Impact Factor ISSN: 1837-9664
J Cancer 2013; 4(7):566-572. doi:10.7150/jca.6638 This issue Cite
Research Paper
TGF-β2: A Novel Target of CD44-Promoted Breast Cancer Invasion
1. Department of Genetics,
2. Department of Family Medicine and Public Health, College of Medicine and Health Sciences, Sultan Qaboos University, Muscat, Sultanate Of Oman
3. Department of Ob GYN, Louisiana State University Health Sciences Center, New Orleans, Louisiana, USA
Received 2013-5-7; Accepted 2013-6-18; Published 2013-8-16
Abstract
We have developed a tetracycline (tet)-off regulated expression of CD44s gene in the breast cancer (BC) cell line MCF-7 (B5 clone) and identified TGF-β2 (Transforming Growth Factor beta-2; 3 fold induction) as a potential CD44-downstream transcriptional target by microarray analysis. To further validate this finding, the same RNA samples, used for microarray analysis and their corresponding protein lysates, collected from the BC cell line MCF-7-B5, were examined for CD44 expression in the presence of HA. Our results showed that TGF-β2 mRNA levels were significantly elevated following the removal of tetracycline at 18, 24, and 48 h post-HA stimulation compared to the parental cells. Furthermore, the TGF-β2 precursor protein increased in a time-dependent pattern upon HA-stimulation and in the absence of tetracycline. More interestingly, inhibition of CD44 gene by RNAi method decreased TGF-β2 expression upon HA-stimulation, and subsequently inhibited BC cell invasion in vitro. In addition to identifying TGF-β2 as a target for HA/CD44 signaling, this data suggests that ATF/CREB might be a potential transcription factor linking HA/CD44 activation to TGF-β2 transcription and additional experiments are required for a better understanding of the molecular mechanisms underpinning the novel function of the CD44/ TGF-β2 signaling pathway in breast cancer metastasis.
Keywords: Hyaluronan, CD44, TGF-β2, CRE/ATF, Tetracycline-inducible, Breast cancer, Metastasis, MCF-7-B5 clone
Introduction
BC is the most common cancer afflicting females worldwide. There are numerous risk factors involved, including but not limited to: early and extended hormonal exposure and the exposure to ionizing radiation, especially at early ages1-2. Metastasis is the most devastating aspect of tumor malignancy. Along with tumor formation, it is the primary cause of treatment failure and cancer-related death. BC cells metastasize mainly to the bone, liver, lung and the brain, but they also invade, and exfoliate into, body cavities, for example as malignant pleural effusions3. Although markers such as steroid receptor status, tumor size, nodal status and age at the time of diagnosis have been used for several years; none have demonstrated remarkable prognostic value4. Key to increasing the patient's quality of life and long-term survival is to develop improved early detection methods and therapies for controlling cancer progression5. The current therapeutic measures used to treat cancer metastasis are limited in efficacy and also have multiple adverse effects stemming from their lack of specificity. As a crucial first step, we seek to gain an understanding of the unique disease processes associated with BC metastasis.
A crucial step involved in cancer cell metastasis requires the interaction of several classes of molecules between the tumor cell and the surrounding tissue including cell adhesion molecules6. The major receptor for hyaluronic acid (HA), CD44, has been shown to initiate the metastatic spread of tumor cells7. HA-CD44 interaction initiates a downstream cascade of events that modulate cell adhesion, motility, proliferation, and survival8. However, the complete biomolecular pathway of CD44 activation has not been elucidated. CD44 is the most studied alternatively-spliced gene in cancer and is associated with wide structural diversity9-10. The standard form of CD44, CD44s, lacks the 10 variably spliced exons and codes for a typical type1 transmembrane protein11. As a transmembrane matrix receptor with a cytoplasmic domain, CD44s has the basic structure to function in signal transduction. HA binding initiates extracellular clustering of CD44s, resulting in the activation of kinases12. CD44s can serve as, a co-receptor physically linked to other classical signaling receptors13, a docking protein for other proteins14, and the trans-membrane domain of CD44s can be cleaved and translocated to the nucleus, where it functions as a transcription factor15.
