migration and invasion by the miR-21-PDCD4-AP-1 feedback loop in human hepatocellular carcinoma. Oncol Rep. 2012;27:1660-8
26. Zhou L, Yang ZX, Song WJ. et al. MicroRNA-21 regulates the migration and invasion of a stem-like population in hepatocellular carcinoma. International journal of oncology. 2013;43:661-9
27. Ribas J, Ni X, Haffner M. et al. miR-21: an androgen receptor-regulated microRNA that promotes hormone-dependent and hormone-independent prostate cancer growth. Cancer research. 2009;69:7165-9
28. Watahiki A, Macfarlane RJ, Gleave ME. et al. Plasma miRNAs as Biomarkers to Identify Patients with Castration-Resistant Metastatic Prostate Cancer. International journal of molecular sciences. 2013;14:7757-70
29. Maugeri-Sacca M, Coppola V, Bonci D, De Maria R. MicroRNAs and prostate cancer: from preclinical research to translational oncology. Cancer journal. 2012;18:253-61
30. Chen Y, Tseng SH. The potential of RECK inducers as antitumor agents for glioma. Anticancer research. 2012;32:2991-8
31. Reis ST, Pontes-Junior J, Antunes AA. et al. miR-21 may acts as an oncomir by targeting RECK, a matrix metalloproteinase regulator, in prostate cancer. BMC urology. 2012;12:14
Author contact
![]() Corresponding author: Katia Ramos Moreira Leite, MD, PhD, Av. Dr. Arnaldo 455, Room 2145 01246-903, Sao Paulo, Brazil. Tel.: +55 11 30617183; katiaramoscom.br
Corresponding author: Katia Ramos Moreira Leite, MD, PhD, Av. Dr. Arnaldo 455, Room 2145 01246-903, Sao Paulo, Brazil. Tel.: +55 11 30617183; katiaramoscom.br

 Global reach, higher impact
Global reach, higher impact