Although a number of studies have implicated CD44s in breast tumor invasion, the underlying mechanism is nascent16. Nevertheless, recent studies carried out in our laboratory and others provide strong evidence, using both in vitro and in vivo models, that CD44s-mediated adhesion and signaling are required for cell growth and the dissemination of breast-derived tumors5,17-19. In the present study we demonstrate a novel relationship between HA/CD44s signaling and the transcription of TGF- β2 in BC cells. To better understand the molecular mechanisms implicated in CD44s-mediated breast tumor invasion and metastasis, various molecular and functional techniques were used and TGF- β2, a CD44s-downstream transcriptional target gene was identified, which is involved in CD44-promoted BC cell motility.
Materials and Methods
Cells
The MCF7 and MDA-MB-231 breast cancer cell lines were purchased from American Type Culture Collection (Manassas, VA). All cells were cultured in DMEM media containing 10% (v/v) FBS, 2 mM L-glutamine and 1 mM sodium pyruvate (Gibco, Gaithersburg, MD). The MCF7-CD44 tet-off (MCF7-B5) BC cell line with regulated CD44s expression was maintained in DMEM supplemented with 10% fetal bovine serum, 2.5 µg/ml doxycycline (dox), 100 µg/ml G418 (Roche Diagnostics Ltd. (GmBH), Lewes, UK) and 1 µg/ml puromycin (5).
Chemicals and reagents
All chemicals were supplied by the Sigma Chemical Co. (St Louis, MO), unless otherwise stated. Hyaluronan of molecular weight 220 kDa and medical grade purity was purchased from Lifecore Biomedical Inc. (Chaska, MN).
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
Total RNA was harvested from MCF7-B5 cells cultured in the presence (+dox) and absence (-dox) of dox following stimulation with 100 µg/ml HA (MW 220 kDa) for 18, 24, and 48 h using the Qiagen RNeasy Mini Kit (Cat no. 74104) as per manufacturer's instructions. For RT-PCR analysis, 1.0 µg of total RNA was reverse transcribed using standard reagents. Samples were incubated in the PTC-200 Thermal Cycler for reverse transcription at 50°C for 30 min. The initial PCR activation step at 95°C for 5 min was followed by 27 cycles. Each cycle consisted of 94˚C for 30 seconds, 55˚C for 30 seconds, and 68˚C for 1 min. Final annealing was at 68˚C for 10 min. The annealing temperatures for various genes were calculated according to their sequence and were optimized. The annealing temperatures and oligonucleotide primers used for CD44s, TGF-β2 and GAPDH genes are listed in Table 1. The PCR products were examined by electrophoresis in a 2% agarose gel, containing 0.2 μg/ml ethidium bromide.
Temperatures and Oligonucleotide Primers used for PCR
| Genes | Temperature | Oligonucleotide Primers |
|---|---|---|
| CD44 | 55˚C | TTT GCA TTG CAG TCA ACA GTC (sense) TTA CAC CCC AAT CTT CAT GTC CAC (antisense) |
| TGF-β2 | 56˚C | GGA GGT TTA CAA AAT AGA CAT GCC (sense) AAG ACT CTG AAC TCT GCT TTC ACC (antisense) |
| GAPDH | 58˚C | ACC ACA GTC CAT GCC ATC AC (sense) TCC ACC ACC CTG TTG CTG TA (antisense) |
Western-blot analysis
The CD44 expression in MCF7-B5 cells was induced by removal of dox at different time points, and protein lysates were collected as previously described (5). Protein lysates were collected centrifuged and the final concentration was determined using the Bicinchoninic Acid (BCA) protein assay reagent (Pierce, Rockford, IL). Protein samples (60 µg) were boiled for 5 minutes in an equal volume of reducing buffer (5 mmol/L of Tris/HCl (pH 7.4), 4% (w/v) sodium dodecyl sulfate, 20% (v/v) glycerol, 10% (v/v) mercaptoethanol, 0.2% (w/v) bromophenol blue), resolved on 12% polyacrylamide gels, and electroblotted onto a nitrocellulose membrane. The membrane was probed with mouse anti-human CD44s monoclonal antibody (1:500 dilution; R&D Systems,) and then treated with a sheep anti-mouse IgG/horseradish peroxidase conjugate (1:2000 dilution; Santa Cruz Biotechnology). The same samples were also examined for TGF-β2 using rabbit anti-human TGF-β2 polyclonal antibody (1:2000 dilution; Santa Cruz Biotechnology) and then treated with sheep anti-rabbit IgG/horseradish peroxidase conjugate (1:2000 dilution; Santa Cruz Biotechnology). Immunoreactivity was detected using chemiluminescence (Super signal, Pierce). Equal loading of protein samples was assessed by reprobing the membranes with β-tubulin antibody (1:2000 dilution; Sigma).
RNAi-mediated depletion of CD44s and TGF-β2
Oligonucleotides specific for human CD44 and TGF-β2, along with a Silencer® Negative Control siRNA were synthesized commercially (Ambion, TX) for siRNA inhibition of CD44 and TGF-β2. The sequences used to inhibit CD44 are, 5'-GGAAAUGGUGCAUUUGGUGTT-3' (sense) and 5'-TTCCUUUACCACGUAAACCAC-3' (antisense), and for TGF-β2 are, 5'-CACUCGAUAUGGACCAGUUTT-3' (sense) and 5'-TTGUGAGCUAUACCUGGUCAA-3' (antisense). Cells were seeded and grown to 50% confluency, washed twice with sterile PBS, then incubated at 37°C for 6 h, with a transfection cocktail compromising OPTI-MEM1, Lipofectamin-2000 (Invitrogen, NY) and the siRNA anti-CD44, anti- TGF-β2 or scrambled oligonucleotides at a final concentration of 100 nM or 200nM. After incubation, the media was replaced with 20% (v/v) FBS-enriched growth media, and subsequently incubated overnight at 37°C. Transfected cells were retransfected with CD44 siRNA transfection cocktail as described above. Protein lysates were harvested within 24h after the 2nd hit, and depletion of protein expression was confirmed by immunoblotting analysis.
Cell invasion assay
MCF7-B5 cells were cultured in the absence of (dox) and the presence of HA (100μg/ml) for a period of 24 hr for continuous induction of CD44 expression. Cells were treated with TGF-β2 specific siRNA, as described above, to inhibit TGF-β2 expression, and 24 hr later the cells were washed twice with sterile PBS and harvested by trypsinization. The harvested cells were then resuspended in DMEM supplemented with 0.5% BSA and plated in Millicell culture inserts (12 µm pore size, Millipore, MA), which were previously coated with a thin layer of 200 µg/ml of Matrigel™ (BD Biosciences, MA). The inserts containing the cells were placed into a tissue culture dish (lower chamber) with the “attracting medium” containing the MCF7-B5 special medium, supplemented with 100μg/ml of HA. Cells were incubated for 22h at 37°C and the Millicell culture insert was removed to allow wiping quickly the upper surface of the insert with a cotton swab in order to remove non-invasive cells. The cell culture inserts were dried under laminar flow hood for 4 h, and the cells that invaded and are present on the bottom of the filter were stained using the Diff-Quick staining kit, according to the manufacturers' protocol (Dade Behring Inc.), and the stained cells were counted under a phase-contrast microscope, equipped with ocular grids.
Statistical analysis of data
Differences between data points in functional assays (invasion assay) were assessed for statistical significance by SPSS program using Student's t-test two-tailed. Data was presented as Mean ± Standard deviation of at least triplicates from three experiments. The difference is considered statistically significant when p <0.05.
Results
In Vitro Characterization of the Tet Off-Regulated CD44s Expression System
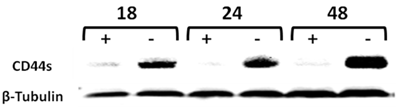
We have previously generated a tet-off-regulated CD44s expression system in the parental MCF-7 cell line, expressing minimal amounts of CD44s to obtain an inducible MCF7-B5 BC cell line (10). To validate that this system functions properly in vitro, upregulation of both CD44s mRNA and protein was examined in the MCF7-B5 clone throughout a 48-h time course. Total RNA and protein lysates were collected from MCF7F-B5 cells at 18, 24, and 48 h in the presence and absence of dox and examined for CD44s expression by western blot. In the absence of dox, the level of expression of CD44s showed a time-dependent increase and was maintained up to 48 h. A single immunoreactive band was detected at a molecular mass equivalent to 85 kDa, which is consistent with the presence of CD44s (Fig. 1). However in the presence of dox, the MCF7F-B5 cells cultured showed barely detectable CD44s expression, indicating that the tet-regulated CD44s suppression system was properly functional in the MCF7F-B5 cell line.
Time-course western-blot validation of the tetracycline (tet) regulated CD44 expression in MCF7F-B5 breast cancer cell line. Protein lysates were collected at different time points (18, 24 and 48hrs) following withdrawal of tet, in the presence of hyaluronan (HA). The level of expression of CD44s showed a time-dependent increase and was maintained up to 48 hrs.
Validation of TGF-β2 as a Transcriptional Target of CD44 Signaling
The mechanisms responsible for upregulation and expression of the downstream target genes of CD44s-promoted metastasis in vivo are poorly understood. Therefore, we have used the tet off-regulated CD44s expression system in the MCF7-B5 cell line to identify novel transcriptional targets of HA/CD44s signaling. Microarray analysis led to the identification of a number of potential CD44s-downstream target genes10,19. Among these target genes, the microarray data obtained showed an upregulation of a 2.9-fold in TGF-β2 mRNA transcript levels, indicating that upon induction of CD44 in MCF7-B5 cells in the presence of dox, TGF-β2 mRNA expression increased by 2.9-fold10. TGF-β2 was chosen for further investigation on the basis that it is a growth factor that promotes BC metastasis, angiogenesis, and cytoskeletal remodeling22-21. Two strategies were employed to validate TGF-β2 as a potential downstream target of CD44s: i) Induction of CD44s upregulates TGF-β2: Time-course RT-PCR and western-blotting analyses were carried out on the same RNA and protein samples collected from MCF7-B5 used for microarray analyses. TGF-β2 RNA levels were significantly induced in the MCF7-B5 cells following the removal of dox at 18, 24, 48 h post-HA stimulation when compared to the MDA-231 cell lines (Fig. 2). Similarly, TGF-β2 protein was upregulated when CD44s expression was induced at different time points (Fig. 3); and ii) RNAi inhibition of CD44s downregulates TGF-β2: When the expression of CD44s was inhibited using the RNAi method, it showed that 24 h inhibition of CD44s was parallel to a significant decrease in TGF-β2 (Fig. 4).
Time-course RT-PCR validation of TGF-beta2 as a potential target of CD44 in MCF7F-B5 breast cancer cell line. RNA samples were collected at different time points (18, 24 and 48hrs) following withdrawal of tet in the presence of hyaluronan (HA). The highly metastatic breast cancer cell line MDA-231 which expresses high levels of CD44 as well as TGF-beta2 was used as a positive control.
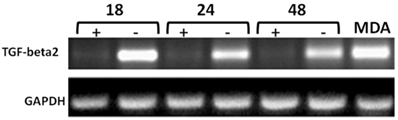
Time-course western blot validation of TGF-beta2 as a potential target of CD44 in MCF7F-B5 breast cancer cell line. Protein lysates were collected at different time points (18, 24 and 48hrs) following withdrawal of tet in the presence of hyaluronan (HA). The levels of expression of TGF-beta2 showed a time-dependent increase that was parallel to the increase of CD44 expression.
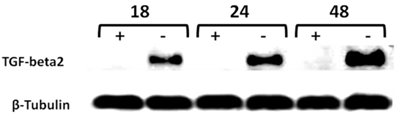
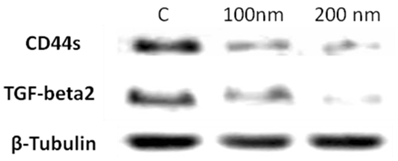
Validation of TGF-beta2 as a potential target of CD44 in MCF7F-B5 breast cancer cell line using RNAi strategy. As described in materials and methods, CD44 expression was inhibited by specific siRNA oligo-nucleotides (100 and 200 nM), protein lysates were collected, and western blot analysis was performed. Inhibition of CD44 was shown to downregulate TGF-beta2. C; represents the control untreated MCF7F-B5 cells.

TGF-β2 underpins CD44 metastatic function in breast cancer cells
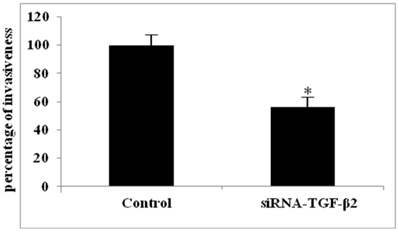
The functional relevance of HA/CD44 induced TGF-β2 expression to BC cell invasiveness was investigated using the MCF7-B5 clone expressing CD44. Specific inhibition of CD44 expression by siRNA resulted in a specific decrease in TGF-β2 expression that correlated with a significant loss of BC cell invasiveness (data not shown). More interestingly, the invasiveness of MCF7-B5 cells post HA stimulation, was significantly attenuated following RNAi-mediated reduction of TGF-β2 expression in these cells (Fig. 5). This reduction in invasiveness ability of MCF7-B5 transfected with specific anti- TGF-β2 RNAi was determined in comparison to cell controls, which were treated with scrambled RNAi, using t-test (Fig. 5).
Functional validation of TGF-β2 expression in breast cancer cell line. Invasion of MCF7F-B5 breast cancer cells as demonstrated by Boyden chamber invasion assay. Graphical representation of the percentage of invasiveness of untreated MCF7F-B5 cells (control) compared to anti-TGF-β2 siRNA-treated MCF7F-B5 cells (student's two-tailed t-test, *P<0.001).
Discussion
Previous studies have indicated that HA/CD44s signaling plays an important role in BC metastasis10. However, the signaling events that underpin HA/CD44s-promoted BC metastasis are poorly understood. The purpose of this study was to identify novel transcriptional targets of HA/CD44s signaling using a tetracycline-regulated (Tet-off) CD44s expression system in a weakly invasive BC cell line, MCF7F-B5, that normally expresses low or no endogenous CD44s. Following validation of the tet-regulated-CD44s expression system, RNA was extracted from HA-stimulated MCF7F-B5 cells and further examined by microarray analysis to identify possible downstream-transcriptional targets of HA/CD44s signaling. We have identified a number of potential downstream target genes and have previously validated Cortactin and Survivin as a target gene of CD44s-promoted breast tumor cell invasion10,19. To better understand CD44 signal transduction mechanisms involved in breast tumor cell invasion and metastasis, the present study employed various molecular approaches to validate the transforming growth factor-β2, TGF-β2 (2.9 fold induction by microarray) as a novel potential CD44s-target mediating BC invasion. TGF-β2 belongs to a large family of polypeptide growth factors that is involved in regulating multiple biological processes including cell proliferation, angiogenesis, immune response, apoptosis, extracellular matrix synthesis, and differentiation21. There are three isoforms of TGF, which are all secreted as latent precursor molecules22. The latent forms of TGF are activated by proteolytic cleavage, pH changes, or interaction with integrins. The active forms of TGF bind to specific receptors, transmembrane serine/threonine kinases, which have been found to activate the SMAD signaling cascade. Bourguignon et al (2008) have established a molecular link between CD44-signaling and TGF-β223; the study provided evidence that activation of TGF-beta2 is an essential CD44-downstream event required for tumor cell survival and metastasis23. Furthermore, a polymorphism in the promoter of TGF-β2 that enhances expression of the protein was associated with lymph node metastasis in breast cancer patients, pointing to a role of TGF-β2 in the process of invasion24.
Embryogenesis and metastasis share one common major obstacle that must be overcome by epithelial cells: survival following the detachment of the cell from the primary site and migration through the extracellular matrix. The process of epithelial-mesenchymal transition (EMT) enables epithelial cells to overcome these restrictions, paving the stage for physiologically programmed cell migration/invasion. Several studies have provided evidence that EMT plays an important role in cancer invasion and metastasis. During the process of EMT, cancer cells gain motility properties using developmental processes that involve reorganization of the actin cytoskeleton, and subsequently the formation of membrane protrusions required for the cell to invade. Takahashi et al (2010) have recently demonstrated that the hyaluronan-CD44-moesin complex associated with TGF-β receptor II and clathrin at actin microdomains, activated TGF-β signaling and caused both cell-cell dissociation and increased cellular motility through actin remodeling25. Other studies have demonstrated that factors regulating developmental EMT, including the TGF-β signaling pathway, play roles in breast tumor progression25-26. Also, activation of the TGF-β pathway in human cancer cell lines and in mouse tumor models can induce EMT and promote tumor cells to invade the ECM in culture and to metastasize to distant organs in mice26-28.
Here, we provide evidence that TGF-β2 is a downstream transcriptional target of CD44-mediated MCF-7 cell invasion. The next question is how does CD44 induce the transcription of TGF-β2? CD44 can induce transcription of TGF-β2 via activation of the transcription factor cAMP response element-binding protein (CREB). Previous study showed that CD44 can undergo sequential metalloprotease- and γ-secretase-mediated proteolytic cleavage, releasing the CD44 intracellular domain (CD44-ICD), which translocates to the nucleus and induce gene transcription29. In fact, recent study clearly demonstrated that the CD44-ICD protein translocated to the nucleus, bound to the transcription factor CREB, and increased S133 phosphorylation and CREB-mediated gene transcription30, which is known to transactivate TGF-beta231. It has been previously reported that a cAMP response element/activating transcription factor (CRE/ATF) site and an E-box motif are located within the TGF-beta2 gene promoter just upstream the transcription start site, and are essential for the transcription of the TGF-beta2 gene31.
In conclusion, our results clearly show that TGF-β2 was upregulated when CD44 was induced in the MCF-7B5 clone. Furthermore, inhibition of CD44 gene by RNAi method decreased TGF-beta2 expression upon HA-stimulation, and subsequently inhibited BC cell invasion in vitro. More interestingly, previous studies have demonstrated that overexpression of TGF- β2 promotes invasion of various tumors, including ovarian cancer32, melanoma33, pancreatic cancer34 and malignant epidermal keratinocytes35. Therefore, our data put together strongly suggest that TGF-β2 is a potential downstream target gene mediating CD44s-promoted breast tumor cell invasion. Ongoing experiments in our laboratory using various experimental strategies are aiming to validate in vivo, the functional relevance of the CD44/CREB/TGF-β2 signaling pathway in promoting breast tumour growth and metastasis.
Acknowledgements
This work was supported by His Majesty Sultan Qaboos University Strategic Grant SR/MED/GNET/10/01 (A.O.). We thank Dr. David J. Waugh for supplying the CD44-B5 clone.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ronckers CM, Erdmann CA, Land CE. Radiation and BC: a review of current evidence. BC Res. 2005;7:21-32
2. Carmichael A, Sami AS, Dixon JM. BC risk among the survivors of atomic bomb and patients exposed to therapeutic ionizing radiation. Eur J Surg Oncol. 2003;29:475-9
3. Shulman LN, Sugarbaker DJ. Malignant effusions. In: (ed.) Harris JR, Lippman ME, Morrow M, Hellman S. Diseases of the breast. Philadelphia: Lippincott-Raven. 1996:833-840
4. Berner HS, Suo Z, Risberg B, Villman K, Karlsson MG, Nesland JM. Clinicopathological associations of CD44 mRNA and protein expression in primary breast carcinomas. Histopathology. 2006;42:546-554
5. Ouhtit A, Elmageed Z, Abdraboh M, Lioe T, Raj M. In vivo Evidence for the Role of CD44s in Promoting BC Metastasis to the Liver. American Journal of Pathology. 2007;171:2033-2039
6. Liotta LA, Steeg PS, Stetler-Stevenson WG. Cancer metastasis and angiogenesis: an imbalance of positive and negative regulation. Cell. 1991;64:327-336
7. Toole BP. Hyaluronan: from extracellular glue to pericellular cue. Nature Reviews Cancer. 2004;4:528-539
8. Marhaba R, Zöller M. CD44 in Cancer Progression: Adhesion, Migration and Growth Regulation. Journal of Molecular Histology. 2004;35(3):211-231
9. Bajorath J. Molecular organisation, structural features, and ligand binding characteristics of CD44, a highly variable cell surface glycoprotein with multiple functions. Proteins. 2000;39:103-111
10. Hill A, McFarlane S, Mulligan K. et al. Cortactin underpins CD44-promoted invasion and adhesion of BC cells to bone marrow endothelial cells. Oncogene. 2006;25:6079-91
11. Naor D, Sionov RV, Ish-Shalom D. CD44: structure, function, and association with the malignant process. Adv Cancer Res. 1997;71:241-319
12. Ito T, Williams JD, Fraser DJ, Phillips AO. Hyaluronan regulates transforming growth factor-β1 receptor compartmentalization. J Biol Chem. 2004;279:25326-25332
13. Lesley J, Hyman R. CD44 structure and function. Front Biosci. 1998;3:616-630
14. Lopez JI, Camenisch TD, Stevens MV, Sands BJ, McDonald J, Schroeder JA. CD44 attenuates metastatic invasion during BC progression. Cancer Res. 2005;65:6755-63
15. Mori H, Tomari T, Koshikawa N. et al. CD44 directs membrane-type 1 matrix metalloproteinase to lamellipodia by associating with its hemopexin-like domain. Embo J. 2002;21:3949-3959
16. Murai T, Miyazaki Y, Nishinakamura H. et al. Engagement of CD44 promotes Rac activation and CD44 cleavage during tumor cell migration. J Biol Chem. 2004;279:4541-4550
17. Murakami D, Okamoto I, Nagano O. et al. Presenilin-dependent g-secretase activity mediates the intramembranous cleavage of CD44. Oncogene. 2003;22:1511-1516
18. Ouhtit A, Matrougui K, Bengrine A, Koochekpour S, Zerfaoui M, Yousief Z. Survivin is Not only a Death Encounter but a Survival Protein for Invading Tumor Cells. Front Biosci. 2007;12:1260-1270
19. Abdraboh ME, Gaur RL, Hollenbach AD, Sandquist D, Raj MH, Ouhtit A. Survivin, a Novel Target of CD44-Promoted Breast Tumor Invasion. The American Journal of Pathology. 2012;180:1324
20. Buck MB, Fritz P, Dippon J, Zugmaier G, Knabbe C. Prognostic Significance of Transforming Growth Factor Β Receptor II in Estrogen Receptor-Negative BC Patients. Clinical Cancer Research. 2004;10:491-8
21. Huang S, Huang J. TGF-beta Control of Cell Proliferation. Journal of Cellular Biochemistry. 2005;96:447-462
22. Schlingensiepen KH, Fischer-Blass B, Schmaus S, Ludwig S. Antisense therapeutics for tumor treatment: the TGF-β2 inhibitor AP 12009 in clinical development against malignant tumors. Recent Results Cancer Research. 2008;177:137-50
23. Bourguignon LY, Singleton PA, Zhu H, Zhou B. Hyaluronan promotes signaling interaction between CD44 and the transforming growth factor beta receptor I in metastatic breast tumor cells. J Biol Chem. 2002;277:39703-12
24. Buck MB, Knabbe C. TGF-β signaling in breast cancer. Ann N Y Acad Sci. 2006;1089:119
25. Takahashi E, Nagano O, Ishimoto T. et al. Tumor necrosis factor-alpha regulates transforming growth factor-beta-dependent epithelial-mesenchymal transition by promoting hyaluronan-CD44-moesin interaction. J Biol Chem. 2010;285:4060-73
26. Zhang J, Liang Q, Lei Y. et al. SOX4 Induces Epithelial-Mesenchymal Transition and Contributes to Breast Cancer Progression. Cancer Res. 2012;72:4597-608
27. Oft M, Heider KH, Beug H. TGFbeta signaling is necessary for carcinoma cell invasiveness and metastasis. Curr Biol. 1998;8:1243-52
28. Janda E, Lehmann K, Killisch I. et al. Ras and TGF[beta] cooperatively regulate epithelial cell plasticity and metastasis: dissection of Ras signaling pathways. J Cell Biol. 2002;156:299-313
29. Okamoto I, Kawano Y, Murakami D. et al. Proteolytic release of CD44 intracellular domain and its role in the CD44 signaling pathway. J Cell Biol. 2001;155:755-62
30. De Falco V, Tamburrino A, Ventre S. et al. CD44 proteolysis increases CREB phosphorylation and sustains proliferation of thyroid cancer cells. Cancer Res. 2012;72:1449-58
31. Kingsley-Kallesen ML, Kelly D, Rizzino A. Transcriptional regulation of the transforming growth factor-beta2 promoter by cAMP-responsive element-binding protein (CREB) and activating transcription factor-1 (ATF-1) is modulated by protein kinases and the coactivators p300 and CREB-binding protein. J Biol Chem. 1999;274:34020-8
32. Do TV, Kubba LA, Du H, Sturgis CD, Woodruff TK. Transforming growth factor-beta1, transforming growth factor-beta2, and transforming growth factor-beta3 enhance ovarian cancer metastatic potential by inducing a Smad3-dependent epithelial-to-mesenchymal transition. Mol Cancer Research. 2008;6:695-705
33. Zhang C, Zhang F, Tsan R, Fidler IJ. Transforming growth factor-beta2 is a molecular determinant for site-specific melanoma metastasis in the brain. Cancer Res. 2009;69:828-35
34. Hilbig A, Oettle H. Transforming growth factor beta in pancreatic cancer. Curr Pharm Biotechnol. 2011;12(12):2158-64
35. Davies M, Prime SS, Eveson JW, Price N, Ganapathy A, D'Mello A, Paterson IC. Transforming growth factor-β enhances invasion and metastasis in Ras-transfected human malignant epidermal keratinocytes. Int J Exp Pathol. 2012;93(2):148-56
Author contact
Corresponding author: Allal Ouhtit, PhD, MPh. Head of Department, Genetics, College of Medicine and Health Sciences, Sultan Qaboos University; P.O Box: 35, Postcode: 123, Muscat, Sultanate Of Oman. E-mail: aouhtitedu.om

 Global reach, higher impact
Global reach, higher impact